

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
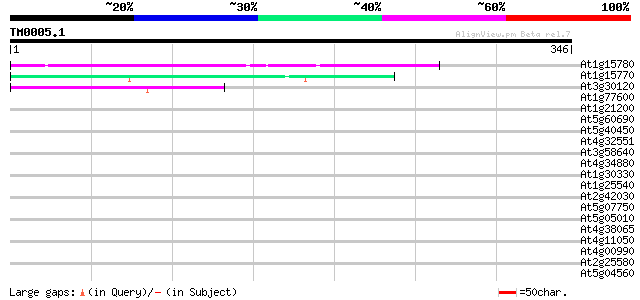
Score E
Sequences producing significant alignments: (bits) Value
At1g15780 unknown protein 52 4e-07
At1g15770 hypothetical protein 52 4e-07
At3g30120 hypothetical protein 42 4e-04
At1g77600 hypothetical protein 35 0.046
At1g21200 unknown protein 33 0.23
At5g60690 REVOLUTA or interfascicular fiberless 1 32 0.51
At5g40450 unknown protein 32 0.51
At4g32551 Leunig protein 32 0.67
At3g58640 unknown protein 31 0.87
At4g34880 amidase - like protein 30 1.5
At1g30330 putative protein 30 1.9
At1g25540 hypothetical protein 30 1.9
At2g42030 putative RING zinc finger protein 29 3.3
At5g07750 putative protein 29 4.3
At5g05010 coatomer delta subunit (delta-coat protein) (delta-COP) 29 4.3
At4g38065 hypothetical protein 29 4.3
At4g11050 glucanase like protein 29 4.3
At4g00990 unknown protein 29 4.3
At2g25580 putative selenium-binding protein 29 4.3
At5g04560 putative protein 28 5.6
>At1g15780 unknown protein
Length = 1335
Score = 52.4 bits (124), Expect = 4e-07
Identities = 59/268 (22%), Positives = 119/268 (44%), Gaps = 9/268 (3%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+T+ LN IYQ++ +KL+Q +S P + ++ +EK R K + E + + KS I
Sbjct: 604 MKETYLPDLNEIYQRVAAKLQQ-DSMPQQQRSDQLEKLRQFKTMLERMIQFLSVSKSNIM 662
Query: 61 PEISERVDRVE-TLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTS-Q 118
P + ++V E +I + H+ Q Q + PQ Q+ QT+ Q
Sbjct: 663 PALKDKVAYYEKQIIGFLNMHRPRKPVQQGQLPQSQMQPMQQPQSQTVQDQSHDNQTNPQ 722
Query: 119 RNNMCLQ-EQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSS 177
+M +Q A+Q + N LS + PG + +Q + + AS + + +
Sbjct: 723 MQSMSMQGAGPRAQQSSMTNMQSNVLSSR--PGVSAPQQNI-PSSIPASSLESGQGNTLN 779
Query: 178 SALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
+ G++ + + L++ ++A L + ++++ L +S+ + + + D
Sbjct: 780 NGQQVAMGSMQQ--NTSQLVNNSSASAQSGLSTLQSNVNQPQLSSSLLQHQHLKQQQDQQ 837
Query: 238 PAAKFIDEPLEMAGQQNQPSRIPQGRKM 265
K + +M QQ Q + Q +++
Sbjct: 838 MQLKQQFQQRQMQQQQLQARQQQQQQQL 865
Score = 35.0 bits (79), Expect = 0.060
Identities = 57/257 (22%), Positives = 98/257 (37%), Gaps = 38/257 (14%)
Query: 96 SKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSI---------- 145
S Q+L + HL+ Q Q+N + + Q NS V S
Sbjct: 919 SSPQLLQGASPQMSQHLSPQVDQKNTV--NKMGTPLQPANSPFVVPSPSSTPLAPSPMQV 976
Query: 146 -KEGPGKA------VQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLN-EVSHKPALI 197
E PG + + RQ ++ G S+S L++ + + + + +
Sbjct: 977 DSEKPGSSSLSMGNIARQQATGMQGVVQSLAIGTPGISASPLLQEFTSPDGNILNSSTIT 1036
Query: 198 SEKPSAA---MQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQ-- 252
S KPSA ++RLI S+S +AL +++ I VV + D I + + G+
Sbjct: 1037 SGKPSATELPIERLIRAVKSISPQALSSAVSDIGSVVSMVDRIAGSAPGNGSRASVGEDL 1096
Query: 253 --------QNQPSRIPQG----RKMARNINAMAFDTSRSCARTHESFNQLTDVEEPDLDL 300
Q + +G +KM R+ AM + +++ Q E DL+
Sbjct: 1097 VAMTKCRLQARNFMTQEGMMATKKMKRHTTAMPLSVASLGGSVGDNYKQFAGSETSDLES 1156
Query: 301 VA-SKVKRSRIEVIYLL 316
A S K++R E + L
Sbjct: 1157 TATSDGKKARTETEHAL 1173
Score = 27.7 bits (60), Expect = 9.6
Identities = 45/229 (19%), Positives = 84/229 (36%), Gaps = 18/229 (7%)
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQ 138
Q Q L Q + ++QQ++ Q Q H +Q N LQ+ Q +Q NS+
Sbjct: 407 QQQPLQQPQQQQKQQPPAQQQLMSQQNSLQATHQNPLGTQSNVAGLQQPQ--QQMLNSQV 464
Query: 139 GVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALIS 198
G + L + + + T+ Q T +G G SS + ++ P L S
Sbjct: 465 GNSSLQNNQHSVHMLSQPTVGLQRTHQAG-----HGLYSSQGQQSQNQPSQQQMMPQLQS 519
Query: 199 EKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSR 258
+Q+ + + L AS G++ +P +D+ ++ Q
Sbjct: 520 HHQQLGLQQQPNLLQQDVQQRLQAS-GQV-----TGSLLPPQNVVDQQRQLYQSQRTLPE 573
Query: 259 IPQGR--KMARNINAMAFDTSRSCARTHESFNQLTDVEEPDLDLVASKV 305
+P A+ +A D ++ + + PDL+ + +V
Sbjct: 574 MPSSSLDSTAQTESANGGDWQE---EVYQKIKSMKETYLPDLNEIYQRV 619
>At1g15770 hypothetical protein
Length = 332
Score = 52.4 bits (124), Expect = 4e-07
Identities = 57/247 (23%), Positives = 98/247 (39%), Gaps = 12/247 (4%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
M++ + + +YQ++ KL ES P + ++ + +KF+ K E + + LRK I
Sbjct: 20 MRERYLPHVTDVYQRLADKLNHEESLPQQQRSKHFDKFQSMKTNIEQLIQVLSLRKRNIM 79
Query: 61 PEISE-RVDRVET-----LINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAK 114
P + + R D E LI ++ HQ + Q L Q L L
Sbjct: 80 PILKDCRWDYYEKRIIYFLITLRTGWEDNQIHQRDLNEIYQRVAAKLQQSLRKTVQKLQL 139
Query: 115 QTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMG 174
S+ M Q + + +Q Q+ G + Q + ++ + T G
Sbjct: 140 TKSEIQPMQQPLSQTVQDQSHDDQTTLQMQSMSMQGAGSRVQQIRQGVLQSLEIGT--PG 197
Query: 175 FSSSALI----KGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREV 230
S+S L+ GN+ S ++RLI S+S +AL +++ IR V
Sbjct: 198 ISASPLLPELTSPDGNIINPLTSTCGKSSATELPIERLIRAMKSISPQALSSAVCDIRSV 257
Query: 231 VHLNDAI 237
V + D I
Sbjct: 258 VSMVDRI 264
>At3g30120 hypothetical protein
Length = 750
Score = 42.4 bits (98), Expect = 4e-04
Identities = 32/138 (23%), Positives = 62/138 (44%), Gaps = 6/138 (4%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEP-KTFNVEKFRGHKRLYEDIFAMFRLRKSQI 59
+K+ HF L+ ++++I KLRQ ES P +P + ++EK + K E + +++S +
Sbjct: 149 LKEMHFPVLSLMHKRIAEKLRQTESLPPQPMQAQSIEKLKAGKLSMEHLMFFLSVQRSNV 208
Query: 60 TPEISERVDRVETLINSIIQHQNL-----SSHQGKHSADVQSKQQVLPQVLGTQNDHLAK 114
+ + ++ E I + Q + QG+ + Q PQV +Q+ +
Sbjct: 209 SEKHRDKFSLYEHHILKFTKSQTMVQRSTQQQQGQFPPSQTAMQSQSPQVHVSQSLDKEQ 268
Query: 115 QTSQRNNMCLQEQQVAKQ 132
S+ C E + Q
Sbjct: 269 MRSRLMPSCQNEASSSLQ 286
Score = 35.0 bits (79), Expect = 0.060
Identities = 41/170 (24%), Positives = 73/170 (42%), Gaps = 17/170 (10%)
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQ--------EQQV- 129
QHQ S+Q DV+ +Q++L +++ + + KQ S+ ++ +Q +QQ+
Sbjct: 350 QHQMQRSNQTNEMNDVRIRQRLLEKLVSSSQLQVPKQVSKVSSPQIQNHSSPQLVDQQIL 409
Query: 130 ---AKQYGNSEQGVNQLSIKEGPGKAV-QRQTLCAQATEASGVSTNDMGFSSSALIKGSG 185
+ G + + P + + + SGV N SSS L
Sbjct: 410 PATVNKTGTPLKSSGSPFVAPAPSPVPGDSEMPISVESPVSGVEINSTLDSSSKLGTQET 469
Query: 186 NLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLND 235
L V P I+E+P + RLI F + S ++L S+ +I V+ + D
Sbjct: 470 PLLSVP-PPEPITERP---IDRLIKAFQAASPKSLAESVSEISSVISMVD 515
>At1g77600 hypothetical protein
Length = 1303
Score = 35.4 bits (80), Expect = 0.046
Identities = 30/119 (25%), Positives = 56/119 (46%), Gaps = 13/119 (10%)
Query: 89 KHSADVQSKQQVLPQVLGTQNDHL---AKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSI 145
K S V S+ +VLP +LG Q + + + SQ N C ++ + + N LS+
Sbjct: 1031 KDSLAVGSEDKVLPPLLGNQIETSITGSTEASQNNTRCSRK----RTHLGEHISCNSLSL 1086
Query: 146 K----EGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEK 200
+ E P K ++R T CA+ + + VS S ++ +++ +H A+I ++
Sbjct: 1087 RTVESEIPIKKLERHTTCAKESVKASVSNKITSSKHSGVVSALKDIS--NHGEAIIGQR 1143
>At1g21200 unknown protein
Length = 443
Score = 33.1 bits (74), Expect = 0.23
Identities = 23/81 (28%), Positives = 37/81 (45%), Gaps = 7/81 (8%)
Query: 78 IQHQNLSSHQGKHSADVQSKQQVLP--QVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGN 135
+ HQ+ + Q +H+ + + + LP V G DH Q NM + EQQ A++ N
Sbjct: 29 VHHQDSMNQQHRHNPNSRPLHEGLPFTMVTGQTCDH-----HQNQNMSMSEQQKAEREKN 83
Query: 136 SEQGVNQLSIKEGPGKAVQRQ 156
S ++ S E G V +
Sbjct: 84 SVSDDDEPSFTEEGGDGVHNE 104
>At5g60690 REVOLUTA or interfascicular fiberless 1
Length = 842
Score = 32.0 bits (71), Expect = 0.51
Identities = 25/102 (24%), Positives = 45/102 (43%), Gaps = 3/102 (2%)
Query: 20 LRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQITPEISERVDRVETLINSIIQ 79
+ +H S + + E+ +R+Y + LR+ Q+ E S + I Q
Sbjct: 16 MNRHLDSSGKYVRYTAEQVEALERVYAECPKPSSLRRQQLIRECSILANIEPKQIKVWFQ 75
Query: 80 HQNLSSHQGKHSADVQS---KQQVLPQVLGTQNDHLAKQTSQ 118
++ Q K ++ +QS K + ++L +ND L KQ SQ
Sbjct: 76 NRRCRDKQRKEASRLQSVNRKLSAMNKLLMEENDRLQKQVSQ 117
>At5g40450 unknown protein
Length = 2910
Score = 32.0 bits (71), Expect = 0.51
Identities = 41/225 (18%), Positives = 80/225 (35%), Gaps = 21/225 (9%)
Query: 93 DVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKA 152
D + P +L + +++ N + E + + E+ VN + K K
Sbjct: 1981 DAHVETPTAPIILEENDSETLIAEAKKGNEEINETERTVALDHEEEFVNHEAPKLEETKD 2040
Query: 153 VQRQTL--CAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIG 210
+ Q + A+ATE + T +G S + P+L+S+K +++
Sbjct: 2041 EKSQEIPETAKATETTIDQTLPIGTS------------QADQTPSLVSDKDDQTPKQVEE 2088
Query: 211 VFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNIN 270
+ + E E + + +P FI+ P+ M PQ A N
Sbjct: 2089 ILEEETKETHKVQA----EDIFSTETVPKESFIEAPVSMLASGEDEPVTPQEGDYAANTQ 2144
Query: 271 A---MAFDTSRSCARTHESFNQLTDVEEPDLDLVASKVKRSRIEV 312
++ +T T +Q E+ D + K++ +EV
Sbjct: 2145 EERHVSAETEEKVGETKPKESQAEGAEKSDDQVEDESTKKTDVEV 2189
>At4g32551 Leunig protein
Length = 931
Score = 31.6 bits (70), Expect = 0.67
Identities = 33/141 (23%), Positives = 53/141 (37%), Gaps = 13/141 (9%)
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQ 138
QH +S Q + Q +QQ+ Q L Q +Q Q+ + Q+QQ +Q +Q
Sbjct: 95 QHPQVSQQQQQ-----QQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQQQQQQQQQQQ 149
Query: 139 GVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVSHKPALIS 198
Q + P Q+Q Q + S L GS N L+
Sbjct: 150 QQQQQQHQNQPPSQQQQQQSTPQHQQQPTPQQQPQRRDGSHLANGSAN--------GLVG 201
Query: 199 EKPSAAMQRLIGVFTSMSSEA 219
M++ G +S++S+A
Sbjct: 202 NNSEPVMRQNPGSGSSLASKA 222
>At3g58640 unknown protein
Length = 809
Score = 31.2 bits (69), Expect = 0.87
Identities = 19/65 (29%), Positives = 27/65 (41%), Gaps = 6/65 (9%)
Query: 53 RLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHL 112
R R ITPEI + + R +N ++ LS QG + S T++ HL
Sbjct: 413 RRRSISITPEIGDDIVRAVRAMNEALKQNRLSKEQGDDDSSPNSPND------RTESSHL 466
Query: 113 AKQTS 117
K S
Sbjct: 467 QKNVS 471
>At4g34880 amidase - like protein
Length = 466
Score = 30.4 bits (67), Expect = 1.5
Identities = 23/91 (25%), Positives = 41/91 (44%), Gaps = 15/91 (16%)
Query: 177 SSALIKGSGNLNEVSH--------------KPALISEKPSAAMQRLIGVFT-SMSSEALG 221
S A+I G +L+E +H + +++ KPS + GV S+ +++G
Sbjct: 153 SGAVILGKASLSEWAHFRSFSIPDGWSAPSQNSVVGIKPSVGLTSRAGVVPISLRQDSIG 212
Query: 222 ASIGKIREVVHLNDAIPAAKFIDEPLEMAGQ 252
+ + VHL DAI +DE + A +
Sbjct: 213 PICRTVSDAVHLLDAIVGYDPLDEATKTASE 243
>At1g30330 putative protein
Length = 933
Score = 30.0 bits (66), Expect = 1.9
Identities = 29/114 (25%), Positives = 47/114 (40%), Gaps = 9/114 (7%)
Query: 51 MFRLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSADVQ----SKQQVLPQVLG 106
M + + SQ ++S++ + + L Q Q LS Q + + Q S+QQ LG
Sbjct: 481 MLQQQLSQQQQQLSQQQQQQQQLSQQ--QQQQLSQQQQQQLSQQQQQQLSQQQQQQAYLG 538
Query: 107 TQNDHLAK---QTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQT 157
H + Q+ N++ Q+QQV + S +S G A Q T
Sbjct: 539 VPETHQPQSQAQSQSNNHLSQQQQQVVDNHNPSASSAAVVSAMSQFGSASQPNT 592
Score = 27.7 bits (60), Expect = 9.6
Identities = 26/120 (21%), Positives = 47/120 (38%), Gaps = 6/120 (5%)
Query: 72 TLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAK 131
+L+ + Q LS Q + S Q +QQ+ Q + +Q SQ+ L +QQ +
Sbjct: 475 SLVQPQMLQQQLSQQQQQLSQQQQQQQQLSQQQQQQLSQQQQQQLSQQQQQQLSQQQQQQ 534
Query: 132 QYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLNEVS 191
Y GV + + ++ L Q + V ++ SS+A++ S
Sbjct: 535 AY----LGVPETHQPQSQAQSQSNNHLSQQQQQV--VDNHNPSASSAAVVSAMSQFGSAS 588
>At1g25540 hypothetical protein
Length = 809
Score = 30.0 bits (66), Expect = 1.9
Identities = 21/84 (25%), Positives = 36/84 (42%), Gaps = 2/84 (2%)
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P+I + + + ++ Q Q Q + +Q QQ +PQ+ Q H +Q Q
Sbjct: 649 PQIPNQQQQQQQQLHQQQQQQQQIQQQQQQQQHLQ--QQQMPQLQQQQQQHQQQQQQQHQ 706
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLS 144
LQ Q +Q +Q +QL+
Sbjct: 707 LSQLQHHQQQQQQQQQQQQQHQLT 730
Score = 28.5 bits (62), Expect = 5.6
Identities = 23/86 (26%), Positives = 35/86 (39%), Gaps = 4/86 (4%)
Query: 79 QHQ--NLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNS 136
QHQ L HQ + Q +QQ Q+ Q+ H +Q + N Q+ Q
Sbjct: 704 QHQLSQLQHHQQQQQQQQQQQQQ--HQLTQLQHHHQQQQQASPLNQMQQQTSPLNQMQQQ 761
Query: 137 EQGVNQLSIKEGPGKAVQRQTLCAQA 162
+NQ+ ++ P + V AQA
Sbjct: 762 TSPLNQMQQQQQPQQMVMGGQAFAQA 787
>At2g42030 putative RING zinc finger protein
Length = 425
Score = 29.3 bits (64), Expect = 3.3
Identities = 33/141 (23%), Positives = 61/141 (42%), Gaps = 14/141 (9%)
Query: 164 EASGVSTNDMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGAS 223
E S ND+ + + +S AL +++ A M + FT S+E L +
Sbjct: 281 EPSSDEINDIDLILNNAPENEEENENLSSSRALATQRRWAQMYGRVSSFTLSSAERLADT 340
Query: 224 ------IGKIREVVHLNDAIPAA-KFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDT 276
+G+ +E N + P + D + G N S++ + A IN+M +
Sbjct: 341 YLITHALGRNQEQ---NSSPPVGVEDRDSFSSIVGVINSESQV----ETAAEINSMLTVS 393
Query: 277 SRSCARTHESFNQLTDVEEPD 297
+ S R HE+ ++++DV+ D
Sbjct: 394 TSSSVRRHENSSRVSDVDSAD 414
>At5g07750 putative protein
Length = 1289
Score = 28.9 bits (63), Expect = 4.3
Identities = 22/72 (30%), Positives = 38/72 (52%), Gaps = 4/72 (5%)
Query: 140 VNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKG-SGNLNEVSHKPALIS 198
V + I +G + +R+T+ A+ ++S V T G S ++ S N +KP IS
Sbjct: 426 VKDIGIDDGDEQR-KRRTVEAKENDSSTVQTQSKGDEESNDLESMSQKTNTSLNKP--IS 482
Query: 199 EKPSAAMQRLIG 210
EKP A +++ +G
Sbjct: 483 EKPQATLRKQVG 494
>At5g05010 coatomer delta subunit (delta-coat protein) (delta-COP)
Length = 527
Score = 28.9 bits (63), Expect = 4.3
Identities = 20/78 (25%), Positives = 34/78 (42%), Gaps = 1/78 (1%)
Query: 74 INSIIQHQNLSSHQGK-HSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQ 132
+ + Q+ + SH+ K H +QSK V+ + + + K ++N +
Sbjct: 127 VAQVKQYCEMESHEEKLHKLVMQSKINDTKDVMKRKANEIDKSKIEKNKPGGFSSMGSMG 186
Query: 133 YGNSEQGVNQLSIKEGPG 150
G E G N+LSI G G
Sbjct: 187 SGRLESGFNELSISSGGG 204
>At4g38065 hypothetical protein
Length = 1050
Score = 28.9 bits (63), Expect = 4.3
Identities = 24/115 (20%), Positives = 51/115 (43%), Gaps = 14/115 (12%)
Query: 19 KLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQITPEISERVDRVETLINSII 78
K+ + +S E + E+F+ + YE + +F+ K + E S+ +D + +L
Sbjct: 174 KMEEEKSQVEEKLKWKKEQFKHLEEAYEKLKNLFKDSKKEWEEEKSKLLDEIYSL----- 228
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQY 133
Q + S D+Q K Q+ N L ++ ++R ++ +Q + +Y
Sbjct: 229 --QTKLDSVTRISEDLQKKLQMC-------NGALTQEETRRKHLEIQVSEFKAKY 274
>At4g11050 glucanase like protein
Length = 626
Score = 28.9 bits (63), Expect = 4.3
Identities = 27/116 (23%), Positives = 53/116 (45%), Gaps = 10/116 (8%)
Query: 127 QQVAKQYGNSEQGVNQLSIKEGPGKAVQRQT-----LCAQATEASGVSTNDMGFSSSALI 181
QQ A+Q+ S G + +IK+ PG + RQ+ A+ + V ++ + +S L+
Sbjct: 308 QQKAEQFMCSLLGKSTKNIKKTPGGLIFRQSWNNMQFVTSASFLATVYSDYLSYSKRDLL 367
Query: 182 KGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
GN++ P+ + E + + ++G +S +G R+V H +I
Sbjct: 368 CSQGNIS-----PSQLLEFSKSQVDYILGDNPRATSYMVGYGENYPRQVHHRGSSI 418
>At4g00990 unknown protein
Length = 454
Score = 28.9 bits (63), Expect = 4.3
Identities = 23/80 (28%), Positives = 37/80 (45%), Gaps = 15/80 (18%)
Query: 78 IQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAK-----Q 132
+++QN+ HQ K++ + KQQ QV K+ S+ N ++E +K +
Sbjct: 181 VKYQNIKVHQKKYAEAMLQKQQYSGQV---------KEASELENKSMKEVDESKKDLKDK 231
Query: 133 YGNSEQGVNQLSIKEGPGKA 152
N EQ N S G G+A
Sbjct: 232 AANEEQS-NNSSRPSGSGEA 250
>At2g25580 putative selenium-binding protein
Length = 567
Score = 28.9 bits (63), Expect = 4.3
Identities = 19/76 (25%), Positives = 33/76 (43%), Gaps = 1/76 (1%)
Query: 81 QNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGV 140
QN + G+ S + ++ Q L G +N+ KQ+ +N +Q Q YG +
Sbjct: 12 QNRTGSSGEVSESIHTQSQSLGSNQG-RNEQSWKQSPSLSNSQVQSQYQGNWYGTNSDYQ 70
Query: 141 NQLSIKEGPGKAVQRQ 156
N + GK + R+
Sbjct: 71 NGVGSSWHSGKTIDRE 86
>At5g04560 putative protein
Length = 1017
Score = 28.5 bits (62), Expect = 5.6
Identities = 16/63 (25%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Query: 218 EALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTS 277
+A+G + G + V D +++PLE++ NQP ++ G K+AR+ +
Sbjct: 399 DAIGGTNGSFLDSVSQIDKTNGLGAMNQPLEVS-MGNQPDKLSTGAKLARDQQPDLLTRN 457
Query: 278 RSC 280
+ C
Sbjct: 458 QQC 460
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,889,924
Number of Sequences: 26719
Number of extensions: 261535
Number of successful extensions: 789
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 754
Number of HSP's gapped (non-prelim): 47
length of query: 346
length of database: 11,318,596
effective HSP length: 100
effective length of query: 246
effective length of database: 8,646,696
effective search space: 2127087216
effective search space used: 2127087216
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0005.1


