

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
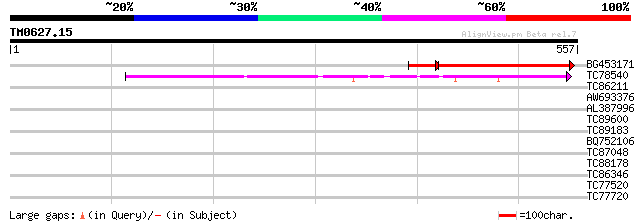
Score E
Sequences producing significant alignments: (bits) Value
BG453171 similar to GP|16604326|gb At4g36400/C7A10_960 {Arabidop... 242 4e-74
TC78540 similar to GP|15010680|gb|AAK73999.1 AT5g06580/F15M7_11 ... 150 9e-37
TC86211 homologue to GP|13194621|gb|AAK15493.1 brassinosteroid b... 32 0.49
AW693376 weakly similar to GP|21536978|gb| unknown {Arabidopsis ... 32 0.64
AL387996 31 1.4
TC89600 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {... 30 1.9
TC89183 weakly similar to PIR|F70591|F70591 probable kefB protei... 30 2.4
BQ752106 similar to GP|172327|gb|AA DNA helicase {Saccharomyces ... 30 3.2
TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsi... 30 3.2
TC88178 weakly similar to GP|10176891|dbj|BAB10121. berberine br... 29 4.2
TC86346 homologue to GP|22136752|gb|AAM91695.1 unknown protein {... 29 4.2
TC77520 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MS... 29 5.4
TC77720 similar to GP|18176302|gb|AAL60019.1 putative berberine ... 28 7.1
>BG453171 similar to GP|16604326|gb At4g36400/C7A10_960 {Arabidopsis
thaliana}, partial (35%)
Length = 594
Score = 242 bits (618), Expect(2) = 4e-74
Identities = 115/134 (85%), Positives = 128/134 (94%)
Frame = +3
Query: 422 EGITEALMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNAANVVGYGHLGDGNLHLNISVP 481
EGI+EALM+ GAVYKYD+SIP+ENLYNLVEEMRSRLG+AANV+GYGHLGDGNLHLN+SV
Sbjct: 93 EGISEALMKAGAVYKYDVSIPVENLYNLVEEMRSRLGDAANVIGYGHLGDGNLHLNVSVS 272
Query: 482 QYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDP 541
QYD+KILSQIEPFVYEWTSK GSISAEHG+GLMKANEI YS+S ETVQ+MASIKNL+DP
Sbjct: 273 QYDEKILSQIEPFVYEWTSKKRGSISAEHGVGLMKANEIFYSQSHETVQMMASIKNLMDP 452
Query: 542 NHILNPYKVLPHSL 555
NHILNPYKVLPHS+
Sbjct: 453 NHILNPYKVLPHSI 494
Score = 54.7 bits (130), Expect(2) = 4e-74
Identities = 26/32 (81%), Positives = 29/32 (90%)
Frame = +2
Query: 392 FLLHSMENELISDGVLAQDINQASSFWRLREG 423
FLL SMENELI+DGVLAQDINQAS+FWR+ G
Sbjct: 2 FLLGSMENELIADGVLAQDINQASTFWRIX*G 97
>TC78540 similar to GP|15010680|gb|AAK73999.1 AT5g06580/F15M7_11
{Arabidopsis thaliana}, partial (77%)
Length = 2163
Score = 150 bits (380), Expect = 9e-37
Identities = 123/453 (27%), Positives = 213/453 (46%), Gaps = 14/453 (3%)
Frame = +1
Query: 114 DEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSV 173
DE + + HK +++ P + ++VS+I+K CN+ + +VP GG T + G ++
Sbjct: 427 DERYIHGKPQNSFHKAVNIPDVIVYPRSEEEVSKIVKSCNNHKIPIVPYGGATSIEGHTL 606
Query: 174 PVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQI 233
V + +SSM +V + +V + G + +L+ G PLD G S I
Sbjct: 607 SPNGGVCIDMSSMKRVKALHVDDMDVVVDPGIGWMELNEYLEPYGLFFPLDPGPGAS--I 780
Query: 234 GGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG 293
GG +T G VRYG++ +V+ ++ V A+G ++ RK GYDL L IGSEG
Sbjct: 781 GGMCATRCSGSLAVRYGTMRDNVISLKVVFANGDIVKTASRARKSAAGYDLTRLVIGSEG 960
Query: 294 SLGIVTKVAILTPPKLSSV-NVALLACKDYNCCQKLLQEAKRKL--GEILSAFEFLDSQS 350
+LG++T+V + +L + +++A ++ + A + G +S E LD
Sbjct: 961 TLGVITEVTL----RLQKIPQFSVVAMCNFPSVKDAADVAITTMNSGIQVSRVELLDEVQ 1128
Query: 351 MNLVINHLEGARNP-LPTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQ 409
+ IN G P PTL + E G++ + +Q + S N SD V A+
Sbjct: 1129VK-AINIANGKNLPESPTL-----MFEFIGTEAYAREQTQIVRKIVSEHNG--SDFVFAE 1284
Query: 410 DINQASSFWRLREGITEALMRVGAVYK------YDLSIPLENLYNLVEEMRSRLGNAANV 463
+ W++R+ EAL A+ D+ +PL +L +++ + + L + V
Sbjct: 1285EPEAKKELWKVRK---EALWACFAMEPNMEAMISDVCVPLSHLADIISKSKKELDASPLV 1455
Query: 464 -VGYGHLGDGNLHLNI---SVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANE 519
H GDGN H I + + ++ F+ G+ + EHG+G K
Sbjct: 1456CTVIAHAGDGNFHTVILFDPAKEEQRREAERLNHFMVHAALSLEGTCTGEHGVGTGKMKY 1635
Query: 520 ILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
+ +E ++ M IK++LDPN+I+NP K++P
Sbjct: 1636LEEELGVEALKTMKKIKSVLDPNNIMNPGKLIP 1734
>TC86211 homologue to GP|13194621|gb|AAK15493.1 brassinosteroid biosynthetic
protein LKB {Pisum sativum}, partial (60%)
Length = 1141
Score = 32.3 bits (72), Expect = 0.49
Identities = 21/60 (35%), Positives = 30/60 (50%)
Frame = +1
Query: 248 RYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPP 307
+YG +V+ E VLADG+ L KDN DL + S+G+LG++ I P
Sbjct: 616 KYGLFSDTVVAFEIVLADGS----LVRATKDNEYSDLFYAIPWSQGTLGLLVAAEIKLIP 783
>AW693376 weakly similar to GP|21536978|gb| unknown {Arabidopsis thaliana},
partial (27%)
Length = 662
Score = 32.0 bits (71), Expect = 0.64
Identities = 21/58 (36%), Positives = 30/58 (51%)
Frame = +3
Query: 312 VNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLH 369
+NV+ Y+ C+KLL+ K E S+ E L S + I +LEG PLP L+
Sbjct: 114 LNVSTRKLSLYDECEKLLECRLLKSDENCSSGESLTFNSHLVDIGNLEGENKPLPDLN 287
>AL387996
Length = 378
Score = 30.8 bits (68), Expect = 1.4
Identities = 16/35 (45%), Positives = 20/35 (56%)
Frame = -1
Query: 95 NDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKY 129
NDD +RY EI+G KN V +EE V W K+
Sbjct: 339 NDDQIRYI-EIIGGKNYVYNEEIRKEVKRVWGWKF 238
>TC89600 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {Arabidopsis
thaliana}, partial (29%)
Length = 700
Score = 30.4 bits (67), Expect = 1.9
Identities = 21/92 (22%), Positives = 42/92 (44%)
Frame = +2
Query: 179 VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVS 238
++V L + + IS D + ++G + + + + ++ G S +GG+++
Sbjct: 395 IVVDLIKL-RGISIDTKTNTAWVQSGATIGEVYYRIYEKSSVLGYPAGLCTSLGVGGHIT 571
Query: 239 TNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD 270
A G + +YG +VL V A G +LD
Sbjct: 572 GGAYGTLMRKYGLGVDNVLNARIVDAKGNILD 667
>TC89183 weakly similar to PIR|F70591|F70591 probable kefB protein -
Mycobacterium tuberculosis (strain H37RV), partial (7%)
Length = 1137
Score = 30.0 bits (66), Expect = 2.4
Identities = 18/40 (45%), Positives = 24/40 (60%), Gaps = 1/40 (2%)
Frame = -2
Query: 225 LGAKGSCQIGGNVS-TNAGGLRLVRYGSLHGSVLGVEAVL 263
LGA+ +IGGNV GGLRL+++G L G+ VL
Sbjct: 197 LGARNRMRIGGNVRYVVDGGLRLMQHGGLGNEDRGL*CVL 78
>BQ752106 similar to GP|172327|gb|AA DNA helicase {Saccharomyces cerevisiae},
partial (9%)
Length = 495
Score = 29.6 bits (65), Expect = 3.2
Identities = 18/69 (26%), Positives = 37/69 (53%), Gaps = 8/69 (11%)
Frame = -3
Query: 389 LEAFLLHSMENE--------LISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLS 440
+E F++ ++ +E L + V+ +D+N+A FW+++ +T+ + G DL
Sbjct: 478 VETFVVSALRSENATKTLGLLTTRTVVRRDLNEARGFWQVKGRVTDLAKKQGV----DLR 311
Query: 441 IPLENLYNL 449
+ LE L N+
Sbjct: 310 VVLEVL*NV 284
>TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsis thaliana},
partial (77%)
Length = 1571
Score = 29.6 bits (65), Expect = 3.2
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +1
Query: 336 LGEILSAFEFLDSQSMNLVINHLEGARNP 364
+G+I+ A E+L SQS N+ H G R+P
Sbjct: 1264 IGDIVVALEYLASQSQNIPEVHRHGVRSP 1350
>TC88178 weakly similar to GP|10176891|dbj|BAB10121. berberine bridge enzyme
{Arabidopsis thaliana}, partial (56%)
Length = 1629
Score = 29.3 bits (64), Expect = 4.2
Identities = 24/114 (21%), Positives = 51/114 (44%), Gaps = 3/114 (2%)
Frame = +1
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG L + + N+ + G+ + +GG+ S G +YG +++ +
Sbjct: 466 QAGATLGELYYAIANKSNLHGFPAGSCPTVGVGGHFSGGGFGTIFRKYGLATDNIIDAQI 645
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG-SLGIVT--KVAILTPPKLSSV 312
+ +G +L+ ++ G DL G G S G++T KV ++ P + ++
Sbjct: 646 IDVNGNILN------REMMGEDLFWAIRGGGGSSFGVITAWKVKLVRVPLIVTI 789
>TC86346 homologue to GP|22136752|gb|AAM91695.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 2436
Score = 29.3 bits (64), Expect = 4.2
Identities = 11/32 (34%), Positives = 25/32 (77%)
Frame = +3
Query: 107 GQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQ 138
GQKNV +++E++ +++++++ Y G + LLL+
Sbjct: 2118 GQKNVHENKEEIEEMDSEYIYIYIGKATLLLR 2213
>TC77520 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MSL3_100
{Arabidopsis thaliana}, partial (27%)
Length = 1880
Score = 28.9 bits (63), Expect = 5.4
Identities = 19/71 (26%), Positives = 33/71 (45%), Gaps = 8/71 (11%)
Frame = -3
Query: 125 WMHKYEGSSKLLLQPWNTDQVSQILKYCNSRS--------LAVVPQGGNTGLVGGSVPVF 176
W+ + SS +L+ PW++ ++ +SRS L +VP T V
Sbjct: 1509 WLTSSDFSSSILITPWHSISIASPGILASSRSARELTSSPLKIVPSVKATPTAATKVATA 1330
Query: 177 DEVIVSLSSMN 187
EVI+S+ S++
Sbjct: 1329 PEVIISVISVS 1297
>TC77720 similar to GP|18176302|gb|AAL60019.1 putative berberine bridge
enzyme {Arabidopsis thaliana}, partial (38%)
Length = 804
Score = 28.5 bits (62), Expect = 7.1
Identities = 28/114 (24%), Positives = 49/114 (42%), Gaps = 3/114 (2%)
Frame = +3
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG + + + + + G S IGG ++ A G + +YG +V+
Sbjct: 447 QAGATIGEVYYKISAKSSVHGFPAGLCTSLGIGGLITGGAYGSMMRKYGLGVDNVVDARI 626
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG-SLGIVT--KVAILTPPKLSSV 312
V A+G +LD + G DL G G S G++ K+ ++ PK +V
Sbjct: 627 VDANGKILD------RKAMGEDLFWAIRGGGGSSFGVILWWKIKLVPVPKTVTV 770
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,870,812
Number of Sequences: 36976
Number of extensions: 228940
Number of successful extensions: 1095
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 1089
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1093
length of query: 557
length of database: 9,014,727
effective HSP length: 101
effective length of query: 456
effective length of database: 5,280,151
effective search space: 2407748856
effective search space used: 2407748856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0627.15


