

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0593a.3
(562 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
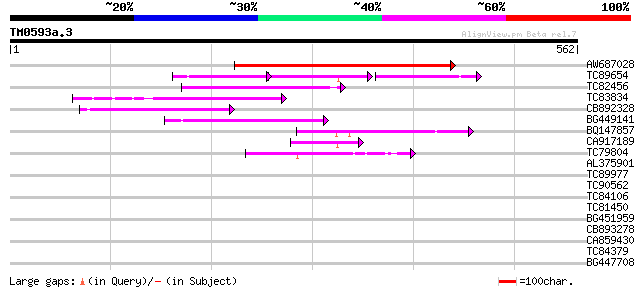
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 227 8e-60
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 76 4e-38
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 116 3e-26
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 97 2e-20
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 92 7e-19
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 91 2e-18
BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {A... 75 9e-14
CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.3... 55 9e-08
TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like... 43 3e-04
AL375901 similar to GP|9759134|dbj mutator-like transposase-like... 33 0.22
TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired re... 32 0.50
TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 32 0.50
TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity ... 32 0.65
TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20... 32 0.85
BG451959 homologue to GP|6175165|gb| Mutator-like transposase {A... 31 1.4
CB893278 weakly similar to PIR|T05645|T056 hypothetical protein ... 31 1.4
CA859430 similar to GP|8131699|dbj cytoplasmic ribosomal protein... 30 1.9
TC84379 similar to GP|18481712|gb|AAL73534.1 putative nucleolar ... 30 2.5
BG447708 28 9.4
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 227 bits (579), Expect = 8e-60
Identities = 112/220 (50%), Positives = 146/220 (65%), Gaps = 1/220 (0%)
Frame = +1
Query: 224 DVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGK 283
DV+ F +TYKKNKYN P+ FSG NHH++T IF AL+ DETIE+YKWVL E M K
Sbjct: 1 DVLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENK 180
Query: 284 QPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKIPDFLNEFKTLIYGNFTP 343
PKAVV DGD SMR A+K VFP+A+HRLC WH+ +N E IK FL F+ +Y NFTP
Sbjct: 181 FPKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQENIKKTPFLEGFRKAMYSNFTP 360
Query: 344 ERFETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKS 403
E+FE W ++I+K + W+ + Y + +WATA++R+ FF IRTTS CE IN+ +K+
Sbjct: 361 EQFEDFWSELIQKNELEGNAWVIKTYANKSLWATAYLRDKFFGRIRTTSQCEAINAIVKT 540
Query: 404 YVLCKNSILDFIYNF-ERAVEEYRHNELASDFKSRYGEPV 442
Y + F R E YR+NEL +DFKS++ EPV
Sbjct: 541 YSRSQRQNF*IYA*FLNRF*EGYRNNELVADFKSKFTEPV 660
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 75.9 bits (185), Expect(3) = 4e-38
Identities = 34/98 (34%), Positives = 54/98 (54%)
Frame = +2
Query: 162 HVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQA 221
H+ ++ +LG G L YL+ ++ FF+ + N+FW D T R +Y
Sbjct: 83 HMSSTRQRTLGGGGH-QVLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSY 259
Query: 222 FGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACA 259
FGD ++F +TYK N+Y P +F+G+NHH + +F CA
Sbjct: 260 FGDTVIFDTTYKTNQYRVPFASFTGFNHHGQPVLFGCA 373
Score = 62.8 bits (151), Expect(3) = 4e-38
Identities = 34/108 (31%), Positives = 55/108 (50%), Gaps = 5/108 (4%)
Frame = +3
Query: 257 ACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHI 316
A L+ +E+ +Y W+ K A+ G+ P ++ D D ++ AV V P HR C W I
Sbjct: 366 AVLLILNESEPSYIWLFKTWLRAVSGRPPVSITTDLDPVIQVAVAQVLPPTRHRFCKWSI 545
Query: 317 QQNCLEKI-----KIPDFLNEFKTLIYGNFTPERFETKWLQVIEKYGI 359
+ K+ P F NEFK ++ + T + FE+ W ++E+Y I
Sbjct: 546 FRENRSKLAHLYQSNPTFDNEFKKCVHESVTIDEFESCWRSLLERYYI 689
Score = 58.5 bits (140), Expect(3) = 4e-38
Identities = 32/105 (30%), Positives = 53/105 (50%)
Frame = +1
Query: 363 KWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAV 422
+W + MY RQ W ++R +FF I E +N F +V ++ + +E+AV
Sbjct: 700 EWPQSMYNARQHWVPVYLRGSFFGEIPLNDGNECLNFFFDGHVNASTTLQLLVRQYEKAV 879
Query: 423 EEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMR 467
+ EL +DF++ PVL T S +E+ A+ L T MF + +
Sbjct: 880 STWHERELKADFETSNSNPVLRTP-SPMEKQAASLYTRKMFMKFQ 1011
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 116 bits (290), Expect = 3e-26
Identities = 54/163 (33%), Positives = 88/163 (53%)
Frame = +2
Query: 171 LGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYS 230
L GD + S+ +++ F+ + N++NLFW D R Y+ FG+VI +
Sbjct: 704 LWRGDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDT 883
Query: 231 TYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVI 290
TY +KY+ P+V F G NHH +T + CA++ + +T W+ E M G P ++
Sbjct: 884 TYLTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFTRWLECMHGHAPNGIIT 1063
Query: 291 DGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKIPDFLNEF 333
+ DK+M+ A++V FP A HR C WHI + K+P+ L ++
Sbjct: 1064EEDKAMKNAIEVAFPKARHRWCLWHIMK------KVPEMLGKY 1174
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 97.1 bits (240), Expect = 2e-20
Identities = 61/212 (28%), Positives = 99/212 (45%)
Frame = +3
Query: 63 IKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDK 122
+ + TR CPA R++ +S +W++ EHNH L A +H L D
Sbjct: 390 VNNLRKETRTGCPAMIRMKL-VESQRWRICEVTLEHNHVL--GAKIHKSIKKNSLPSSD- 557
Query: 123 AQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSY 182
A+ ++K+Y L+ G++++ +D +++L GD + ++
Sbjct: 558 AEGKTIKVYHA--------LVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGDTQAIYNF 713
Query: 183 LQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVV 242
L +P FF+ + E +L N W D SR F DVI F +TY NKY P+V
Sbjct: 714 LCRMQLTNPNFFYLMDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVNKYEIPLV 893
Query: 243 NFSGYNHHTETTIFACALVCDETIETYKWVLK 274
G NHH ++ + C L+ E IE+YKW+ +
Sbjct: 894 ALVGINHHGQSVLLGCGLLAGEIIESYKWLFR 989
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 91.7 bits (226), Expect = 7e-19
Identities = 54/155 (34%), Positives = 82/155 (52%), Gaps = 1/155 (0%)
Frame = +1
Query: 70 TRVSCPAKFRVRFHA-KSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSL 128
+R +C A VRF+ K WKV +HNH P + + S R +++ + ++ SL
Sbjct: 364 SRTNCLAM--VRFNVTKEGVWKVTKLILDHNHEFVPLQQRYLLRSMRNMSNLKEDRIKSL 537
Query: 129 KLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGE 188
+R N+ G L + GG + G + KD++N+V K + AGDA S L++LQ K
Sbjct: 538 VNDAIRVTNVGGYLGKEVGGVDKRGVMLKDMHNYVSTGKLKFIEAGDAQSLLNHLQSKQA 717
Query: 189 KDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFG 223
+D MF++ E L N+FW DG S +DY A G
Sbjct: 718 QDSMFYYSVQLDHESRLNNVFWRDGKSVVDYIATG 822
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 90.5 bits (223), Expect = 2e-18
Identities = 46/163 (28%), Positives = 82/163 (50%)
Frame = +3
Query: 154 FLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDG 213
F +KD+ N + +++ + + L + +KDP F F++T LEN+ W
Sbjct: 171 FTEKDVRNLLQSFRKLD-PEEETLDLLRMCRNIKDKDPNFKFEYTLDANNRLENIAWSYA 347
Query: 214 TSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVL 273
+S Y FGD +VF +T++ ++ P+ + G N++ F C L+ DET+ ++ W +
Sbjct: 348 SSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLLRDETVRSFSWAI 527
Query: 274 KALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHI 316
KA M GK P+ ++ D + ++ A+ P H C W I
Sbjct: 528 KAFLGFMNGKAPQTILTDQNICLKEALSAEMPMTKHAFCIWMI 656
>BQ147857 weakly similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (26%)
Length = 560
Score = 74.7 bits (182), Expect = 9e-14
Identities = 48/180 (26%), Positives = 82/180 (44%), Gaps = 5/180 (2%)
Frame = +1
Query: 285 PKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLE--KIKIPDFLNEFKT---LIYG 339
P+ ++ D + ++ A+ V P + H C WHI + + + +E+K +Y
Sbjct: 4 PQTLLTDHNTWLKEAIAVEMPESKHAFCIWHILSKFSDWXYLLLGSQYDEWKADFHRLYN 183
Query: 340 NFTPERFETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINS 399
E FE W Q+++KYG+ K I +Y R WA F+R FFAG+ + E IN
Sbjct: 184 LEMXEDFEKSWRQMVDKYGLHANKHIISLYSLRTFWALPFLRRYFFAGLTSXCQTESINV 363
Query: 400 FIKSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLT 459
FI+ ++ ++ F+ V ++ A R + + S IE A+ +LT
Sbjct: 364 FIQRFLSAQSQPXRFLQQVADIV-DFNDRAGAKQXMQRKMQKXCLKTGSPIESHAATILT 540
>CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (12%)
Length = 312
Score = 54.7 bits (130), Expect = 9e-08
Identities = 26/77 (33%), Positives = 42/77 (53%), Gaps = 5/77 (6%)
Frame = +1
Query: 279 AMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEK-----IKIPDFLNEF 333
AM G+ P ++ D D +++A+ VFP+ HR C WHI + C EK ++ P+F EF
Sbjct: 7 AMSGRPPLSITTDHDSVIQSAIMQVFPDTRHRFCKWHIFKQCQEKLSHIFLQFPNFEAEF 186
Query: 334 KTLIYGNFTPERFETKW 350
+ + + FE+ W
Sbjct: 187 HKCVNLTDSIDEFESCW 237
>TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like
transposase {Arabidopsis thaliana}, partial (57%)
Length = 2129
Score = 43.1 bits (100), Expect = 3e-04
Identities = 40/173 (23%), Positives = 73/173 (42%), Gaps = 4/173 (2%)
Frame = +1
Query: 234 KNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGK---QPKAVVI 290
K+KY ++ + ++ A A+V E E++ W L L A+ P+ + +
Sbjct: 784 KSKYLSTFLSATSFDADGGLFPLAFAVVDVENDESWTWFLSELHNALEVNTECMPQIIFL 963
Query: 291 -DGDKSMRAAVKVVFPNATHRLCGWHIQQNCLEKIKIPDFLNEFKTLIYGNFTPERFETK 349
DG K + A++ FP ++H C H+ +N ++ K ++ + Y T F K
Sbjct: 964 SDGQKGIVDAIRRKFPRSSHAFCMRHLSENIGKEFKNSRLIHLLWSAAYAT-TINAFREK 1140
Query: 350 WLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIK 402
+ IE+ W++ + ++ WA +F G R L I F K
Sbjct: 1141MAE-IEEVSPNASMWLQHFHPSQ--WALV-----YFEGTRYGHLSSNIEEFNK 1275
>AL375901 similar to GP|9759134|dbj mutator-like transposase-like protein
{Arabidopsis thaliana}, partial (27%)
Length = 493
Score = 33.5 bits (75), Expect = 0.22
Identities = 24/103 (23%), Positives = 46/103 (44%)
Frame = +3
Query: 304 FPNATHRLCGWHIQQNCLEKIKIPDFLNEFKTLIYGNFTPERFETKWLQVIEKYGIGEEK 363
FP+A+H C ++ +N + K +N F +Y T FE+K ++IE + ++
Sbjct: 9 FPSASHGFCLRYVSENFRDTFKNTKLVNIFWNAVYA-LTAAEFESKITEMIE---VSQDV 176
Query: 364 WIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVL 406
+ +WA A +F G+R G+ + ++ L
Sbjct: 177 ISWFQHFPPFLWAVA-----YFDGVRYGHFTLGVTELLYNWAL 290
>TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired response
protein {Arabidopsis thaliana}, partial (35%)
Length = 1523
Score = 32.3 bits (72), Expect = 0.50
Identities = 33/150 (22%), Positives = 64/150 (42%), Gaps = 5/150 (3%)
Frame = +2
Query: 386 AGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLIT 445
AG T E +NSF Y+ K ++ +F+ + + E +DF + + +P L
Sbjct: 2 AGTSTAQRSESMNSFFDKYIHKKITLKEFVKQYRLILLNRYDEEEIADFDTLHKQPAL-K 178
Query: 446 ALSHIEQGASKLLTLNMFREMRHMIQRALNLNLVERSEIGTTVMIKFSICRHPNTQ--LS 503
+ S E+ S + T +F++ + + R E+G +++ + + + L
Sbjct: 179 SPSPWEKQMSTIYTHAIFKKFQVEVLGVAGCQ--SRIEVGDGTAVRYIVQDYEKDEEFLV 352
Query: 504 SWSFMTRIRTC---LIVIVGFLNMLAYLVL 530
+W ++ +C L GFL A VL
Sbjct: 353 TWKELSSEVSCFCRLFEYKGFLCRHALSVL 442
>TC90562 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (66%)
Length = 1169
Score = 32.3 bits (72), Expect = 0.50
Identities = 17/43 (39%), Positives = 20/43 (45%)
Frame = +2
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTP 104
+I + TRV C A VR S W + F EH H LTP
Sbjct: 506 KIVRQRAETRVGCRAMIMVR-KLNSGLWSITKFVKEHTHPLTP 631
>TC84106 similar to GP|9502168|gb|AAF88018.1| contains simlarity to
Arabidopsis thaliana far-red impaired response protein
(GB:AAD51282.1), partial (25%)
Length = 673
Score = 32.0 bits (71), Expect = 0.65
Identities = 19/74 (25%), Positives = 35/74 (46%), Gaps = 2/74 (2%)
Frame = +2
Query: 54 MRSM*IISRIKEAKPVTRVSCPAKFRVRFHAKS--SKWKVVLFEPEHNHGLTPAAHVHFI 111
+R ++ ++ K V R C AK + S+W V+ F HNH L V +
Sbjct: 365 LRGRQMVMHHRDRKSV-RCGCDAKMYLSKDVVDGVSQWTVLQFSNVHNHELLEDDQVRLL 541
Query: 112 PSFRGLTDGDKAQV 125
P++R + + D+ ++
Sbjct: 542 PAYRKIHEADQERI 583
>TC81450 similar to GP|16604382|gb|AAL24197.1 AT3g59470/T16L24_20
{Arabidopsis thaliana}, partial (59%)
Length = 832
Score = 31.6 bits (70), Expect = 0.85
Identities = 16/45 (35%), Positives = 21/45 (46%)
Frame = +2
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAA 106
+I + TRV C A +R S KW + F EH H L P +
Sbjct: 221 KIVRQRAETRVGCRAMIMMR-KINSGKWVITKFVKEHTHPLNPGS 352
>BG451959 homologue to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (28%)
Length = 658
Score = 30.8 bits (68), Expect = 1.4
Identities = 22/92 (23%), Positives = 39/92 (41%), Gaps = 4/92 (4%)
Frame = +2
Query: 232 YKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALD---EAMFGKQPKAV 288
Y K+KY ++ +G++ A +V +E + + W L L E P+
Sbjct: 371 YLKSKYLGTLLLATGFDGDGALFPLAFGVVDEENDDNWMWFLSKLHNLLEINTENMPRLT 550
Query: 289 VI-DGDKSMRAAVKVVFPNATHRLCGWHIQQN 319
++ D + + V+ FP A H C H+ N
Sbjct: 551 ILSDRQQGIVDGVEANFPTAFHGFCMRHLSDN 646
>CB893278 weakly similar to PIR|T05645|T056 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (5%)
Length = 821
Score = 30.8 bits (68), Expect = 1.4
Identities = 19/61 (31%), Positives = 25/61 (40%)
Frame = +3
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGD 121
R+ P TR C A R+ + KW V F EH H L V + S + L D
Sbjct: 507 RVLPPPPATREGCQAMIRLALRDEG-KWVVTKFVKEHTHKLMSPGEVPWRGSGKHLVSED 683
Query: 122 K 122
+
Sbjct: 684 E 686
>CA859430 similar to GP|8131699|dbj cytoplasmic ribosomal protein S13 {Panax
ginseng}, complete
Length = 575
Score = 30.4 bits (67), Expect = 1.9
Identities = 15/55 (27%), Positives = 23/55 (41%)
Frame = -3
Query: 389 RTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVL 443
R T +C+G+ F +C + ++N E V Y ELA P+L
Sbjct: 216 RYTPICDGLMPFFAYLQICSQTSSGVVFNHEGGVRRYGKAELAIPLPGACIRPIL 52
>TC84379 similar to GP|18481712|gb|AAL73534.1 putative nucleolar protein
{Sorghum bicolor}, partial (31%)
Length = 666
Score = 30.0 bits (66), Expect = 2.5
Identities = 24/74 (32%), Positives = 35/74 (46%)
Frame = -1
Query: 385 FAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLI 444
F G+ E I+SF +S + C I N + +EEY N + S KSR+ P L
Sbjct: 657 FHGLLDVHSLEHIHSFXQSRLTC------LIMNQTKPLEEYPKNVVPSATKSRHRSPSLY 496
Query: 445 TALSHIEQGASKLL 458
S + Q A++ L
Sbjct: 495 FC-SIVLQAANRCL 457
>BG447708
Length = 147
Score = 28.1 bits (61), Expect = 9.4
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = +1
Query: 75 PAKFRVRFHAKSSKWKVVLFEPEHNHGLTP 104
P KF FH K K K L +P H G+ P
Sbjct: 13 PEKFXTPFHLKGGKPKKALGKPSHPAGVGP 102
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,387,585
Number of Sequences: 36976
Number of extensions: 264070
Number of successful extensions: 1683
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1673
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1680
length of query: 562
length of database: 9,014,727
effective HSP length: 101
effective length of query: 461
effective length of database: 5,280,151
effective search space: 2434149611
effective search space used: 2434149611
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0593a.3


