

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0001.6
(381 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
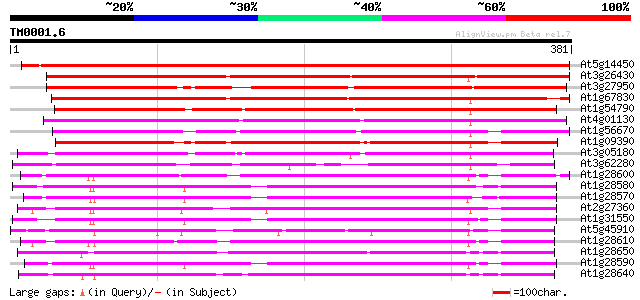
Score E
Sequences producing significant alignments: (bits) Value
At5g14450 early nodule-specific protein - like 517 e-147
At3g26430 nodulin, putative 351 4e-97
At3g27950 early nodule-specific protein, putative 333 9e-92
At1g67830 ENOD8-like protein 330 6e-91
At1g54790 293 1e-79
At4g01130 putative acetyltransferase 276 2e-74
At1g56670 lipase homolog (Lip-4) 253 1e-67
At1g09390 putative lipase 250 8e-67
At3g05180 putative nodulin 248 4e-66
At3g62280 putative protein 189 3e-48
At1g28600 lipase like protein 181 8e-46
At1g28580 lipase like protein 171 6e-43
At1g28570 lipase like protein 163 1e-40
At2g27360 putative lipase 163 2e-40
At1g31550 unknown protein 163 2e-40
At5g45910 GDSL-motif lipase/hydrolase-like protein 162 2e-40
At1g28610 lipase, putative 161 6e-40
At1g28650 lipase, putative 159 3e-39
At1g28590 putative lipase (At1g28590) 157 1e-38
At1g28640 lipase, putative 154 1e-37
>At5g14450 early nodule-specific protein - like
Length = 389
Score = 517 bits (1331), Expect = e-147
Identities = 245/372 (65%), Positives = 288/372 (76%), Gaps = 1/372 (0%)
Query: 9 AFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSA 68
++ L C V + + P C FPAIYNFGDSNSDTGGISAAFEPI PYG+ F +P+
Sbjct: 18 SWLLLCLFAVTT-SVSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDPYGQGFFHRPTG 76
Query: 69 RDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPF 128
RD DGRL +DFIAE+L LPYLSAYLNSLG+N+RHGANFATGGSTIRRQNETIFQYGISPF
Sbjct: 77 RDSDGRLTIDFIAERLGLPYLSAYLNSLGSNFRHGANFATGGSTIRRQNETIFQYGISPF 136
Query: 129 SLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMN 188
SLDMQ QF QFKAR+ L+ + K+ +R KLP EEF+KALYTFDIGQNDLSVGFR M+
Sbjct: 137 SLDMQIAQFDQFKARSALLFTQIKSRYDREKLPRQEEFAKALYTFDIGQNDLSVGFRTMS 196
Query: 189 FDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYG 248
DQ++ ++PDIVN LASAV+NIY+ GGRTFW+HNT P GCLPVN+FY GYLD G
Sbjct: 197 VDQLKATIPDIVNHLASAVRNIYQQGGRTFWVHNTGPFGCLPVNMFYMGTPAPGYLDKSG 256
Query: 249 CVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKIC 308
CVK QN MA+EFN++LK+ V+ LR EL +AAITYVD+Y AKY ++SN K GF +PLK+C
Sbjct: 257 CVKAQNEMAMEFNRKLKETVINLRKELTQAAITYVDVYTAKYEMMSNPKKLGFANPLKVC 316
Query: 309 CGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFT 368
CGYH HIWCG G N +++G +C P M VSWDGVHY EAAN VA+R LNG T
Sbjct: 317 CGYHEKYDHIWCGNKGKVNNTEIYGGSCPNPVMAVSWDGVHYTEAANKHVADRTLNGLLT 376
Query: 369 DPPTLITQACYR 380
DPP IT+ACYR
Sbjct: 377 DPPVPITRACYR 388
>At3g26430 nodulin, putative
Length = 380
Score = 351 bits (900), Expect = 4e-97
Identities = 179/361 (49%), Positives = 241/361 (66%), Gaps = 10/361 (2%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
SP C FPAI+NFGDSNSDTGG+SA+F P P G++F PS R DGRLI+DFIAE+L
Sbjct: 24 SPSCNFPAIFNFGDSNSDTGGLSASFGQAPYPNGQTFFHSPSGRFSDGRLIIDFIAEELG 83
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPYL+A+L+S+G+N+ HGANFAT GST+R N TI Q G+SP SLD+Q VQF F R++
Sbjct: 84 LPYLNAFLDSIGSNFSHGANFATAGSTVRPPNATIAQSGVSPISLDVQLVQFSDFITRSQ 143
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRM-MNFDQMRESMPDIVNQLA 204
+ + + + LP E FS+ALYTFDIGQNDL+ G ++ M DQ++ +PD+ +QL+
Sbjct: 144 LI--RNRGGVFKKLLPKKEYFSQALYTFDIGQNDLTAGLKLNMTSDQIKAYIPDVHDQLS 201
Query: 205 SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQL 264
+ ++ +Y GGR FWIHNTAP+GCLP + + +PA +D +GC +N +A +N +L
Sbjct: 202 NVIRKVYSKGGRRFWIHNTAPLGCLPY-VLDRFPVPASQIDNHGCAIPRNEIARYYNSEL 260
Query: 265 KDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCG----YHVNDTHIWC 320
K RV++LR EL EAA TYVD+Y+ K LI+ K GF PL CCG Y+ N I C
Sbjct: 261 KRRVIELRKELSEAAFTYVDIYSIKLTLITQAKKLGFRYPLVACCGHGGKYNFNKL-IKC 319
Query: 321 GTIGTANGKD-VFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACY 379
G GK+ V +C S VSWDG+H+ E N W+ +I +G+F+DPP + AC
Sbjct: 320 GAKVMIKGKEIVLAKSCNDVSFRVSWDGIHFTETTNSWIFQQINDGAFSDPPLPVKSACT 379
Query: 380 R 380
R
Sbjct: 380 R 380
>At3g27950 early nodule-specific protein, putative
Length = 361
Score = 333 bits (854), Expect = 9e-92
Identities = 172/353 (48%), Positives = 226/353 (63%), Gaps = 21/353 (5%)
Query: 26 SPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLN 85
S C FPA++NFGDSNSDTG ISAA +PPP G +F + + R DGRLI+DFI E L
Sbjct: 25 SSSCNFPAVFNFGDSNSDTGAISAAIGEVPPPNGVAFFGRSAGRHSDGRLIIDFITENLT 84
Query: 86 LPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTK 145
LPYL+ YL+S+G NYRHGANFATGGS IR T+ + SPF L Q QF FK RT
Sbjct: 85 LPYLTPYLDSVGANYRHGANFATGGSCIR---PTLACF--SPFHLGTQVSQFIHFKTRTL 139
Query: 146 QLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLAS 205
LY + +FSKALYT DIGQNDL++GF+ M +Q++ ++P I+
Sbjct: 140 SLYNQT------------NDFSKALYTLDIGQNDLAIGFQNMTEEQLKATIPLIIENFTI 187
Query: 206 AVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLK 265
A+K +Y+ G R F IHNT P GCLP + PA DPYGC+K N +A+EFNKQLK
Sbjct: 188 ALKLLYKEGARFFSIHNTGPTGCLP---YLLKAFPAIPRDPYGCLKPLNNVAIEFNKQLK 244
Query: 266 DRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGT 325
+++ +L+ ELP + TYVD+Y+AKY LI+ K GF+DP CC + + CG
Sbjct: 245 NKITQLKKELPSSFFTYVDVYSAKYNLITKAKALGFIDPFDYCCVGAIG-RGMGCGKTIF 303
Query: 326 ANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
NG +++ ++C+ ++SWDG+HY E AN VANRIL+GS +DPP +AC
Sbjct: 304 LNGTELYSSSCQNRKNFISWDGIHYTETANMLVANRILDGSISDPPLPTQKAC 356
>At1g67830 ENOD8-like protein
Length = 372
Score = 330 bits (847), Expect = 6e-91
Identities = 173/357 (48%), Positives = 233/357 (64%), Gaps = 16/357 (4%)
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI+NFGDSNSDTGG+SAAF PP+G SF P+ R CDGRL++DFIAE L LPY
Sbjct: 26 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 85
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LSA+L+S+G+N+ HGANFAT GS IR N T+ Q G SPFSLD+QFVQF F R++ +
Sbjct: 86 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTV- 144
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
++ + ++ LP + FSKALYTFDIGQNDL+ G F +Q+ +P+I++Q +A+
Sbjct: 145 -RSRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAI 203
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY GGR FWIHNT PIGCL + + A D +GCV N +A +FN LK
Sbjct: 204 KNIYGQGGRYFWIHNTGPIGCL-AYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQA 262
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVNDTHIWCGTIG 324
V++LR+ L EAAITYVD+Y+ K+ L + + GF L CCG+ + + I CG
Sbjct: 263 VIELRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKK 322
Query: 325 TANGKDVF-GNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
GK+V+ G C++P V WDGVH+ +AAN ++ ++I G +++AC R
Sbjct: 323 IVKGKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKR 371
>At1g54790
Length = 383
Score = 293 bits (750), Expect = 1e-79
Identities = 156/347 (44%), Positives = 212/347 (60%), Gaps = 12/347 (3%)
Query: 31 FPAIYNFGDSNSDTGG-ISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYL 89
+PAI NFGDSNSDTG ISA E + PPYG+++ PS R CDGRLIVDF+ ++++LP+L
Sbjct: 30 YPAIINFGDSNSDTGNLISAGIENVNPPYGQTYFNLPSGRYCDGRLIVDFLLDEMDLPFL 89
Query: 90 SAYLNSLGT-NYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
+ YL+SLG N++ G NFA GSTI N T +SPFS D+Q QF +FK+R +L
Sbjct: 90 NPYLDSLGLPNFKKGCNFAAAGSTILPANPT----SVSPFSFDLQISQFIRFKSRAIELL 145
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLASAVK 208
+ E+ LP + +SK LY DIGQND++ F DQ+ S+P I+ + +K
Sbjct: 146 SKTGRKYEKY-LPPIDYYSKGLYMIDIGQNDIAGAFYSKTLDQVLASIPSILETFEAGLK 204
Query: 209 NIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRV 268
+YE GGR WIHNT P+GCL N+ K + LD +GCV N A FN QL
Sbjct: 205 RLYEEGGRNIWIHNTGPLGCLAQNI-AKFGTDSTKLDEFGCVSSHNQAAKLFNLQLHAMS 263
Query: 269 VKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY---HVN-DTHIWCGTIG 324
K + + P+A +TYVD+++ K LI+N GF PL CCG +N D+ I CG
Sbjct: 264 NKFQAQYPDANVTYVDIFSIKSNLIANYSRFGFEKPLMACCGVGGAPLNYDSRITCGQTK 323
Query: 325 TANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPP 371
+G V AC S Y++WDG+HY EAAN +V+++IL G ++DPP
Sbjct: 324 VLDGISVTAKACNDSSEYINWDGIHYTEAANEFVSSQILTGKYSDPP 370
>At4g01130 putative acetyltransferase
Length = 382
Score = 276 bits (705), Expect = 2e-74
Identities = 146/360 (40%), Positives = 205/360 (56%), Gaps = 9/360 (2%)
Query: 24 KNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEK 83
K C F AI+NFGDSNSDTGG AAF P+G ++ +KP+ R DGRLI+DF+A+
Sbjct: 25 KGDSKCDFEAIFNFGDSNSDTGGFWAAFPAQSGPWGMTYFKKPAGRASDGRLIIDFLAKS 84
Query: 84 LNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKAR 143
L +P+LS YL S+G+++RHGANFAT ST+ N ++F GISPFSL +Q Q KQFK
Sbjct: 85 LGMPFLSPYLQSIGSDFRHGANFATLASTVLLPNTSLFVSGISPFSLAIQLNQMKQFKVN 144
Query: 144 TKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQL 203
+ + + L+ LP F K+LYTF IGQND + + ++++ +P ++ Q+
Sbjct: 145 VDESHSLDRPGLK--ILPSKIVFGKSLYTFYIGQNDFTSNLASIGVERVKLYLPQVIGQI 202
Query: 204 ASAVKNIYELGGRTFWIHNTAPIGCLPVNLF-YKHNLPAGYLDPYGCVKDQNVMAVEFNK 262
A +K IY +GGRTF + N AP+GC P L Y H LD YGC+ N +N
Sbjct: 203 AGTIKEIYGIGGRTFLVLNLAPVGCYPAILTGYTHT--DADLDKYGCLIPVNKAVKYYNT 260
Query: 263 QLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTHI 318
L + + RTEL A + Y+D + L + K+ G +K CCGY + + +
Sbjct: 261 LLNKTLSQTRTELKNATVIYLDTHKILLDLFQHPKSYGMKHGIKACCGYGGRPYNFNQKL 320
Query: 319 WCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQAC 378
+CG AC P YVSWDG+H EAANH ++ IL+GS + PP ++ C
Sbjct: 321 FCGNTKVIGNFSTTAKACHDPHNYVSWDGIHATEAANHHISMAILDGSISYPPFILNNLC 380
>At1g56670 lipase homolog (Lip-4)
Length = 373
Score = 253 bits (646), Expect = 1e-67
Identities = 146/358 (40%), Positives = 204/358 (56%), Gaps = 28/358 (7%)
Query: 30 AFPAIYNFGDSNSDTGGISAAFE-PIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
A P I+NFGDSNSDTGG+ A PI P G F ++ + R DGRL++DF+ + LN
Sbjct: 37 ARPVIFNFGDSNSDTGGLVAGLGYPIGFPNGRLFFRRSTGRLSDGRLLIDFLCQSLNTSL 96
Query: 89 LSAYLNSLG-TNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQL 147
L YL+SLG T +++GANFA GS +N PFSL++Q QF FK+R+ +L
Sbjct: 97 LRPYLDSLGRTRFQNGANFAIAGSPTLPKNV--------PFSLNIQVKQFSHFKSRSLEL 148
Query: 148 YQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMN-FDQMRESMPDIVNQLASA 206
+ + + F ALY DIGQND++ F N + Q + +P I+ ++ S+
Sbjct: 149 ASSSNSL--KGMFISNNGFKNALYMIDIGQNDIARSFARGNSYSQTVKLIPQIITEIKSS 206
Query: 207 VKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKD 266
+K +Y+ GGR FWIHNT P+GCLP L + + LD +GC+ N A FN+ L
Sbjct: 207 IKRLYDEGGRRFWIHNTGPLGCLPQKLSM---VKSKDLDQHGCLVSYNSAATLFNQGLDH 263
Query: 267 RVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTHIWCGT 322
+LRTEL +A I Y+D+YA KY LI+N+ GF PL CCGY + + I CG
Sbjct: 264 MCEELRTELRDATIIYIDIYAIKYSLIANSNQYGFKSPLMACCGYGGTPYNYNVKITCGH 323
Query: 323 IGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
G+ N CE+ S ++SWDG+HY E AN VA ++L+ ++ PPT C R
Sbjct: 324 KGS--------NVCEEGSRFISWDGIHYTETANAIVAMKVLSMHYSKPPTPFHFFCRR 373
>At1g09390 putative lipase
Length = 370
Score = 250 bits (639), Expect = 8e-67
Identities = 144/349 (41%), Positives = 214/349 (61%), Gaps = 30/349 (8%)
Query: 32 PAIYNFGDSNSDTGGISAAFE-PIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLS 90
P I+NFGDSNSDTGG+ A I P G SF Q+ + R DGRL++DF+ + LN L+
Sbjct: 36 PVIFNFGDSNSDTGGLVAGLGYSIGLPNGRSFFQRSTGRLSDGRLVIDFLCQSLNTSLLN 95
Query: 91 AYLNSL-GTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQ 149
YL+SL G+ +++GANFA GS+ T+ +Y PF+L++Q +QF FK+R +L
Sbjct: 96 PYLDSLVGSKFQNGANFAIVGSS------TLPRY--VPFALNIQLMQFLHFKSRALEL-- 145
Query: 150 EAKTALERSKLPVPEE-FSKALYTFDIGQNDLSVGF-RMMNFDQMRESMPDIVNQLASAV 207
A + ++ + E F ALY DIGQND++ F + +++ ++ + +P++++++ SA+
Sbjct: 146 -ASISDPLKEMMIGESGFRNALYMIDIGQNDIADSFSKGLSYSRVVKLIPNVISEIKSAI 204
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
K +Y+ GGR FW+HNT P+GCLP L H+ G+ D +GC+ N A FN+ L
Sbjct: 205 KILYDEGGRKFWVHNTGPLGCLPQKLSMVHS--KGF-DKHGCLATYNAAAKLFNEGLDHM 261
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGY----HVNDTHIWCGTI 323
LRTEL EA I YVD+YA KY LI+N+ N GF PL CCGY + + +I CG
Sbjct: 262 CRDLRTELKEANIVYVDIYAIKYDLIANSNNYGFEKPLMACCGYGGPPYNYNVNITCGNG 321
Query: 324 GTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPT 372
G+ +C++ S ++SWDG+HY E AN VA ++L+ + PPT
Sbjct: 322 GS--------KSCDEGSRFISWDGIHYTETANAIVAMKVLSMQHSTPPT 362
>At3g05180 putative nodulin
Length = 379
Score = 248 bits (633), Expect = 4e-66
Identities = 149/373 (39%), Positives = 206/373 (54%), Gaps = 22/373 (5%)
Query: 6 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPP-YGESFPQ 64
LF+A + + S + N FPA++NFGDSNSDTG +S+ +P P Y +F +
Sbjct: 13 LFVAISHTLSPLAGSFRISND----FPAVFNFGDSNSDTGELSSGLGFLPQPSYEITFFR 68
Query: 65 KP-SARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTN-YRHGANFATGGSTIRRQNETIFQ 122
P S R C+GRLIVDF+ E ++ PYL YL+S+ YR G NFA STI++ N +
Sbjct: 69 SPTSGRFCNGRLIVDFLMEAIDRPYLRPYLDSISRQTYRRGCNFAAAASTIQKANAASY- 127
Query: 123 YGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSV 182
SPF +Q QF FK++ QL Q+ + L+R LP FS LY FDIGQND++
Sbjct: 128 ---SPFGFGVQVSQFITFKSKVLQLIQQDEE-LQRY-LPSEYFFSNGLYMFDIGQNDIAG 182
Query: 183 GFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP--VNLFYKHNLP 240
F DQ+ +P I++ +K +Y G R +WIHNT P+GCL V++F +
Sbjct: 183 AFYTKTVDQVLALVPIILDIFQDGIKRLYAEGARNYWIHNTGPLGCLAQVVSIFGEDK-- 240
Query: 241 AGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEG 300
LD +GCV D N A FN QL KL + P + TYVD+++ K LI N G
Sbjct: 241 -SKLDEFGCVSDHNQAAKLFNLQLHGLFKKLPQQYPNSRFTYVDIFSIKSDLILNHSKYG 299
Query: 301 FVDPLKICCGY---HVN-DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANH 356
F + +CCG +N D + CG +NG + C S YV+WDG+HY EAAN
Sbjct: 300 FDHSIMVCCGTGGPPLNYDDQVGCGKTARSNGTIITAKPCYDSSKYVNWDGIHYTEAANR 359
Query: 357 WVANRILNGSFTD 369
+VA IL G +++
Sbjct: 360 FVALHILTGKYSE 372
>At3g62280 putative protein
Length = 343
Score = 189 bits (479), Expect = 3e-48
Identities = 132/375 (35%), Positives = 183/375 (48%), Gaps = 54/375 (14%)
Query: 3 LRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFE-PIPPPYGES 61
L+ F+ LS +C NS S P + NFGDSNSDTGG+ A PI P+G +
Sbjct: 8 LQCFFLVLCLSLLVCSNSETSYKSNKK--PILINFGDSNSDTGGVLAGVGLPIGLPHGIT 65
Query: 62 FPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIF 121
F + + R DGRLIVDF E L + YLS YL+SL N++ G NFA G+T
Sbjct: 66 FFHRGTGRLGDGRLIVDFYCEHLKMTYLSPYLDSLSPNFKRGVNFAVSGATA-------- 117
Query: 122 QYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLS 181
I F L +Q QF FK R+++L R L F ALY DIGQNDL
Sbjct: 118 -LPIFSFPLAIQIRQFVHFKNRSQELISSG-----RRDLIDDNGFRNALYMIDIGQNDLL 171
Query: 182 VGFRMMN--FDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNL 239
+ N + + E +P ++ ++ A++ G +HN +
Sbjct: 172 LALYDSNLTYAPVVEKIPSMLLEIKKAIQ-----GELAIHLHNDSD-------------- 212
Query: 240 PAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNE 299
LDP GC + N +A FNK L +LR++ +A + YVD+Y+ KY L ++ K
Sbjct: 213 ----LDPIGCFRVHNEVAKAFNKGLLSLCNELRSQFKDATLVYVDIYSIKYKLSADFKLY 268
Query: 300 GFVDPLKICCGY----HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAAN 355
GFVDPL CCGY + D CG G+ +DV + + WDGVHY EAAN
Sbjct: 269 GFVDPLMACCGYGGRPNNYDRKATCGQPGSTICRDV--------TKAIVWDGVHYTEAAN 320
Query: 356 HWVANRILNGSFTDP 370
+V + +L ++ P
Sbjct: 321 RFVVDAVLTNRYSYP 335
>At1g28600 lipase like protein
Length = 393
Score = 181 bits (458), Expect = 8e-46
Identities = 129/388 (33%), Positives = 187/388 (47%), Gaps = 37/388 (9%)
Query: 8 IAFFLSCTLCVNSVELKNSPPCA-FPAIYNFGDSNSDTGGISAAFE----PIP--PPYGE 60
+ FL TL V V + PC F +I +FGDS +DTG + + P+ PPYGE
Sbjct: 7 LVIFLFSTLFVTIVS--SETPCPNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPYGE 64
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
+F P+ R CDGR+I+DFIAE + LPY+ Y S N+ G NFA G+T + + +
Sbjct: 65 TFFHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFGSKNRNFDKGVNFAVAGATALK-SSFL 123
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF-DIGQND 179
+ GI P + VQ K FK L S + AL +IG ND
Sbjct: 124 KKRGIQPHTNVSLGVQLKSFKKSLPNLC--------GSPSDCRDMIGNALILMGEIGGND 175
Query: 180 LSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL-FYKH 237
+ F ++ E +P ++ ++S + + +GG+TF + PIGC V L YK
Sbjct: 176 YNFPFFNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKT 235
Query: 238 NLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTK 297
+ Y GC+K N +++LK + +LR P I Y D Y + +
Sbjct: 236 SNKDEYDPSTGCLKWLNKFGEYHSEKLKVELNRLRKLYPHVNIIYADYYNSLLRIFKEPA 295
Query: 298 NEGFVD-PLKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAE 352
GF++ P CCG Y+ N T CG++G +C+ PS YV WDGVH E
Sbjct: 296 KFGFMERPFPACCGIGGPYNFNFTR-KCGSVGV--------KSCKDPSKYVGWDGVHMTE 346
Query: 353 AANHWVANRILNGSFTDPPTLITQACYR 380
AA W+A+ ILNG + +PP ++C R
Sbjct: 347 AAYKWIADGILNGPYANPP--FDRSCLR 372
>At1g28580 lipase like protein
Length = 390
Score = 171 bits (433), Expect = 6e-43
Identities = 120/382 (31%), Positives = 184/382 (47%), Gaps = 31/382 (8%)
Query: 3 LRPLFIAFFLSCTLCVNSVELKNSPPCA-FPAIYNFGDSNSDTGGISAAFEP--IP---- 55
L L + FLS + N + + C F +I +FGDS +DTG + +P +P
Sbjct: 9 LMKLLVFIFLSTFVVTN---VSSETKCREFKSIISFGDSIADTGNLLGLSDPKDLPHMAF 65
Query: 56 PPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRR 115
PPYGE+F P+ R +GRLI+DFIAE L LP + + S N+ G NFA GG+T
Sbjct: 66 PPYGENFFHHPTGRFSNGRLIIDFIAEFLGLPLVPPFYGSHNANFEKGVNFAVGGATALE 125
Query: 116 QN---ETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYT 172
++ + + + SL +Q FK+ + + +E + + + E
Sbjct: 126 RSFLEDRGIHFPYTNVSLGVQLNSFKESLPSICGSPSDCRDMIENALILMGE-------- 177
Query: 173 FDIGQNDLSVGFRM-MNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPV 231
IG ND + F + ++++E MP ++ ++SA+ + +GGRTF + P+GC +
Sbjct: 178 --IGGNDYNYAFFVDKGIEEIKELMPLVITTISSAITELIGMGGRTFLVPGEFPVGCSVL 235
Query: 232 NLFYKHNLPAGYLDPY-GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKY 290
L DP GC+K N +QL+ + +L+ P I Y D Y A +
Sbjct: 236 YLTSHQTSNMEEYDPLTGCLKWLNKFGENHGEQLRAELNRLQKLYPHVNIIYADYYNALF 295
Query: 291 GLISNTKNEGFVD-PLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVH 349
L GF++ PL CCG + T+G G D+ +C+ PS YV+WDGVH
Sbjct: 296 HLYQEPAKFGFMNRPLSACCGAGGPYNY----TVGRKCGTDIV-ESCDDPSKYVAWDGVH 350
Query: 350 YAEAANHWVANRILNGSFTDPP 371
EAA +A ILNG + PP
Sbjct: 351 MTEAAYRLMAEGILNGPYAIPP 372
>At1g28570 lipase like protein
Length = 389
Score = 163 bits (413), Expect = 1e-40
Identities = 119/377 (31%), Positives = 178/377 (46%), Gaps = 34/377 (9%)
Query: 10 FFLSCTLCVNSVELKNSPPCA-FPAIYNFGDSNSDTGGISAAFEP--IPP----PYGESF 62
F + T C+ +V + P C F +I +FGDS +DTG + A +P +P PYGE+F
Sbjct: 12 FLILSTFCLTTVN--SEPQCHNFKSIISFGDSIADTGNLLALSDPTNLPKVAFLPYGETF 69
Query: 63 PQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQN---ET 119
P+ R +GRLI+DFIAE L P + + S N+ G NFA GG+T ++ E
Sbjct: 70 FHHPTGRFSNGRLIIDFIAEFLGFPLVPPFYGSQNANFEKGVNFAVGGATALERSFLEER 129
Query: 120 IFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQND 179
+ + SL +Q FK+ + + +E S + + E IG ND
Sbjct: 130 GIHFPYTNVSLAVQLSSFKESLPNLCVSPSDCRDMIENSLILMGE----------IGGND 179
Query: 180 LSVGFRM-MNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHN 238
+ F + N ++++E +P ++ ++SA+ + +GG+TF + P+GC L
Sbjct: 180 YNYAFFVGKNIEEIKELVPLVIETISSAITELIGMGGKTFLVPGEFPLGCSVAYLSLYQT 239
Query: 239 LPAGYLDPY-GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTK 297
DP GC+K N + ++QL+ + +L+ P I Y D Y L
Sbjct: 240 SNIEEYDPLTGCLKWLNKFSEYHDEQLQAELNRLQKLYPHVNIIYADYYNTLLRLAQEPA 299
Query: 298 NEGFVD-PLKICC--GYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAA 354
GF+ PL CC G N T+G G V C+ PS YVSWDGVH EAA
Sbjct: 300 KFGFISRPLPACCALGGPFN------FTLGRKRGTQV-PECCDDPSKYVSWDGVHMTEAA 352
Query: 355 NHWVANRILNGSFTDPP 371
+A IL G + PP
Sbjct: 353 YRLMAEGILKGPYAIPP 369
>At2g27360 putative lipase
Length = 394
Score = 163 bits (412), Expect = 2e-40
Identities = 122/386 (31%), Positives = 179/386 (45%), Gaps = 44/386 (11%)
Query: 6 LFIAFFLSC---TLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEP--IP----P 56
+ ++FF+S T+ + +N F +I +FGDS +DTG + P +P P
Sbjct: 8 MLLSFFISTFLITVVTSQTRCRN-----FKSIISFGDSITDTGNLLGLSSPNDLPESAFP 62
Query: 57 PYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRR- 115
PYGE+F PS R DGRLI+DFIAE L +P++ + S N+ G NFA GG+T
Sbjct: 63 PYGETFFHHPSGRFSDGRLIIDFIAEFLGIPHVPPFYGSKNGNFEKGVNFAVGGATALEC 122
Query: 116 --QNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF 173
E S SL Q FK E+ L S P + + +
Sbjct: 123 SVLEEKGTHCSQSNISLGNQLKSFK-----------ESLPYLCGSSSPDCRDMIENAFIL 171
Query: 174 --DIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP 230
+IG ND + F N ++++E +P ++ ++SA+ + ++G RTF + P+GC
Sbjct: 172 IGEIGGNDYNFPLFDRKNIEEVKELVPLVITTISSAISELVDMGARTFLVPGNFPLGCSV 231
Query: 231 VNL-FYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAK 289
L Y+ Y GC+ N +V N+QL+ + +LR P I Y D Y
Sbjct: 232 AYLTLYETPNKEEYNPLTGCLTWLNDFSVYHNEQLQAELKRLRNLYPHVNIIYGDYYNTL 291
Query: 290 YGLISNTKNEGFVD-PLKICCGY--HVNDT-HIWCGTIGTANGKDVFGNACEKPSMYVSW 345
L+ G +D PL CCG N T I CG+ G C PS YV+W
Sbjct: 292 LRLMQEPSKFGLMDRPLPACCGLGGPYNFTFSIKCGSKGV--------EYCSDPSKYVNW 343
Query: 346 DGVHYAEAANHWVANRILNGSFTDPP 371
DG+H EAA W++ +L G + PP
Sbjct: 344 DGIHMTEAAYKWISEGVLTGPYAIPP 369
>At1g31550 unknown protein
Length = 394
Score = 163 bits (412), Expect = 2e-40
Identities = 120/384 (31%), Positives = 182/384 (47%), Gaps = 45/384 (11%)
Query: 3 LRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEP--IP----P 56
L LF++ S + C N F +I +FGDS +DTG + + +P P
Sbjct: 17 LSTLFVSIVSSESQCRN-----------FESIISFGDSIADTGNLLGLSDHNNLPMSAFP 65
Query: 57 PYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQ 116
PYGE+F P+ R DGRLI+DFIAE L LPY+ Y S N+ G NFA +T
Sbjct: 66 PYGETFFHHPTGRFSDGRLIIDFIAEFLGLPYVPPYFGSTNGNFEKGVNFAVASATALES 125
Query: 117 N--ETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFD 174
+ E + FSL +Q FKQ L + + + + + + E
Sbjct: 126 SFLEEKGYHCPHNFSLGVQLKIFKQSLPNLCGLPSDCRDMIGNALILMGE---------- 175
Query: 175 IGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL 233
IG ND + F++ D+++E +P +++ ++SA+ + +GGRTF + P+GC L
Sbjct: 176 IGANDYNFPFFQLRPLDEVKELVPLVISTISSAITELIGMGGRTFLVPGGFPLGCSVAFL 235
Query: 234 FYKHNLPAGYLDPY-GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGL 292
DP GC+K N ++QL++ + +LR P I Y D Y A L
Sbjct: 236 TLHQTSNMEEYDPLTGCLKWLNKFGEYHSEQLQEELNRLRKLNPHVNIIYADYYNASLRL 295
Query: 293 ISNTKNEGFVD-PLKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDG 347
GF++ L CCG Y+ N + CG++G AC PS YV+WDG
Sbjct: 296 GREPSKYGFINRHLSACCGVGGPYNFNLSRS-CGSVGV--------EACSDPSKYVAWDG 346
Query: 348 VHYAEAANHWVANRILNGSFTDPP 371
+H EAA+ +A+ ++ G + PP
Sbjct: 347 LHMTEAAHKSMADGLVKGPYAIPP 370
>At5g45910 GDSL-motif lipase/hydrolase-like protein
Length = 372
Score = 162 bits (411), Expect = 2e-40
Identities = 126/395 (31%), Positives = 191/395 (47%), Gaps = 51/395 (12%)
Query: 1 MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPP---- 56
M + LFI F S + V S+ ++ P + +I+NFGDS SDTG + + P
Sbjct: 1 MRINMLFIVAF-SFLVSVRSLPMR--PTLKYESIFNFGDSLSDTGNFLLSGDVDSPNIGR 57
Query: 57 -PYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTN----YRHGANFATGGS 111
PYG++F + + R DGRLI+DFIAE LPY+ YL SL TN ++ GANFA G+
Sbjct: 58 LPYGQTFFNRSTGRCSDGRLIIDFIAEASGLPYIPPYLQSLRTNDSVDFKRGANFAVAGA 117
Query: 112 TIRR----QNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFS 167
T +N + ++ +LD+Q FK+ K +L ++K + F
Sbjct: 118 TANEFSFFKNRGLSVTLLTNKTLDIQLDWFKKL-----------KPSLCKTKPECEQYFR 166
Query: 168 KALYTF-DIGQNDLS---VGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNT 223
K+L+ +IG ND + + FR +F + +P ++N++ + E G T +
Sbjct: 167 KSLFLVGEIGGNDYNYPLLAFR--SFKHAMDLVPFVINKIMDVTSALIEEGAMTLIVPGN 224
Query: 224 APIGCLPVNLFYKHNLPAGYL--DPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAIT 281
PIGC L + N +G+L C N +A N +LK + LR + P A I
Sbjct: 225 LPIGC-SAALLERFNDNSGWLYDSRNQCYMPLNNLAKLHNDKLKKGLAALRKKYPYAKII 283
Query: 282 YVDLYAAKYGLISNTKNEGFV-DPLKICCG-----YHVNDTHIWCGTIGTANGKDVFGNA 335
Y D Y++ ++ GF LK CCG Y+V ++ CG G+
Sbjct: 284 YADYYSSAMQFFNSPSKYGFTGSVLKACCGGGDGRYNV-QPNVRCGEKGS--------TT 334
Query: 336 CEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDP 370
CE PS Y +WDG+H EAA +A +++G FT P
Sbjct: 335 CEDPSTYANWDGIHLTEAAYRHIATGLISGRFTMP 369
>At1g28610 lipase, putative
Length = 383
Score = 161 bits (407), Expect = 6e-40
Identities = 117/380 (30%), Positives = 184/380 (47%), Gaps = 40/380 (10%)
Query: 8 IAFFLSC---TLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFE----PIPP--PY 58
++FFLS T+ + + +N +I +FGDS +DTG + + P+ PY
Sbjct: 8 VSFFLSTLFVTIVSSQTQCRN-----LESIISFGDSITDTGNLVGLSDRNHLPVTAFLPY 62
Query: 59 GESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNE 118
GE+F P+ R C+GR+I+DFIAE L LP++ + S N+ G NFA G+T +
Sbjct: 63 GETFFHHPTGRSCNGRIIIDFIAEFLGLPHVPPFYGSKNGNFEKGVNFAVAGAT-ALETS 121
Query: 119 TIFQYGI-SPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF-DIG 176
+ + GI P S +Q K FK E+ L S + A +IG
Sbjct: 122 ILEKRGIYYPHSNISLGIQLKTFK--------ESLPNLCGSPTDCRDMIGNAFIIMGEIG 173
Query: 177 QNDLSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL-FY 235
ND + F + +++E +P ++ +++SA+ + ++GGRTF + P+GC L Y
Sbjct: 174 GNDFNFAFFVNKTSEVKELVPLVITKISSAIVELVDMGGRTFLVPGNFPLGCSATYLTLY 233
Query: 236 KHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISN 295
+ + Y GC+ N + +N++L+ + +L P I Y D + A L
Sbjct: 234 QTSNKEEYDPLTGCLTWLNDFSEYYNEKLQAELNRLSKLYPHVNIIYGDYFNALLRLYQE 293
Query: 296 TKNEGFVD-PLKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHY 350
GF+D PL CCG Y+ + CG++G C PS YV+WDGVH
Sbjct: 294 PSKFGFMDRPLPACCGLGGPYNFTLSK-KCGSVGV--------KYCSDPSKYVNWDGVHM 344
Query: 351 AEAANHWVANRILNGSFTDP 370
EAA W+A+ +L G +T P
Sbjct: 345 TEAAYKWIADGLLKGPYTIP 364
>At1g28650 lipase, putative
Length = 385
Score = 159 bits (401), Expect = 3e-39
Identities = 124/379 (32%), Positives = 176/379 (45%), Gaps = 31/379 (8%)
Query: 6 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGI----------SAAFEPIP 55
L ++F L V S + +I +FGDS +DTG AAF P
Sbjct: 10 LIVSFLLILYYTTIVVASSESRCRRYKSIISFGDSIADTGNYVHLSNVNNLPQAAFLP-- 67
Query: 56 PPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRR 115
YGESF PS R DGRL++DFIAE L LPY+ Y S ++ G NFA G+T
Sbjct: 68 --YGESFFHPPSGRYSDGRLVIDFIAEFLGLPYVPPYFGSQNVSFNQGINFAVYGATALD 125
Query: 116 QNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF-D 174
+ + Q S F+ VQ FK Q S E +L +
Sbjct: 126 RAFLVKQGIKSDFTNISLSVQLNTFK-------QILPNLCASSTRDCREMLGDSLILMGE 178
Query: 175 IGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL 233
IG ND + F + ++++E +P I+ ++SA+ ++ +LGG+TF + PIGC L
Sbjct: 179 IGGNDYNYPFFEGKSINEIKELVPLIIKAISSAIVDLIDLGGKTFLVPGNFPIGCSTAYL 238
Query: 234 FYKHNLPAGYLDPY-GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGL 292
+ DP+ GC+ N N+QLK + +L+ P I Y D Y + YGL
Sbjct: 239 TLFQTATVEH-DPFTGCIPWLNKFGEHHNEQLKIELKQLQKLYPHVNIIYADYYNSLYGL 297
Query: 293 ISNTKNEGFVD-PLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYA 351
GF + PL CCG V + + TIG G++ + C+ PS YV+WDG H
Sbjct: 298 FQEPAKYGFKNRPLAACCG--VGGQYNF--TIGKECGENGV-SYCQNPSEYVNWDGYHLT 352
Query: 352 EAANHWVANRILNGSFTDP 370
EA +A +LNG +T P
Sbjct: 353 EATYQKMAQGLLNGRYTTP 371
>At1g28590 putative lipase (At1g28590)
Length = 403
Score = 157 bits (396), Expect = 1e-38
Identities = 118/378 (31%), Positives = 178/378 (46%), Gaps = 38/378 (10%)
Query: 11 FLSCTLCVNSVELKNSPPCA-FPAIYNFGDSNSDTGGISAAFEP--IP----PPYGESFP 63
F+ TL V SV + C F +I +FGDS +DTG + +P +P PPYGE+F
Sbjct: 15 FILSTLLVTSVNSQTQ--CRNFKSIISFGDSIADTGNLLGLSDPNDLPASAFPPYGETFF 72
Query: 64 QKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQN---ETI 120
P+ R DGRLI+DFIAE L P + + N++ G NFA G+T + E
Sbjct: 73 HHPTGRYSDGRLIIDFIAEFLGFPLVPPFYGCQNANFKKGVNFAVAGATALEPSFLEERG 132
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDL 180
I+ SL +Q F + + + +E + + + E IG ND
Sbjct: 133 IHSTITNVSLSVQLRSFTESLPNLCGSPSDCRDMIENALILMGE----------IGGNDY 182
Query: 181 SVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL-FYKHN 238
+ F+ ++ E +P ++ ++SA+ + +GGRTF + PIG L YK +
Sbjct: 183 NFALFQRKPVKEVEELVPFVIATISSAITELVCMGGRTFLVPGNFPIGYSASYLTLYKTS 242
Query: 239 LPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKN 298
Y GC+K N + +NKQL++ + LR P I Y D Y A L
Sbjct: 243 NKEEYDPLTGCLKWLNDFSEYYNKQLQEELNGLRKLYPHVNIIYADYYNALLRLFQEPAK 302
Query: 299 EGFVD-PLKICCG----YHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEA 353
GF++ PL CCG Y+ N + CG++G C+ PS YV++DG+H EA
Sbjct: 303 FGFMNRPLPACCGVGGSYNFNFSR-RCGSVGV--------EYCDDPSQYVNYDGIHMTEA 353
Query: 354 ANHWVANRILNGSFTDPP 371
A ++ +L G + PP
Sbjct: 354 AYRLISEGLLKGPYAIPP 371
>At1g28640 lipase, putative
Length = 390
Score = 154 bits (388), Expect = 1e-37
Identities = 120/374 (32%), Positives = 179/374 (47%), Gaps = 25/374 (6%)
Query: 7 FIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGIS--AAFEPIPP----PYGE 60
F+ S T+ V S E + F +I +FGDS +DTG + +P PYGE
Sbjct: 12 FLLVLYSTTIIVASSESRCR---RFKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGE 68
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
SF PS R DGRLI+DFIAE L LPY+ +Y S ++ G NFA G+T + +
Sbjct: 69 SFFHPPSGRYSDGRLIIDFIAEFLGLPYVPSYFGSQNVSFDQGINFAVYGATALDRVFLV 128
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF-DIGQND 179
+ S F+ VQ F KQ+ T+ R E +L +IG ND
Sbjct: 129 GKGIESDFTNVSLSVQLNIF----KQILPNLCTSSSRD---CREMLGDSLILMGEIGVND 181
Query: 180 LSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHN 238
+ F + +++++ +P ++ ++SA+ ++ +LGG+TF + P+GC P L
Sbjct: 182 YNYPFFEGKSINEIKQLVPLVIKAISSAIVDLIDLGGKTFLVPGNFPLGCYPAYLTLFQT 241
Query: 239 LPAGYLDPY-GCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTK 297
DP+ GC+ N N+QLK + +L+ I Y D Y + + L
Sbjct: 242 AAEEDHDPFTGCIPRLNEFGEYHNEQLKTELKRLQELYDHVNIIYADYYNSLFRLYQEPV 301
Query: 298 NEGFVD-PLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANH 356
GF + PL CCG V + + TIG G + C+ PS YV+WDG H EA +
Sbjct: 302 KYGFKNRPLAACCG--VGGQYNF--TIGKECGHRGV-SCCQNPSEYVNWDGYHLTEATHQ 356
Query: 357 WVANRILNGSFTDP 370
+A ILNG++ P
Sbjct: 357 KMAQVILNGTYASP 370
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,252,147
Number of Sequences: 26719
Number of extensions: 415431
Number of successful extensions: 1451
Number of sequences better than 10.0: 111
Number of HSP's better than 10.0 without gapping: 101
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1007
Number of HSP's gapped (non-prelim): 121
length of query: 381
length of database: 11,318,596
effective HSP length: 101
effective length of query: 280
effective length of database: 8,619,977
effective search space: 2413593560
effective search space used: 2413593560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0001.6


