

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0004a.3
(148 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
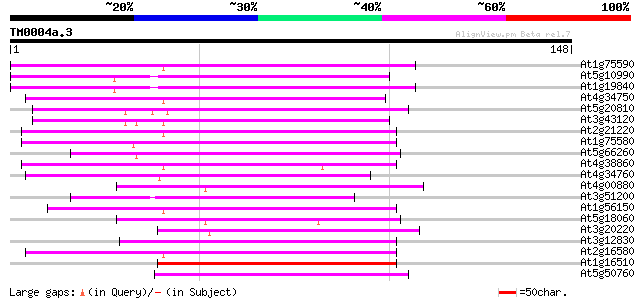
Score E
Sequences producing significant alignments: (bits) Value
At1g75590 unknown protein 79 9e-16
At5g10990 unknown protein 77 3e-15
At1g19840 hypothetical protein 77 4e-15
At4g34750 putative protein 69 7e-13
At5g20810 unknown protein 69 1e-12
At3g43120 unknown protein 69 1e-12
At2g21220 auxin-regulated like protein 67 5e-12
At1g75580 auxin induced protein, putative 66 8e-12
At5g66260 auxin-induced protein-like 64 2e-11
At4g38860 putative auxin-induced protein 64 2e-11
At4g34760 putative auxin-regulated protein 64 2e-11
At4g00880 62 1e-10
At3g51200 putative protein 62 1e-10
At1g56150 unknown protein 61 2e-10
At5g18060 auxin-induced protein-like 60 3e-10
At3g20220 unknown protein 60 3e-10
At3g12830 unknown protein 59 7e-10
At2g16580 putative auxin-induced protein 59 1e-09
At1g16510 auxin-induced like protein 59 1e-09
At5g50760 putative protein 58 2e-09
>At1g75590 unknown protein
Length = 154
Score = 79.0 bits (193), Expect = 9e-16
Identities = 42/110 (38%), Positives = 63/110 (57%), Gaps = 3/110 (2%)
Query: 1 MAGAMKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAI 57
MAG + K KIR IVRL+Q++ RW+ + S P PSG + VYVG +RF +
Sbjct: 1 MAGGLGKCSKIRHIVRLRQMLRRWRDQARMSSSFSRCVPSDVPSGHVAVYVGSSCRRFVV 60
Query: 58 PARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNEN 107
A +LN PV LL + EEEFG G L++PC+ F ++ + ++++
Sbjct: 61 RATYLNHPVLRNLLVQAEEEFGFVNQGPLVIPCEESVFEESIRFISRSDS 110
>At5g10990 unknown protein
Length = 148
Score = 77.4 bits (189), Expect = 3e-15
Identities = 42/104 (40%), Positives = 61/104 (58%), Gaps = 6/104 (5%)
Query: 1 MAGAMKKVDKIRQIVRLKQVMMRWKQ----ISLRRHDSSDPRPPSGCLYVYVGHERQRFA 56
MAG + K KIR IV+L+Q++ +W+ S+RR SD PSG + VYVG +RF
Sbjct: 1 MAGGIGKCSKIRHIVKLRQMLRQWRNKARMSSVRRSVPSDV--PSGHVAVYVGRSCRRFV 58
Query: 57 IPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVK 100
+ A +LN P+ LL + EEEFG G L++PC+ F ++
Sbjct: 59 VLATYLNHPILMNLLVKAEEEFGFANQGPLVIPCEESVFEESIR 102
>At1g19840 hypothetical protein
Length = 153
Score = 76.6 bits (187), Expect = 4e-15
Identities = 42/111 (37%), Positives = 66/111 (58%), Gaps = 6/111 (5%)
Query: 1 MAGAMKKVDKIRQIVRLKQVMMRWKQ----ISLRRHDSSDPRPPSGCLYVYVGHERQRFA 56
MAG++ K KIR IVRL+Q++ RW+ S+ R SD PSG + V VG +RF
Sbjct: 1 MAGSLVKCSKIRHIVRLRQMLRRWRNKARLSSVSRCVPSDV--PSGHVAVCVGSGCRRFV 58
Query: 57 IPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRLHKNEN 107
+ A +LN P+ + LL + EEEFG G L++PC+ F ++ + ++++
Sbjct: 59 VRASYLNHPIISNLLVQAEEEFGFANQGPLVIPCEESVFEEAIRFISRSDS 109
>At4g34750 putative protein
Length = 150
Score = 69.3 bits (168), Expect = 7e-13
Identities = 36/97 (37%), Positives = 57/97 (58%), Gaps = 2/97 (2%)
Query: 5 MKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP--PSGCLYVYVGHERQRFAIPARFL 62
M K +KI +VR++Q++ +W++ + ++DP P G + V VG R+R+ + A+ L
Sbjct: 1 MGKNNKIGSVVRIRQMLKQWQKKAHIGSSNNDPVSDVPPGHVAVSVGENRRRYVVRAKHL 60
Query: 63 NLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
N P+F LL E EEE+G G L +PCD F I+
Sbjct: 61 NHPIFRRLLAEAEEEYGFANVGPLAIPCDESLFEDII 97
>At5g20810 unknown protein
Length = 165
Score = 68.6 bits (166), Expect = 1e-12
Identities = 44/138 (31%), Positives = 67/138 (47%), Gaps = 39/138 (28%)
Query: 7 KVDKIRQIVRLKQVMMRWKQISL------------------------------RRHDSSD 36
K+ IRQIVRLK+++ +W+ +++ + DS +
Sbjct: 8 KLTGIRQIVRLKEILQKWQTVTIGPKSEVPPLAAGKQAVAMISPAINKRLLDVKNGDSDE 67
Query: 37 -----PRPP----SGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLI 87
P PP G L VYVG E +RF IP +L+ +F LL++ EEEFG SG L
Sbjct: 68 ETCQSPEPPHDVPKGNLAVYVGPELRRFIIPTSYLSHSLFKVLLEKAEEEFGFDQSGALT 127
Query: 88 LPCDVCFFTHIVKRLHKN 105
+PC+V F +++K + N
Sbjct: 128 IPCEVETFKYLLKCMENN 145
>At3g43120 unknown protein
Length = 160
Score = 68.6 bits (166), Expect = 1e-12
Identities = 43/133 (32%), Positives = 66/133 (49%), Gaps = 39/133 (29%)
Query: 7 KVDKIRQIVRLKQVMMRWKQISL-----------RRH----------------------- 32
K+ I+QIVRLK+++ +W+ +++ R+H
Sbjct: 8 KLTGIKQIVRLKEILQKWQTVTIGSKSDDGELGARKHTAIISPVINKRLLDLKTCDSDEE 67
Query: 33 ---DSSDPRP--PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLI 87
S +P P P G L VYVG E +RF IP FL+ +F LL++ EEE+G SG L
Sbjct: 68 TTCQSPEPPPDVPKGYLAVYVGPELRRFIIPTNFLSHSLFKVLLEKAEEEYGFDHSGALT 127
Query: 88 LPCDVCFFTHIVK 100
+PC+V F +++K
Sbjct: 128 IPCEVETFKYLLK 140
>At2g21220 auxin-regulated like protein
Length = 104
Score = 66.6 bits (161), Expect = 5e-12
Identities = 35/102 (34%), Positives = 51/102 (49%), Gaps = 3/102 (2%)
Query: 4 AMKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPAR 60
A+K+ K+ Q LKQ++ R ++ + D P P G VYVG +R R+ +P
Sbjct: 2 AVKRSSKLTQTAMLKQILKRCSSLAKNQCYDEDGLPVDVPKGHFPVYVGEKRSRYIVPIS 61
Query: 61 FLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRL 102
FL P F LL + EEEFG GL +PC+ F + +
Sbjct: 62 FLTHPKFKSLLQQAEEEFGFNHDMGLTIPCEEVVFRSLTSMI 103
>At1g75580 auxin induced protein, putative
Length = 108
Score = 65.9 bits (159), Expect = 8e-12
Identities = 36/106 (33%), Positives = 51/106 (47%), Gaps = 7/106 (6%)
Query: 4 AMKKVDKIRQIVRLKQVMMRWKQISLRR-------HDSSDPRPPSGCLYVYVGHERQRFA 56
AMKK +K+ Q +KQ++ R + ++ + S P G VYVG R R+
Sbjct: 2 AMKKANKLTQTAMIKQILKRCSSLGKKQSNVYGEDENGSPLNVPKGHFVVYVGENRVRYV 61
Query: 57 IPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRL 102
+P FL P F LL + EEEFG GL +PC+ F + L
Sbjct: 62 VPISFLTRPEFQLLLQQAEEEFGFDHDMGLTIPCEEVVFRSLTSML 107
>At5g66260 auxin-induced protein-like
Length = 99
Score = 64.3 bits (155), Expect = 2e-11
Identities = 34/88 (38%), Positives = 47/88 (52%), Gaps = 1/88 (1%)
Query: 17 LKQVMMRWKQISLRRH-DSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETE 75
LKQ++ R + + D + P G VYVGH R R IP FL P+F LL ++E
Sbjct: 11 LKQMLKRCSSLGKKSSVDVNFNGVPKGHFVVYVGHSRSRHVIPISFLTHPIFQMLLQQSE 70
Query: 76 EEFGLRGSGGLILPCDVCFFTHIVKRLH 103
EEFG GL +PCD FF ++ ++
Sbjct: 71 EEFGFFQDNGLTIPCDEHFFRALISSIN 98
>At4g38860 putative auxin-induced protein
Length = 105
Score = 64.3 bits (155), Expect = 2e-11
Identities = 36/103 (34%), Positives = 53/103 (50%), Gaps = 4/103 (3%)
Query: 4 AMKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP---PSGCLYVYVGHERQRFAIPAR 60
A+K+ K+ Q LKQ++ R + ++ + P P G VYVG +R R+ +P
Sbjct: 2 AVKRSSKLTQTAMLKQILKRCSSLGKKQCYDEEGLPLDVPKGHFPVYVGEKRTRYIVPIS 61
Query: 61 FLNLPVFAGLLDETEEEFGLR-GSGGLILPCDVCFFTHIVKRL 102
FL P F LL + EEEFG R GGL +PC+ F + +
Sbjct: 62 FLTHPEFLILLQQAEEEFGFRHDMGGLTIPCEEVVFLSLTSMI 104
>At4g34760 putative auxin-regulated protein
Length = 107
Score = 64.3 bits (155), Expect = 2e-11
Identities = 36/96 (37%), Positives = 44/96 (45%), Gaps = 5/96 (5%)
Query: 5 MKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPR-----PPSGCLYVYVGHERQRFAIPA 59
MKK K+ Q LKQ++ R + + D P G VYVG R R+ +P
Sbjct: 4 MKKTSKLTQTAMLKQILKRCSSLGKKNGGGYDEDCLPLDVPKGHFPVYVGENRSRYIVPI 63
Query: 60 RFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFF 95
FL P F LL EEEFG GL +PCD F
Sbjct: 64 SFLTHPEFQSLLQRAEEEFGFDHDMGLTIPCDELVF 99
>At4g00880
Length = 122
Score = 61.6 bits (148), Expect = 1e-10
Identities = 30/83 (36%), Positives = 44/83 (52%), Gaps = 2/83 (2%)
Query: 29 LRRHDSSDPRPPSGCLYVYVGH--ERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGL 86
L H+ + P GCL V VG E++RF IP + N P+F LL E EEEFG G +
Sbjct: 18 LHHHEHDHEKVPKGCLAVKVGQGEEQERFVIPVMYFNHPLFGQLLKEAEEEFGFAQKGTI 77
Query: 87 ILPCDVCFFTHIVKRLHKNENKY 109
+PC V F ++ + + ++
Sbjct: 78 TIPCHVEEFRYVQGLIDRENTRF 100
>At3g51200 putative protein
Length = 106
Score = 61.6 bits (148), Expect = 1e-10
Identities = 32/75 (42%), Positives = 43/75 (56%), Gaps = 1/75 (1%)
Query: 17 LKQVMMRWKQISLRRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEE 76
LKQ++M+ +++ + D P G VYVGH R R IP LN P F +L ++EE
Sbjct: 19 LKQMLMKRCSSFVKKSNEEDV-PKKGYFAVYVGHFRDRHVIPITSLNHPTFKMMLQKSEE 77
Query: 77 EFGLRGSGGLILPCD 91
EFG R GL +PCD
Sbjct: 78 EFGFRQESGLTIPCD 92
>At1g56150 unknown protein
Length = 110
Score = 61.2 bits (147), Expect = 2e-10
Identities = 36/101 (35%), Positives = 52/101 (50%), Gaps = 9/101 (8%)
Query: 11 IRQIVRLKQVMMRWKQISLRRHDSSDPRP---------PSGCLYVYVGHERQRFAIPARF 61
++Q++R + Q SL R +S R P G + VYVGHE +RF + A
Sbjct: 1 MKQLIRRLSRVADSTQYSLLRSESQRGRTKKEKHKSWVPEGHVPVYVGHEMERFVVNAEL 60
Query: 62 LNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRL 102
LN PVF LL ++ +E+G G L +PC V F I++ L
Sbjct: 61 LNHPVFVALLKQSAQEYGYEQQGVLRIPCHVLVFERILESL 101
>At5g18060 auxin-induced protein-like
Length = 90
Score = 60.5 bits (145), Expect = 3e-10
Identities = 32/77 (41%), Positives = 45/77 (57%), Gaps = 2/77 (2%)
Query: 29 LRRHDSSDPRPPSGCLYVYVGH-ERQRFAIPARFLNLPVFAGLLDETEEEFGL-RGSGGL 86
L R ++ PP G L VYVG +++R+ +P +LN P F LL ++EEEFG GGL
Sbjct: 14 LSRSAAAVSAPPKGFLAVYVGESQKKRYLVPLSYLNQPSFQALLSKSEEEFGFDHPMGGL 73
Query: 87 ILPCDVCFFTHIVKRLH 103
+PC F ++ RLH
Sbjct: 74 TIPCPEDTFINVTSRLH 90
>At3g20220 unknown protein
Length = 118
Score = 60.5 bits (145), Expect = 3e-10
Identities = 31/70 (44%), Positives = 40/70 (56%), Gaps = 1/70 (1%)
Query: 40 PSGCLYVYVGHE-RQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHI 98
P G L VYVG E RQRF IP ++L P F L+DE +EFG GG+ +PC+ F I
Sbjct: 48 PRGHLAVYVGREERQRFVIPTKYLQYPEFRSLMDEVADEFGYDHEGGIHIPCEESVFEEI 107
Query: 99 VKRLHKNENK 108
+ R + K
Sbjct: 108 LIRYMSCDKK 117
>At3g12830 unknown protein
Length = 132
Score = 59.3 bits (142), Expect = 7e-10
Identities = 30/73 (41%), Positives = 40/73 (54%)
Query: 30 RRHDSSDPRPPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILP 89
RR P G + VYVG E +RF + A LN PVF GLL+ + +E+G G L +P
Sbjct: 41 RRSKKQTSSVPEGHVPVYVGDEMERFVVSAELLNHPVFIGLLNRSAQEYGYEQKGVLQIP 100
Query: 90 CDVCFFTHIVKRL 102
C V F I++ L
Sbjct: 101 CHVLVFERIMESL 113
>At2g16580 putative auxin-induced protein
Length = 108
Score = 58.5 bits (140), Expect = 1e-09
Identities = 34/104 (32%), Positives = 46/104 (43%), Gaps = 6/104 (5%)
Query: 5 MKKVDKIRQIVRLKQVMMRWKQISLRRHDSSDPRP------PSGCLYVYVGHERQRFAIP 58
+KK K+ Q L+Q++ R + + P G VYVGH R R+ +P
Sbjct: 4 LKKSTKLAQTAMLRQILKRCSSLGKKNGGGGYEEVDLPLDVPKGHFPVYVGHNRSRYIVP 63
Query: 59 ARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIVKRL 102
FL F LL EEEFG GL +PCD FF + +
Sbjct: 64 ISFLTNLDFQCLLRRAEEEFGFDHDMGLTIPCDELFFQDLTSMI 107
>At1g16510 auxin-induced like protein
Length = 147
Score = 58.5 bits (140), Expect = 1e-09
Identities = 26/63 (41%), Positives = 39/63 (61%)
Query: 40 PSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHIV 99
P+G + VYVG E +RF + A +N P+F GLL+ + +E+G G L +PC V F +V
Sbjct: 55 PAGHVPVYVGEEMERFVVSAELMNHPIFVGLLNRSAQEYGYAQKGVLHIPCHVIVFERVV 114
Query: 100 KRL 102
+ L
Sbjct: 115 ETL 117
>At5g50760 putative protein
Length = 183
Score = 57.8 bits (138), Expect = 2e-09
Identities = 26/67 (38%), Positives = 38/67 (55%)
Query: 39 PPSGCLYVYVGHERQRFAIPARFLNLPVFAGLLDETEEEFGLRGSGGLILPCDVCFFTHI 98
P G VYVG +QR + + LN P+F LL++ E E+G R G ++LPC+V FF
Sbjct: 55 PSHGFFTVYVGPTKQRIVVKTKLLNHPLFKNLLEDAETEYGYRRDGPIVLPCEVDFFFKA 114
Query: 99 VKRLHKN 105
+ + N
Sbjct: 115 LADMKSN 121
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.141 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,360,894
Number of Sequences: 26719
Number of extensions: 138097
Number of successful extensions: 462
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 353
Number of HSP's gapped (non-prelim): 82
length of query: 148
length of database: 11,318,596
effective HSP length: 90
effective length of query: 58
effective length of database: 8,913,886
effective search space: 517005388
effective search space used: 517005388
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0004a.3


