

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0002.6
(260 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
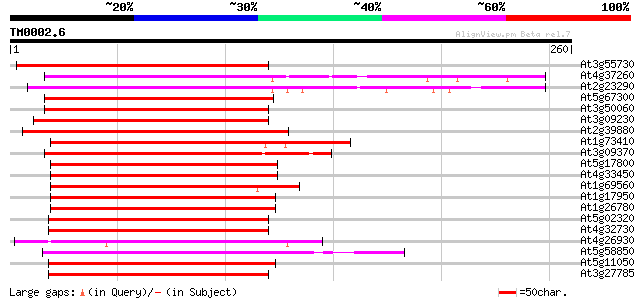
Score E
Sequences producing significant alignments: (bits) Value
At3g55730 MYB transcription factor - like protein 179 1e-45
At4g37260 myb-related protein 177 4e-45
At2g23290 putative MYB family transcription factor 174 3e-44
At5g67300 myb-related protein, 33.3K (pir||S71284) 173 7e-44
At3g50060 R2R3-MYB transcription factor 170 6e-43
At3g09230 unknown protein 164 3e-41
At2g39880 putative MYB family transcription factor 164 6e-41
At1g73410 putative protein 143 1e-34
At3g09370 MYB family like transcription factor 134 6e-32
At5g17800 MYB56 R2R3-MYB factor family member 133 1e-31
At4g33450 putative transcription factor (MYB69) 132 2e-31
At1g69560 myb-related transcription factor, putative 131 3e-31
At1g17950 unknown protein 131 3e-31
At1g26780 putative transcription factor MYB117 (MYB117) 129 2e-30
At5g02320 myb -like protein 127 8e-30
At4g32730 putative myb-protein 126 1e-29
At4g26930 putative myb-related protein 125 2e-29
At5g58850 putative transcription factor MYB119 (MYB119) 124 4e-29
At5g11050 putative transcription factor MYB64 (MYB64) 123 9e-29
At3g27785 putative transcription factor MYB118 (MYB118) 123 9e-29
>At3g55730 MYB transcription factor - like protein
Length = 399
Score = 179 bits (455), Expect = 1e-45
Identities = 81/117 (69%), Positives = 94/117 (80%)
Query: 4 GGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWC 63
GG GGGGG R +VKG WS +ED L KLV + GPRNWS+I+ GIPGRSGKSCRLRWC
Sbjct: 40 GGGGCGGGGGGIRSKVKGPWSTEEDAVLTKLVRKLGPRNWSLIARGIPGRSGKSCRLRWC 99
Query: 64 NQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
NQL P ++ +PF+ ED +II AHAVHGNKWA I++LL GRTDNAIKNHWNSTLRR+
Sbjct: 100 NQLDPCLKRKPFSDEEDRMIISAHAVHGNKWAVIAKLLTGRTDNAIKNHWNSTLRRK 156
>At4g37260 myb-related protein
Length = 320
Score = 177 bits (450), Expect = 4e-45
Identities = 112/269 (41%), Positives = 148/269 (54%), Gaps = 43/269 (15%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
+R+KG WSP+ED+ L +LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP+V+HR F+
Sbjct: 10 ERIKGPWSPEEDDLLQRLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSPEVEHRAFS 69
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR---------------- 120
ED II+AHA GNKWATISRLL GRTDNAIKNHWNSTL+R+
Sbjct: 70 QEEDETIIRAHARFGNKWATISRLLNGRTDNAIKNHWNSTLKRKCSVEGQSCDFGGNGGY 129
Query: 121 ---RGESLPLPLASPSPSPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCVEKD 177
GE PL + S S L S P G ++ ++ G G + V +
Sbjct: 130 DGNLGEEQPLK-RTASGGGGVSTGLYMSPGS-PSGSDVSEQSSG----GAHVFKPTVRSE 183
Query: 178 DEESRDGVEEAETMT-----EEEVLASSVPTSLS------------LFPMGEKKMVAVAE 220
S G + ++ +E + + P L+ LFP+ +++ V V E
Sbjct: 184 VTASSSGEDPPTYLSLSLPWTDETVRVNEPVQLNQNTVMDGGYTAELFPVRKEEQVEVEE 243
Query: 221 KEEKKMDMQ-EGLMRTMMQRMIAEEVRAY 248
+E K + G T++Q MI EVR+Y
Sbjct: 244 EEAKGISGGFGGEFMTVVQEMIRTEVRSY 272
>At2g23290 putative MYB family transcription factor
Length = 309
Score = 174 bits (442), Expect = 3e-44
Identities = 114/272 (41%), Positives = 147/272 (53%), Gaps = 37/272 (13%)
Query: 9 NGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSP 68
+G + DR+KG WSP+ED+ L LV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP
Sbjct: 2 SGSTRKEMDRIKGPWSPEEDDLLQSLVQKHGPRNWSLISKSIPGRSGKSCRLRWCNQLSP 61
Query: 69 DVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-------- 120
+V+HR FT ED II AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+
Sbjct: 62 EVEHRGFTAEEDDTIILAHARFGNKWATIARLLNGRTDNAIKNHWNSTLKRKCSGGGGGG 121
Query: 121 -RGESLPL--------PLASPSP---SPSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
G+S L P S + A P G ++ ++ P
Sbjct: 122 EEGQSCDFGGNGGYDGNLTDEKPLKRRASGGGGVVVVTALSPTGSDVSEQSQS-SGSVLP 180
Query: 169 FKRHC-VEKDDEESRDGVEEAETMTEEE-----VLASSVP------TSLSLFPMGEKKMV 216
C V K + V E+ + EEE L S+P T LFP+ ++
Sbjct: 181 VSSSCHVFKPTARAGGVVIESSSPEEEEKDPMTCLRLSLPWVNESTTPPELFPVKREE-- 238
Query: 217 AVAEKEEKKMDMQEGLMRTMMQRMIAEEVRAY 248
E++E+++ G T++Q MI EVR+Y
Sbjct: 239 --EEEKEREISGLGGDFMTVVQEMIKTEVRSY 268
>At5g67300 myb-related protein, 33.3K (pir||S71284)
Length = 305
Score = 173 bits (439), Expect = 7e-44
Identities = 77/106 (72%), Positives = 90/106 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WSP+EDE L +LV ++GPRNW+VIS IPGRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 3 DRIKGPWSPEEDEQLRRLVVKYGPRNWTVISKSIPGRSGKSCRLRWCNQLSPQVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRG 122
ED I +AHA GNKWATI+RLL GRTDNA+KNHWNSTL+R+ G
Sbjct: 63 AEEDETIARAHAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRKCG 108
>At3g50060 R2R3-MYB transcription factor
Length = 301
Score = 170 bits (431), Expect = 6e-43
Identities = 75/104 (72%), Positives = 88/104 (84%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DRVKG WS +EDE L ++V ++GPRNWS IS IPGRSGKSCRLRWCNQLSP+V+HRPF+
Sbjct: 3 DRVKGPWSQEEDEQLRRMVEKYGPRNWSAISKSIPGRSGKSCRLRWCNQLSPEVEHRPFS 62
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
P ED I+ A A GNKWATI+RLL GRTDNA+KNHWNSTL+R+
Sbjct: 63 PEEDETIVTARAQFGNKWATIARLLNGRTDNAVKNHWNSTLKRK 106
>At3g09230 unknown protein
Length = 393
Score = 164 bits (416), Expect = 3e-41
Identities = 75/109 (68%), Positives = 86/109 (78%)
Query: 12 GGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ 71
G G RDRVKG WS +ED+ L +LV + G RNWS I+ IPGRSGKSCRLRWCNQL+P++
Sbjct: 47 GRGRRDRVKGPWSKEEDDVLSELVKRLGARNWSFIARSIPGRSGKSCRLRWCNQLNPNLI 106
Query: 72 HRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
FT ED II AHA+HGNKWA I++LLPGRTDNAIKNHWNS LRRR
Sbjct: 107 RNSFTEVEDQAIIAAHAIHGNKWAVIAKLLPGRTDNAIKNHWNSALRRR 155
>At2g39880 putative MYB family transcription factor
Length = 367
Score = 164 bits (414), Expect = 6e-41
Identities = 72/123 (58%), Positives = 93/123 (75%)
Query: 7 SSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQL 66
+ + GG + +VKG W P++DE L +LV GPRNW++IS GIPGRSGKSCRLRWCNQL
Sbjct: 37 AKSDANGGGKSKVKGPWLPEQDEALTRLVKMCGPRNWNLISRGIPGRSGKSCRLRWCNQL 96
Query: 67 SPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLP 126
P ++ +PF+ E+ +I+ A AV GNKW+ I++LLPGRTDNAIKNHWNS LRR+ E
Sbjct: 97 DPILKRKPFSDEEEHMIMSAQAVLGNKWSVIAKLLPGRTDNAIKNHWNSNLRRKPAEQWK 156
Query: 127 LPL 129
+PL
Sbjct: 157 IPL 159
>At1g73410 putative protein
Length = 243
Score = 143 bits (360), Expect = 1e-34
Identities = 70/142 (49%), Positives = 88/142 (61%), Gaps = 3/142 (2%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P EDE L LV Q+GP NW+ I+ +PGRSGKSCRLRW NQL P + PFT E
Sbjct: 6 RGHWRPAEDEKLKDLVEQYGPHNWNAIALKLPGRSGKSCRLRWFNQLDPRINRNPFTEEE 65
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTL-RRRRGESLP--LPLASPSPSP 136
+ ++ AH +HGN+W+ I+RL PGRTDNA+KNHW+ + RR R S P LP + S S
Sbjct: 66 EERLLAAHRIHGNRWSIIARLFPGRTDNAVKNHWHVIMARRTRQTSKPRLLPSTTSSSSL 125
Query: 137 SPSPPLPFSMAKRPHGYELLDR 158
S + S H Y DR
Sbjct: 126 MASEQIMMSSGGYNHNYSSDDR 147
>At3g09370 MYB family like transcription factor
Length = 505
Score = 134 bits (336), Expect = 6e-32
Identities = 62/133 (46%), Positives = 91/133 (67%), Gaps = 2/133 (1%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +KG W+ +EDE +++LV ++GP WS+I+ +PGR GK CR RW N L+PD+ +T
Sbjct: 127 DLIKGPWTHEEDEKIVELVEKYGPAKWSIIAQSLPGRIGKQCRERWHNHLNPDINKDAWT 186
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSP 136
E+ ++ AH HGNKWA I+++LPGRTDNAIKNHWNS+L +++ E L P P+
Sbjct: 187 TEEEVALMNAHRSHGNKWAEIAKVLPGRTDNAIKNHWNSSL-KKKSEFYLLTGRLPPPTT 245
Query: 137 SPSPPLPFSMAKR 149
+ + +P S+ KR
Sbjct: 246 TRN-GVPDSVTKR 257
>At5g17800 MYB56 R2R3-MYB factor family member
Length = 323
Score = 133 bits (334), Expect = 1e-31
Identities = 60/105 (57%), Positives = 74/105 (70%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P ED L +LV Q GP+NW++IS + GRSGKSCRLRW NQL P + R FT E
Sbjct: 93 RGHWRPTEDAKLKELVAQFGPQNWNLISNHLLGRSGKSCRLRWFNQLDPRINKRAFTEEE 152
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGES 124
+ ++ AH +GNKWA ISRL PGRTDNA+KNHW+ + RR ES
Sbjct: 153 EFRLLAAHRAYGNKWALISRLFPGRTDNAVKNHWHVIMARRTRES 197
>At4g33450 putative transcription factor (MYB69)
Length = 250
Score = 132 bits (332), Expect = 2e-31
Identities = 55/105 (52%), Positives = 80/105 (75%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P ED+ L +LV Q+GP+NW+ I+ + GRSGKSCRLRW NQL P++ +PFT E
Sbjct: 19 RGHWRPVEDDNLRQLVEQYGPKNWNFIAQHLYGRSGKSCRLRWYNQLDPNITKKPFTEEE 78
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGES 124
+ +++AH + GN+WA+I+RL PGRTDNA+KNH++ + RR+ E+
Sbjct: 79 EERLLKAHRIQGNRWASIARLFPGRTDNAVKNHFHVIMARRKREN 123
>At1g69560 myb-related transcription factor, putative
Length = 330
Score = 131 bits (330), Expect = 3e-31
Identities = 61/126 (48%), Positives = 83/126 (65%), Gaps = 11/126 (8%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P ED L +LV +GP+NW++I+ + GRSGKSCRLRW NQL P + R FT E
Sbjct: 107 RGHWRPAEDTKLKELVAVYGPQNWNLIAEKLQGRSGKSCRLRWFNQLDPRINRRAFTEEE 166
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHW-----------NSTLRRRRGESLPLP 128
+ ++QAH ++GNKWA I+RL PGRTDN++KNHW +S+ RRR+ P
Sbjct: 167 EERLMQAHRLYGNKWAMIARLFPGRTDNSVKNHWHVIMARKFREQSSSYRRRKTMVSLKP 226
Query: 129 LASPSP 134
L +P+P
Sbjct: 227 LINPNP 232
>At1g17950 unknown protein
Length = 249
Score = 131 bits (330), Expect = 3e-31
Identities = 55/104 (52%), Positives = 72/104 (68%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P EDE L +LV Q GP NW+ I+ + GRSGKSCRLRW NQL P + PFT E
Sbjct: 5 RGHWRPAEDEKLRELVEQFGPHNWNAIAQKLSGRSGKSCRLRWFNQLDPRINRNPFTEEE 64
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGE 123
+ ++ +H +HGN+W+ I+R PGRTDNA+KNHW+ + RR E
Sbjct: 65 EERLLASHRIHGNRWSVIARFFPGRTDNAVKNHWHVIMARRGRE 108
>At1g26780 putative transcription factor MYB117 (MYB117)
Length = 280
Score = 129 bits (323), Expect = 2e-30
Identities = 55/104 (52%), Positives = 75/104 (71%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPTE 79
+G W P ED L +LV +GP+NW++I+ + GRSGKSCRLRW NQL P + R FT E
Sbjct: 98 RGHWRPAEDVKLKELVSIYGPQNWNLIAEKLQGRSGKSCRLRWFNQLDPRINRRAFTEEE 157
Query: 80 DTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGE 123
+ ++QAH ++GNKWA I+RL PGRTDN++KNHW+ + R+ E
Sbjct: 158 EERLMQAHRLYGNKWAMIARLFPGRTDNSVKNHWHVVMARKYRE 201
>At5g02320 myb -like protein
Length = 529
Score = 127 bits (318), Expect = 8e-30
Identities = 51/102 (50%), Positives = 79/102 (77%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
VKG W+ +ED+ +++LV ++GP WSVI+ +PGR GK CR RW N L+P ++ +T
Sbjct: 107 VKGPWTQEEDDKIVELVKKYGPAKWSVIAKSLPGRIGKQCRERWHNHLNPGIRKDAWTVE 166
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E++ ++ +H ++GNKWA I+++LPGRTDNAIKNHWNS+L+++
Sbjct: 167 EESALMNSHRMYGNKWAEIAKVLPGRTDNAIKNHWNSSLKKK 208
Score = 30.0 bits (66), Expect = 1.3
Identities = 13/52 (25%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ 71
K +W+ +E+ LM +G + W+ I+ +PGR+ + + W + L ++
Sbjct: 160 KDAWTVEEESALMNSHRMYGNK-WAEIAKVLPGRTDNAIKNHWNSSLKKKLE 210
>At4g32730 putative myb-protein
Length = 776
Score = 126 bits (317), Expect = 1e-29
Identities = 51/102 (50%), Positives = 75/102 (73%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
VKG WS +ED T++ LV ++GP+ WS IS +PGR GK CR RW N L+P + +T
Sbjct: 86 VKGPWSKEEDNTIIDLVEKYGPKKWSTISQHLPGRIGKQCRERWHNHLNPGINKNAWTQE 145
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +I+AH ++GNKWA + + LPGR+DN+IKNHWNS+++++
Sbjct: 146 EELTLIRAHQIYGNKWAELMKFLPGRSDNSIKNHWNSSVKKK 187
>At4g26930 putative myb-related protein
Length = 389
Score = 125 bits (315), Expect = 2e-29
Identities = 71/159 (44%), Positives = 91/159 (56%), Gaps = 17/159 (10%)
Query: 3 GGGDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNW-SVISAGIPGRSGKSCRLR 61
GGG S +G GGG + KG W+ EDETL V ++G NW SV R GKSCRLR
Sbjct: 5 GGGASEDGEGGGVVLK-KGPWTVAEDETLAAYVREYGEGNWNSVQKKTWLARCGKSCRLR 63
Query: 62 WCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRR 121
W N L P+++ FTP E+ +IIQ H+ GNKWA ++ LPGRTDN IKN+WN+ L+R +
Sbjct: 64 WANHLRPNLRKGSFTPEEERLIIQLHSQLGNKWARMAAQLPGRTDNEIKNYWNTRLKRFQ 123
Query: 122 GESLPL---------------PLASPSPSPSPSPPLPFS 145
+ LPL P SP PS +P F+
Sbjct: 124 RQGLPLYPPEYSQNNHQQQMYPQQPSSPLPSQTPASSFT 162
>At5g58850 putative transcription factor MYB119 (MYB119)
Length = 430
Score = 124 bits (312), Expect = 4e-29
Identities = 61/168 (36%), Positives = 101/168 (59%), Gaps = 13/168 (7%)
Query: 16 RDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPF 75
++ +KG W+ +ED L++LV QHG R W++IS + GR+GK CR RW N L PD++ +
Sbjct: 101 KNLIKGQWTAEEDRKLIRLVRQHGERKWAMISEKLEGRAGKQCRERWHNHLRPDIKKDGW 160
Query: 76 TPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPS 135
+ E+ +++++H GNKWA I++L+PGRT+N+IKNHWN+T RR+ + ++ +
Sbjct: 161 SEEEERVLVESHMRIGNKWAEIAKLIPGRTENSIKNHWNATKRRQNSKRKHKRESNADNN 220
Query: 136 PSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCVEKDDEESRD 183
+ P AKRP L D +R+ + KD++E ++
Sbjct: 221 DRDASP----SAKRP---------CILQDYIKSIERNNINKDNDEKKN 255
>At5g11050 putative transcription factor MYB64 (MYB64)
Length = 423
Score = 123 bits (309), Expect = 9e-29
Identities = 52/105 (49%), Positives = 75/105 (70%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+ ED L+KLV QHG R W+VIS + GR+GK CR RW N L PD++ ++
Sbjct: 104 IKGQWTADEDRKLIKLVMQHGERKWAVISEKLEGRAGKQCRERWHNHLRPDIKKDSWSEE 163
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGE 123
E+ ++++AH GNKWA I++L+ GRT+N+IKNHWN+T RR+ +
Sbjct: 164 EERLLVEAHTRIGNKWAEIAKLIQGRTENSIKNHWNATKRRQNSK 208
Score = 27.3 bits (59), Expect = 8.4
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRW 62
D K SWS +E+ L++ + G + W+ I+ I GR+ S + W
Sbjct: 154 DIKKDSWSEEEERLLVEAHTRIGNK-WAEIAKLIQGRTENSIKNHW 198
>At3g27785 putative transcription factor MYB118 (MYB118)
Length = 437
Score = 123 bits (309), Expect = 9e-29
Identities = 54/102 (52%), Positives = 73/102 (70%)
Query: 19 VKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFTPT 78
+KG W+P+ED+ L++LV HG + WS I+ + GR GK CR RW N L PD++ +T
Sbjct: 188 IKGQWTPEEDKLLVQLVDLHGTKKWSQIAKMLQGRVGKQCRERWHNHLRPDIKKDGWTEE 247
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED I+I+AH GN+WA I+R LPGRT+N IKNHWN+T RR+
Sbjct: 248 EDIILIKAHKEIGNRWAEIARKLPGRTENTIKNHWNATKRRQ 289
Score = 30.0 bits (66), Expect = 1.3
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRW 62
D K W+ +ED L+K + G R W+ I+ +PGR+ + + W
Sbjct: 238 DIKKDGWTEEEDIILIKAHKEIGNR-WAEIARKLPGRTENTIKNHW 282
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,869,162
Number of Sequences: 26719
Number of extensions: 346865
Number of successful extensions: 4016
Number of sequences better than 10.0: 236
Number of HSP's better than 10.0 without gapping: 188
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 3164
Number of HSP's gapped (non-prelim): 568
length of query: 260
length of database: 11,318,596
effective HSP length: 97
effective length of query: 163
effective length of database: 8,726,853
effective search space: 1422477039
effective search space used: 1422477039
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0002.6


