

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
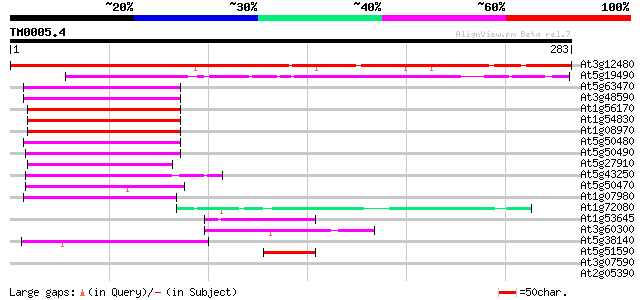
At3g12480 unknown protein 287 4e-78
At5g19490 putative protein 156 1e-38
At5g63470 transcription factor Hap5a-like protein 59 2e-09
At3g48590 transcription factor Hap5a 59 2e-09
At1g56170 transcription factor, putative 59 2e-09
At1g54830 unknown protein 59 2e-09
At1g08970 transcription factor like protein 59 2e-09
At5g50480 transcription factor Hap5a-like 58 7e-09
At5g50490 unknown protein 57 1e-08
At5g27910 transcription factor - like protein 53 2e-07
At5g43250 unknown protein 51 8e-07
At5g50470 putative protein 49 4e-06
At1g07980 unknown protein 46 3e-05
At1g72080 hypothetical protein 44 1e-04
At1g53645 unknown protein 43 2e-04
At3g60300 unknown protein 43 2e-04
At5g38140 putative protein 42 3e-04
At5g51590 unknown protein 41 6e-04
At3g07590 putative small nuclear ribonucleoprotein (Sm-D1) 39 0.003
At2g05390 putative retroelement pol polyprotein 39 0.004
>At3g12480 unknown protein
Length = 293
Score = 287 bits (735), Expect = 4e-78
Identities = 168/300 (56%), Positives = 202/300 (67%), Gaps = 24/300 (8%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSK+LELFLQDLCDRTYEITL+RGAK
Sbjct: 1 MRKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKSLELFLQDLCDRTYEITLERGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSH--GHGHSEPSADDRTIPKRRKAAGDDCN 118
T++SLHLKHCV+ YNVFDFLR+VVSKVPDY H G GH + + DDR+I KRRK D+ N
Sbjct: 61 TVSSLHLKHCVERYNVFDFLREVVSKVPDYGHSQGQGHGDVTMDDRSISKRRKPISDEVN 120
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG----RTAVRETPHQAVEPEPCASVQ 174
DSDEE K+ K E+ ++GRG GRGRGRGRGRG + A RE ++ +E E S Q
Sbjct: 121 DSDEEYKKSKTQEIGSAKTSGRG-GRGRGRGRGRGGRAAKAAEREGLNREMEVEAANSGQ 179
Query: 175 QGIQHDTNTDMTIHDSSETKELPK--ENAAVPAESAEFH--------NLDLNANTNE-NE 223
+ N M +SS ++ K + A E + H + DLNA + + NE
Sbjct: 180 P--PPEDNVKMHASESSPQEDEKKGIDGTAASNEDTKQHLQSPKEGIDFDLNAESLDLNE 237
Query: 224 DKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
K A T + T+S EE GW + D+ KM D QLA+LG I+EDEEDYDEEG
Sbjct: 238 TKLAPATGTTTTTTAATDS--EEYSGWPMMDISKM--DPAQLASLGKRIDEDEEDYDEEG 293
>At5g19490 putative protein
Length = 236
Score = 156 bits (394), Expect = 1e-38
Identities = 107/255 (41%), Positives = 137/255 (52%), Gaps = 25/255 (9%)
Query: 29 LAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCVQSYNVFDFLRDVVSKVP 88
+AVP+LVSKALELFLQDLC+ TY++TL RGAKT+N+ HLK CVQ+ NVFDFLRD V+KVP
Sbjct: 1 MAVPLLVSKALELFLQDLCNHTYDVTLSRGAKTVNAFHLKQCVQATNVFDFLRDTVAKVP 60
Query: 89 DYSHGHGHSEPSADDRTIPKRRKAA-GDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRG 147
D G S+ +D++ KRRK G CND D +K +M E+ HT S GRGR RGRG
Sbjct: 61 DL----GGSD--TEDQSATKRRKVVDGSSCNDED-MIKTTQMHEVKHT-SCGRGR-RGRG 111
Query: 148 RGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSSETKELPKENAAVPAES 207
RGR GRT + + E + N ++ D+S K N
Sbjct: 112 RGRSSGRTGSGLSLKFEEDLEDGSPESSRTPSPENGSLSHDDTSWKKVASHNNHHSSNSE 171
Query: 208 AEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLAN 267
+ N DLN +EN D E+Q E P + L ++++M ID
Sbjct: 172 VKVRNFDLNVELDENGDNATW-----------LETQLERSPDYPL-EINEMKIDPDDQQQ 219
Query: 268 LGTHIEEDEEDYDEE 282
DEEDYDEE
Sbjct: 220 ASA---SDEEDYDEE 231
>At5g63470 transcription factor Hap5a-like protein
Length = 250
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/79 (37%), Positives = 48/79 (59%)
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+ P ARIKKIM+ADEDV I+ P+L +KA ELF+ +L R++ + +T+ +
Sbjct: 78 QLPLARIKKIMKADEDVRMISAEAPILFAKACELFILELTIRSWLHAEENKRRTLQKNDI 137
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 138 AAAITRTDIFDFLVDIVPR 156
>At3g48590 transcription factor Hap5a
Length = 234
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/79 (37%), Positives = 48/79 (59%)
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+ P ARIKKIM+ADEDV I+ P+L +KA ELF+ +L R++ + +T+ +
Sbjct: 65 QLPLARIKKIMKADEDVRMISAEAPILFAKACELFILELTIRSWLHAEENKRRTLQKNDI 124
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ ++FDFL D+V +
Sbjct: 125 AAAITRTDIFDFLVDIVPR 143
>At1g56170 transcription factor, putative
Length = 199
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/77 (38%), Positives = 47/77 (60%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ +KA E+F+ +L R + T + +T+ +
Sbjct: 78 PLARIKKIMKADEDVRMISAEAPVIFAKACEMFILELTLRAWIHTEENKRRTLQKNDIAA 137
Query: 70 CVQSYNVFDFLRDVVSK 86
+ +VFDFL D++ +
Sbjct: 138 AISRTDVFDFLVDIIPR 154
>At1g54830 unknown protein
Length = 217
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/77 (38%), Positives = 48/77 (61%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T + +T+ +
Sbjct: 72 PLARIKKIMKADEDVRMISAEAPVVFARACEMFILELTLRSWNHTEENKRRTLQKNDIAA 131
Query: 70 CVQSYNVFDFLRDVVSK 86
V ++FDFL D+V +
Sbjct: 132 AVTRTDIFDFLVDIVPR 148
>At1g08970 transcription factor like protein
Length = 231
Score = 59.3 bits (142), Expect = 2e-09
Identities = 30/77 (38%), Positives = 48/77 (61%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P ARIKKIM+ADEDV I+ PV+ ++A E+F+ +L R++ T + +T+ +
Sbjct: 82 PLARIKKIMKADEDVRMISAEAPVVFARACEMFILELTLRSWNHTEENKRRTLQKNDIAA 141
Query: 70 CVQSYNVFDFLRDVVSK 86
V ++FDFL D+V +
Sbjct: 142 AVTRTDIFDFLVDIVPR 158
>At5g50480 transcription factor Hap5a-like
Length = 202
Score = 57.8 bits (138), Expect = 7e-09
Identities = 30/79 (37%), Positives = 46/79 (57%)
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+ P ARIKKIM+AD DV ++ P++ +KA E+F+ DL R++ + T+ +
Sbjct: 54 QLPLARIKKIMKADPDVHMVSAEAPIIFAKACEMFIVDLTMRSWLKAEENKRHTLQKSDI 113
Query: 68 KHCVQSYNVFDFLRDVVSK 86
+ V S +DFL DVV K
Sbjct: 114 SNAVASSFTYDFLLDVVPK 132
>At5g50490 unknown protein
Length = 186
Score = 57.0 bits (136), Expect = 1e-08
Identities = 32/78 (41%), Positives = 44/78 (56%)
Query: 9 FPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLK 68
FP +RIK+IM+ D DV IA P L+SKA E+F+ DL R++ + T+ +
Sbjct: 37 FPISRIKRIMKFDPDVSMIAAEAPNLLSKACEMFVMDLTMRSWLHAQESNRLTIRKSDVD 96
Query: 69 HCVQSYNVFDFLRDVVSK 86
V +FDFLRD V K
Sbjct: 97 AVVSQTVIFDFLRDDVPK 114
>At5g27910 transcription factor - like protein
Length = 187
Score = 52.8 bits (125), Expect = 2e-07
Identities = 29/73 (39%), Positives = 41/73 (55%)
Query: 10 PAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKH 69
P RIKKIM+ D DV IA P+L+SKA E+F+ DL R++ + T+ ++
Sbjct: 38 PITRIKKIMKYDPDVTMIASEAPILLSKACEMFIMDLTMRSWLHAQESKRVTLQKSNVDA 97
Query: 70 CVQSYNVFDFLRD 82
V +FDFL D
Sbjct: 98 AVAQTVIFDFLLD 110
>At5g43250 unknown protein
Length = 130
Score = 50.8 bits (120), Expect = 8e-07
Identities = 33/100 (33%), Positives = 56/100 (56%), Gaps = 5/100 (5%)
Query: 9 FPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLK 68
FP R+KKIM+ D+D+ KI +++ + ELFL L +++ +T ++ KT+N HL+
Sbjct: 12 FPIGRVKKIMKLDKDINKINSEALHVITYSTELFLHFLAEKSAVVTAEKKRKTVNLDHLR 71
Query: 69 HCVQSYN-VFDFLRDVVSKVPDYSHGHGHSEPSADDRTIP 107
V+ + DFL D +P + H++ S D+ IP
Sbjct: 72 IAVKRHQPTSDFLLD---SLPLPAQPVKHTK-SVSDKKIP 107
>At5g50470 putative protein
Length = 212
Score = 48.5 bits (114), Expect = 4e-06
Identities = 29/86 (33%), Positives = 45/86 (51%), Gaps = 6/86 (6%)
Query: 9 FPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRG------AKTM 62
FP RIKKIM+++ +V + PVL+SKA E+ + DL R++ T++ G + T+
Sbjct: 64 FPLTRIKKIMKSNPEVNMVTAEAPVLISKACEMLILDLTMRSWLHTVEGGRQTLKRSDTL 123
Query: 63 NSLHLKHCVQSYNVFDFLRDVVSKVP 88
+ F FL DVV + P
Sbjct: 124 TRSDISAATTRSFKFTFLGDVVPRDP 149
>At1g07980 unknown protein
Length = 206
Score = 45.8 bits (107), Expect = 3e-05
Identities = 23/77 (29%), Positives = 43/77 (54%)
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
+FP RI++IM++D +I LV+KA E+F++ + Y+ +++ K ++ HL
Sbjct: 109 KFPMNRIRRIMRSDNSAPQIMQDAVFLVNKATEMFIERFSEEAYDSSVKDKKKFIHYKHL 168
Query: 68 KHCVQSYNVFDFLRDVV 84
V + ++FL D V
Sbjct: 169 SSVVSNDQRYEFLADSV 185
>At1g72080 hypothetical protein
Length = 243
Score = 43.9 bits (102), Expect = 1e-04
Identities = 51/193 (26%), Positives = 71/193 (36%), Gaps = 38/193 (19%)
Query: 85 SKVPDYSHGHGHSEPSADDRT--------------IPKRRKAAGDDCNDSDEEVKRGKMP 130
S+ P +S+G G P A T I + AGDD D RG+
Sbjct: 71 SRSPSHSNG-GREPPFATINTEEHLTMQIESPSTIINTEERRAGDD--DGSRSRGRGR-- 125
Query: 131 ELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDS 190
GRGRGRGRGRGRGR R+ R H EP + H
Sbjct: 126 --GRGRGRGRGRGRGRGRGRGRNRSRSRSPSHSNGGREPPTTTLNTEDH----------- 172
Query: 191 SETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGW 250
T ++ + + E+ N ++ A N D K T + E E + +
Sbjct: 173 -LTMQIEPPSTIINRENGTDANDNVVAVNNIESDCKIDETVPPKTDEEILEDEDD----- 226
Query: 251 SLSDVDKMAIDSV 263
S +DK+ I S+
Sbjct: 227 IRSLLDKLPIPSM 239
Score = 28.1 bits (61), Expect = 5.6
Identities = 18/49 (36%), Positives = 18/49 (36%), Gaps = 11/49 (22%)
Query: 132 LSHTGSTGRGRGRGR-----------GRGRGRGRTAVRETPHQAVEPEP 169
L G R R RGR GRGRGR R R H EP
Sbjct: 35 LGRRGGNNRNRSRGRDDNQSPSNGGRGRGRGRSRRRSRSPSHSNGGREP 83
>At1g53645 unknown protein
Length = 523
Score = 43.1 bits (100), Expect = 2e-04
Identities = 27/57 (47%), Positives = 34/57 (59%), Gaps = 2/57 (3%)
Query: 99 PSADDRTIPKRRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRG-RGRGRGR 154
P+ D T PK + +A + + E+ RG+ S G GRGRGRGRG RGRGRGR
Sbjct: 257 PTVKDGT-PKPQLSAEEAGRRARSELSRGEAEGSSVGGRGGRGRGRGRGARGRGRGR 312
Score = 30.4 bits (67), Expect = 1.1
Identities = 18/37 (48%), Positives = 19/37 (50%)
Query: 120 SDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTA 156
S EE R ELS + G G GRGRGRGR A
Sbjct: 269 SAEEAGRRARSELSRGEAEGSSVGGRGGRGRGRGRGA 305
Score = 28.1 bits (61), Expect = 5.6
Identities = 15/34 (44%), Positives = 18/34 (52%), Gaps = 5/34 (14%)
Query: 133 SHTGSTGRGRGRGRGR-----GRGRGRTAVRETP 161
S + S+GRGRGRG G GRG+ V P
Sbjct: 36 SSSDSSGRGRGRGSGEDGGFPAAGRGQFGVNREP 69
>At3g60300 unknown protein
Length = 253
Score = 42.7 bits (99), Expect = 2e-04
Identities = 29/88 (32%), Positives = 37/88 (41%), Gaps = 5/88 (5%)
Query: 99 PSADDRTIPKRRKAAGDDCNDSDEEVKRGKMP--ELSHTGSTGRGRGRGRGRGRGRGRTA 156
P A T + AG+ +EE K + S T GRGR RGRGR RGRG T
Sbjct: 168 PPAHASTSRNEEEEAGESQEQGEEEPKEAESETNSSSSTNRRGRGRWRGRGRSRGRGPTV 227
Query: 157 VRETPHQAVEPEPCASVQQGIQHDTNTD 184
P+ +P +Q +Q D
Sbjct: 228 NERKPN---SQDPRKPTRQWVQRAREGD 252
>At5g38140 putative protein
Length = 191
Score = 42.4 bits (98), Expect = 3e-04
Identities = 29/100 (29%), Positives = 48/100 (48%), Gaps = 6/100 (6%)
Query: 7 TRFPAARIKKIMQADEDVG------KIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
T P +R++KI+++D +V KI+ VP L SKA E F+ ++ R + T +
Sbjct: 56 THLPLSRVRKILKSDPEVKIYVNFQKISCDVPALFSKACEYFILEVTLRAWMHTQSCTRE 115
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPS 100
T+ + V++ +DFL D V P G P+
Sbjct: 116 TIRRCDIFQAVKNSGTYDFLIDRVPFGPHCVTHQGVQPPA 155
>At5g51590 unknown protein
Length = 419
Score = 41.2 bits (95), Expect = 6e-04
Identities = 19/26 (73%), Positives = 20/26 (76%)
Query: 129 MPELSHTGSTGRGRGRGRGRGRGRGR 154
+P S GS RGRGRGRGRGRGRGR
Sbjct: 106 VPLTSEFGSRKRGRGRGRGRGRGRGR 131
Score = 28.9 bits (63), Expect = 3.3
Identities = 33/117 (28%), Positives = 37/117 (31%), Gaps = 7/117 (5%)
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQ 178
D V MP S T R RGRGRGRGR R EP
Sbjct: 90 DGSLAVTLSPMPISSSVPLTSEFGSRKRGRGRGRGRGRGRGRGQGQGSREP-----NNNN 144
Query: 179 HDTNTDMTIHDSSETKELPKENAAVPAE--SAEFHNLDLNANTNENEDKKASTTAKQ 233
+D N P PAE S F L N E+ K T ++Q
Sbjct: 145 NDNNWLKNPQMFEFNNNTPTSGGGGPAEIVSPSFTPHVLTVNAGEDVTMKIMTFSQQ 201
>At3g07590 putative small nuclear ribonucleoprotein (Sm-D1)
Length = 114
Score = 38.9 bits (89), Expect = 0.003
Identities = 16/18 (88%), Positives = 17/18 (93%)
Query: 137 STGRGRGRGRGRGRGRGR 154
+ GRGRGRGRGRGRGRGR
Sbjct: 97 AVGRGRGRGRGRGRGRGR 114
Score = 35.4 bits (80), Expect = 0.035
Identities = 16/25 (64%), Positives = 16/25 (64%)
Query: 128 KMPELSHTGSTGRGRGRGRGRGRGR 152
K P GRGRGRGRGRGRGR
Sbjct: 90 KKPVAGKAVGRGRGRGRGRGRGRGR 114
Score = 32.0 bits (71), Expect = 0.39
Identities = 13/16 (81%), Positives = 14/16 (87%)
Query: 139 GRGRGRGRGRGRGRGR 154
G+ GRGRGRGRGRGR
Sbjct: 95 GKAVGRGRGRGRGRGR 110
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 38.5 bits (88), Expect = 0.004
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Query: 117 CNDSDEEVKRGKM----PELSHTGSTGRGRGRGRGRGRGRG 153
C++ D ++GK+ E S+ GRGRGRGR GRGRG
Sbjct: 161 CDEDDSPEEQGKLMYANSESSYDTRGGRGRGRGRSSGRGRG 201
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,760,059
Number of Sequences: 26719
Number of extensions: 313676
Number of successful extensions: 2275
Number of sequences better than 10.0: 154
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 71
Number of HSP's that attempted gapping in prelim test: 1535
Number of HSP's gapped (non-prelim): 414
length of query: 283
length of database: 11,318,596
effective HSP length: 98
effective length of query: 185
effective length of database: 8,700,134
effective search space: 1609524790
effective search space used: 1609524790
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0005.4


