

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.8
(919 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
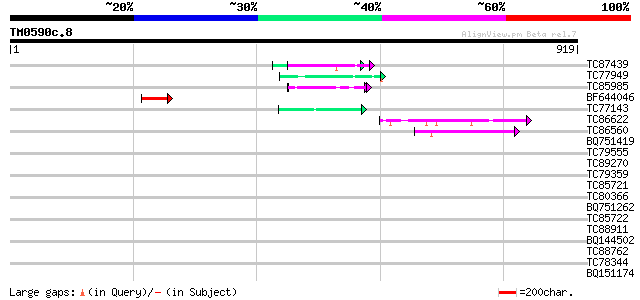
Sequences producing significant alignments: (bits) Value
TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in... 55 1e-07
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 53 5e-07
TC85985 similar to GP|5139695|dbj|BAA81686.1 expressed in cucumb... 47 3e-05
BF644046 45 1e-04
TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-pro... 44 2e-04
TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myo... 44 2e-04
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 43 6e-04
BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania maj... 42 0.001
TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishman... 41 0.002
TC89270 similar to GP|22773423|gb|AAL66352.2 catalase-peroxidase... 41 0.002
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 41 0.002
TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 40 0.003
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 40 0.003
BQ751262 similar to GP|6715100|gb|A versicolorin B synthase {Asp... 40 0.004
TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 40 0.004
TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2... 40 0.005
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 40 0.005
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 39 0.007
TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens cu... 39 0.009
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 39 0.012
>TC87439 weakly similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (57%)
Length = 897
Score = 55.1 bits (131), Expect = 1e-07
Identities = 42/144 (29%), Positives = 62/144 (42%), Gaps = 3/144 (2%)
Frame = +3
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAA--ATEPTAVPASSPASAG-ATAAFAAESSIGA 507
G P A PTT+ + P + VAA T T P +SP S+ AT+ AA +
Sbjct: 96 GGQSPSAAPTTSPSTPAATTPVSSPVAAPPTTPTTPAPVASPKSSPPATSPKAAAPTATP 275
Query: 508 TAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEK 567
AAS+ T + P+ K P S P P + P+T A P P+ P +G+K
Sbjct: 276 PAASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSPPAVTPVAAPTTTPAVPAPA-PSKGKK 452
Query: 568 SCPGAATTSEAAQIEQAPAPEVGS 591
+ + + + PAP G+
Sbjct: 453 NKKKHGAPAPSPALLGPPAPPAGA 524
Score = 44.7 bits (104), Expect = 2e-04
Identities = 51/167 (30%), Positives = 67/167 (39%), Gaps = 16/167 (9%)
Frame = +3
Query: 426 GGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAA--AAPKGKNVA---EASVAAAT 480
GG A T +P T S + P PTT A A+PK A +A+ AT
Sbjct: 96 GGQSPSAAPTTSPSTPAATTPVSSP-VAAPPTTPTTPAPVASPKSSPPATSPKAAAPTAT 272
Query: 481 EPTA--VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPI--------GDK 530
P A PA +P S A +S S+ T AA + TP G K
Sbjct: 273 PPAASSPPAVTPVSTPPPAPVPVKSPPTPAPVSSPPAVTPVAAPTTTPAVPAPAPSKGKK 452
Query: 531 -EKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTS 576
+K++ P P PP+PP+ P PS + S PG ATT+
Sbjct: 453 NKKKHGAPAPSPALLGPPAPPA---GAPGPSE----DASSPGPATTA 572
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 53.1 bits (126), Expect = 5e-07
Identities = 45/179 (25%), Positives = 69/179 (38%), Gaps = 6/179 (3%)
Frame = +3
Query: 437 APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVP-ASSPASAGA 495
+P+ + + + S G S P PTT +P G + P A+P ASSP
Sbjct: 486 SPRTPKSSPSPSAGGLSPPSPSPTTTTPSPSG---------SPPSPVAIPPASSPVPTSG 638
Query: 496 TAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDA 555
A + + A + ++ + S P G + +P P +P PST
Sbjct: 639 PTASSPSPVVSTPPAGGPMASSPSPVVSTPPAGGPMASSPSPAGGPPALSPAGGPSTAAG 818
Query: 556 GPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNA-----IEPSEFLL 609
GP P+P G + PG + A AP GS + P+ + PS FL+
Sbjct: 819 GP-PAPGPGGAATSPGPGGAGASGPGGAAAAP--GSPGSNSTAPSGNSGSFVAPSNFLV 986
>TC85985 similar to GP|5139695|dbj|BAA81686.1 expressed in cucumber
hypocotyls {Cucumis sativus}, partial (42%)
Length = 892
Score = 47.0 bits (110), Expect = 3e-05
Identities = 39/137 (28%), Positives = 59/137 (42%), Gaps = 1/137 (0%)
Frame = +3
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAAATEP-TAVPASSPASAGATAAFAAESSIGATA 509
G P + PTT+ P + A VAA T+P + P +SP S+ ++ A + A +
Sbjct: 114 GGQSPSSAPTTSP--PTVTTPSAAPVAAPTKPKSPAPVASPKSSPPASSPTAATVTPAVS 287
Query: 510 ASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSC 569
+A V K+ A+S + TP P +PP+P P+ SPP S
Sbjct: 288 PAAPVPVAKSPAASSPVVAPV----STPPKPAPVSSPPAPV------PVSSPPTPVPVSS 437
Query: 570 PGAATTSEAAQIEQAPA 586
P A+T + PA
Sbjct: 438 PPTASTPAVTPSAEVPA 488
Score = 45.8 bits (107), Expect = 7e-05
Identities = 35/130 (26%), Positives = 54/130 (40%)
Frame = +3
Query: 453 SKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASA 512
S P P + P + A V ++ PT VP SSP +A +T A + + A A S
Sbjct: 327 SSPVVAPVSTPPKPAPVSSPPAPVPVSSPPTPVPVSSPPTA-STPAVTPSAEVPAAAPSK 503
Query: 513 GVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGA 572
TK K K++ P P + PP+P P+ +P + S PG
Sbjct: 504 SKKKTK-----------KGKKHSAPAPSPALEGPPAP-------PVGAPGPSLDASSPGP 629
Query: 573 ATTSEAAQIE 582
A+ ++ + E
Sbjct: 630 ASAADESGAE 659
>BF644046
Length = 597
Score = 45.4 bits (106), Expect = 1e-04
Identities = 20/51 (39%), Positives = 31/51 (60%)
Frame = +3
Query: 214 YEYMFSELGIRLPFSPFVQTVLRDINTAPCQLHPNAWAFIRCFEILSAAVG 264
Y ++F ++G + PF+ F L+ +N A QLHPN AF+ FEI ++G
Sbjct: 15 YSFVFEDIGFKFPFTNFECDFLKALNVASSQLHPNCCAFMCGFEISCESLG 167
>TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-protein {Pyrus
communis}, partial (30%)
Length = 956
Score = 44.3 bits (103), Expect = 2e-04
Identities = 44/151 (29%), Positives = 57/151 (37%), Gaps = 8/151 (5%)
Frame = +3
Query: 436 KAPKRRRLTKASSDAGTSK---PGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
+ PK +TK +S A G T+ A G + A T PT P ++P +
Sbjct: 78 QTPKPLSITKMASYAAVQVFLILGLLATSCFAQAPGAAPTQPPSATPTPPTPAPVATPPT 257
Query: 493 AGATAAFAAESSIGATAASAGVNATKAAASSD---TPIGDKEKENETPKSPPR--QDAPP 547
A T A + A+A AT A A+S TP D + T SPP P
Sbjct: 258 A--TPPTATPPTATPPPAAAPTPATPAPATSPPAPTPTSDAPTPDSTSSSPPAPGPGGPA 431
Query: 548 SPPSTHDAGPMPSPPHQGEKSCPGAATTSEA 578
P + DA P PS K A S A
Sbjct: 432 PGPGSTDAPPSPSAAFSINKPIMAATALSAA 524
>TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myosin heavy
chain {Arabidopsis thaliana}, partial (17%)
Length = 2546
Score = 44.3 bits (103), Expect = 2e-04
Identities = 68/285 (23%), Positives = 117/285 (40%), Gaps = 39/285 (13%)
Frame = +2
Query: 600 NAIEPSEFLLAGLNRDV-------IEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFN 652
NAIE + LA RD+ +E E L + L E+ L EKFN
Sbjct: 362 NAIEEN---LAVRGRDIELTSARNLELESLHESLTRDSEQKLQ----------EAIEKFN 502
Query: 653 AANVEAERLKAESAKHQEAAAA-----------WEKRFDKLATQAGKDK------VYADK 695
+ + E + L + +E A +E+ KLA+ +++ V A+K
Sbjct: 503 SKDSEVQSLLEKIKILEENIAGAGEQSISLKSEFEESLSKLASLQSENEDLKRQIVEAEK 682
Query: 696 MIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEAL--------- 746
+ + L +K + DEL SL + EKE ++L++ L
Sbjct: 683 KTSQSFSENELLVGTNIQLKTKIDELQESLNSVVSEKEVTAQELVSHKNLLAELNDVQSK 862
Query: 747 ---IAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETH 803
I +E+ IL E +L ++AL + +K +E+ ++ + +IK E
Sbjct: 863 SSEIHSANEVRILEVESKL-QEALQKHTEKESET-----KELNEKLNTLEGQIKIYEEQA 1024
Query: 804 AEALA---TKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADI 845
EA+A + AE+ L K+K LE + ++ K++E + A I
Sbjct: 1025HEAVAAAENRKAELEESLIKLKHLEAAVEEQQNKSLERETETAGI 1159
Score = 34.3 bits (77), Expect = 0.22
Identities = 41/195 (21%), Positives = 76/195 (38%), Gaps = 33/195 (16%)
Frame = +3
Query: 613 NRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAA 672
N D ++ EV + + + ++ L L +EA+ KAES H+E
Sbjct: 1464 NEDSLKSEVETLKIEIAEKSALQSRLH---------------EIEAQLAKAESRLHEEVG 1598
Query: 673 AAWEKRFDKLATQAGKDKVYADKM--IGTAGIKIGELEDQLALMK--------EEAD--E 720
+ + + K + Y K+ I K+ ELE +L L + EE+ E
Sbjct: 1599 SVQAAASQREVDLSSKFEDYEQKISEINVLNGKVAELEKELHLAQDTIANQKGEESQKLE 1778
Query: 721 LDASLQACKKEKE---------------------QAEKDLIARGEALIAKESELAILCAE 759
L+A+L+ +E E QA++ + +GE + K+ L + +
Sbjct: 1779 LEAALKNSVEELETKKNEISLLQKQVIEFEQKLQQADEKISVKGEEAVDKKDALEVKSRD 1958
Query: 760 LELVKKALAEQEKKS 774
+ + + +KKS
Sbjct: 1959 FSISSPSKRKSKKKS 2003
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 42.7 bits (99), Expect = 6e-04
Identities = 52/191 (27%), Positives = 84/191 (43%), Gaps = 21/191 (10%)
Frame = +2
Query: 656 VEAERLKAESAKHQEAAAAWEKRFDKL------------ATQAGKDKVY--ADKMIGTAG 701
VE E +K E ++ +E + E L A A + KV +++MI T
Sbjct: 1301 VELENVKKEHSELKEKESELESTVGNLHVKLRKSKSELEACSADESKVRGASEEMILTLS 1480
Query: 702 IKIGELED---QLALMKEEADELDASLQACKKEKEQAEKDL-IARGEALIAKESELAILC 757
E E+ ++ MK + DEL +A K E+AEK L A EA AK +E + +
Sbjct: 1481 RLTSETEEARREVEDMKNKTDELKKEAEATKLALEEAEKKLKEATEEAEAAKAAEASAIE 1660
Query: 758 AELELVKKALAEQEKKSAES---LALAKSDMEAVMQATSEEIKKATETHAEALATKDAEI 814
L ++ A + S+ES + ++ + E++ + E K A A A +A
Sbjct: 1661 QITVLTERTSARRASTSSESGAAMTISTEEFESLKRKVEESDKLADMKVDAAKAQVEAVK 1840
Query: 815 ASQLAKIKSLE 825
AS+ +K LE
Sbjct: 1841 ASENEVLKKLE 1873
Score = 38.1 bits (87), Expect = 0.016
Identities = 69/314 (21%), Positives = 119/314 (36%), Gaps = 22/314 (7%)
Frame = +2
Query: 547 PSPPSTHDAGPMP-SPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPS 605
PSP + G + SPP Q K + E A + P + P A
Sbjct: 329 PSPSLKPEVGEIDTSPPFQSVKDA--VSLFGEGAFSGERPI---------HKKPKAYSAE 475
Query: 606 EFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAES 665
L V EKE LN KE+ + ++LK +
Sbjct: 476 RVLAKETQLHVAEKE-----LNKLKEQVKNAETTKAQALVELERAQRTVDDLTQKLKLIT 640
Query: 666 AKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASL 725
+ A A E + + G+ G G ELE+ + ELDA+
Sbjct: 641 ESRESAVKATEAAKSQAKQKYGESD-------GVNGAWKEELENAVQRYASIMTELDAAK 799
Query: 726 QACKKEKEQAEKDLIARGEAL-IAKESELAI---------LCAELELVKKA-----LAEQ 770
Q +K +++ + AR A+ +E+E A+ L E+ VK++ LA
Sbjct: 800 QDLRKTRQEYDSSSDARVSAVKRTEEAENAMKENTERVSELSKEISAVKESIEQTKLAYV 979
Query: 771 EKKSAESLALAKSD-MEAVMQATSEEIKK-----ATETHAEALATKDAEIASQLAKIKSL 824
E + ++L L + D + +AT E+ KK E + E + +A++A + +I +L
Sbjct: 980 ESQQQQALVLTEKDALRQSYKATLEQSKKKLLALKKEFNPEITKSLEAQLAETMNEIAAL 1159
Query: 825 EDELATEKAKAIEA 838
EL +++ +++
Sbjct: 1160HTELENKRSSDLDS 1201
>BQ751419 weakly similar to GP|21629340|gb L509.2 {Leishmania major}, partial
(1%)
Length = 766
Score = 41.6 bits (96), Expect = 0.001
Identities = 37/153 (24%), Positives = 54/153 (35%), Gaps = 12/153 (7%)
Frame = +1
Query: 447 SSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIG 506
+S + +++P PTTA++ P G A T + ++PA A TA +
Sbjct: 52 ASSSPSARPSPPPTTASSTPPG---APTPSPTPTRRATLNGNTPAPASPTAQATPPTGPS 222
Query: 507 ATAASAGVNATKAAASSDTPIGDKEKENE--TPKSPPRQDAPPSPPSTHDAGPMPSPP-- 562
A S T SS T + T SP PP+P + PP
Sbjct: 223 TAAPSPSTCTTTGPTSSSTSASSPAATSPRTTSPSPHSSGTPPAPAPSASRSSTSRPPRA 402
Query: 563 ---HQGEKSCP-----GAATTSEAAQIEQAPAP 587
+S P A TT+ + Q P P
Sbjct: 403 RARRAASRSSPWATPARACTTAPTSASRQTPPP 501
Score = 37.0 bits (84), Expect = 0.035
Identities = 31/108 (28%), Positives = 42/108 (38%)
Frame = +1
Query: 445 KASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESS 504
+A+ G S P+T + AS AAT P + SP S+G A A +S
Sbjct: 196 QATPPTGPSTAAPSPSTCTTTGPTSSSTSASSPAATSPRTT-SPSPHSSGTPPAPAPSAS 372
Query: 505 IGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPST 552
+T+ A +AA+ S P S RQ PP P T
Sbjct: 373 RSSTSRPPRARARRAASRSSPWATPARACTTAPTSASRQ-TPPPSPRT 513
Score = 35.8 bits (81), Expect = 0.077
Identities = 37/129 (28%), Positives = 49/129 (37%), Gaps = 13/129 (10%)
Frame = +1
Query: 478 AATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP 537
A++ P+A P+ P +A +T A S T +A + +TP P
Sbjct: 52 ASSSPSARPSPPPTTASSTPPGAPTPSPTPTR--------RATLNGNTP---------AP 180
Query: 538 KSPPRQDAPPSPPSTHDAGPMPSP-----PHQGEKSCPGAATTS--------EAAQIEQA 584
SP Q PP+ PST A P PS P S A TS ++ A
Sbjct: 181 ASPTAQATPPTGPST--AAPSPSTCTTTGPTSSSTSASSPAATSPRTTSPSPHSSGTPPA 354
Query: 585 PAPEVGSSS 593
PAP SS
Sbjct: 355 PAPSASRSS 381
Score = 29.6 bits (65), Expect = 5.5
Identities = 25/95 (26%), Positives = 40/95 (41%), Gaps = 14/95 (14%)
Frame = +1
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAF------ 499
++S + TS+P + AA + A + A T PT+ +P + TAA
Sbjct: 364 SASRSSTSRP-PRARARRAASRSSPWATPARACTTAPTSASRQTPPPSPRTAAKPKASRC 540
Query: 500 --------AAESSIGATAASAGVNATKAAASSDTP 526
S + A AA+AG AT+ A++ P
Sbjct: 541 TPSRSRAPTEPSKMAAAAAAAGTRATRPRATAAAP 645
>TC79555 homologue to GP|10567859|gb|AAG18593.1 L8019.5 {Leishmania major},
partial (1%)
Length = 1410
Score = 41.2 bits (95), Expect = 0.002
Identities = 46/167 (27%), Positives = 66/167 (38%), Gaps = 15/167 (8%)
Frame = +2
Query: 436 KAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGA 495
K+P R R + S S G +P AAAA + A+ PT +S A G
Sbjct: 722 KSPPRPRPARLPSARRASPRGGRPLAAAAA-----AGATTSTTASRPTRSARTSAARRGR 886
Query: 496 TAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK---------ENETPKSPPRQD-- 544
A S G+ G + T+A + + + + N P SP R+
Sbjct: 887 PAI*DGLCSSGSMRTCRG*SRTRALTFTSSTVSVRSNGSFGASSGPSNTCPSSPRRRSGR 1066
Query: 545 ---APPSPPSTHDAGPMPSPPHQGEKSCPGA-ATTSEAAQIEQAPAP 587
A S P+T+ P P PP S P A TS++A ++PAP
Sbjct: 1067LPAATTSSPATNANSP-PFPPASPSSSVPPAKPPTSKSA---RSPAP 1195
Score = 35.8 bits (81), Expect = 0.077
Identities = 31/114 (27%), Positives = 49/114 (42%), Gaps = 8/114 (7%)
Frame = +2
Query: 418 TEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSK------PGAQPTTAA--AAPKGK 469
+ G G N S+ + RL A++ + + P A P+++ A P
Sbjct: 992 SNGSFGASSGPSNTCPSSPRRRSGRLPAATTSSPATNANSPPFPPASPSSSVPPAKPPTS 1171
Query: 470 NVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASS 523
A + + P+ P+SSP SA A+A+ +S A SAG T+ ASS
Sbjct: 1172 KSARSPAPV*SRPSPPPSSSPGSA-ASASSRCSASSRARPTSAGSRRTRTTASS 1330
>TC89270 similar to GP|22773423|gb|AAL66352.2 catalase-peroxidase
{Neurospora crassa}, partial (25%)
Length = 792
Score = 40.8 bits (94), Expect = 0.002
Identities = 46/177 (25%), Positives = 65/177 (35%), Gaps = 11/177 (6%)
Frame = +1
Query: 434 STKAPKRRRLTKASSDAG-----TSKPGAQPT---TAAAAPKGKNVAEASVAAATEPTAV 485
+T P R T+AS T + G PT TA + A A T+ V
Sbjct: 259 TTSPPSSRSTTRASRRTSRP**PTPRTGGLPTLATTAVCSSAWPGTAPARTEFTTDAEVV 438
Query: 486 PASSPASAGATAAFAAESS---IGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPR 542
+S AS +TA +S +G S+ AT++ + P SP
Sbjct: 439 ERASNASHRSTAGRTMSASTRPVGCCGPSSKSTATRSRGPTS*-----SWPATWPSSPWV 603
Query: 543 QDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLP 599
P SP + G P+ P G PG+ATTS +A A S++ P
Sbjct: 604 SRRPASPEAVPTPGK-PTSPSTGAARTPGSATTSATPTATRARADRASSTARRRQSP 771
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 40.8 bits (94), Expect = 0.002
Identities = 34/153 (22%), Positives = 58/153 (37%)
Frame = +3
Query: 452 TSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAAS 511
T+KP P++ + +P + A + P+ PA SP+ + +T AT+
Sbjct: 255 TAKP---PSSVSVSPTNSPASPAKSPTLSPPSQTPAVSPSGSASTPP-------PATSPP 404
Query: 512 AGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPG 571
A A + +S I + TP +PP+ + + P+ +P
Sbjct: 405 AKSPAVQPPSSVSPAISPSNNVSSTPPVSSPASSPPTAAVSPVSSPVEAPSVSSPPEASS 584
Query: 572 AATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEP 604
A S +A APA + SS P + P
Sbjct: 585 AGIPSSSATPADAPAATLPSSKSPGTSPASSSP 683
>TC85721 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (60%)
Length = 995
Score = 40.4 bits (93), Expect = 0.003
Identities = 41/118 (34%), Positives = 57/118 (47%), Gaps = 16/118 (13%)
Frame = +1
Query: 442 RLTKASSDAGT--SKPGAQPTTAAAA-PKGKNV-AEASVAAATEP-TAVPASSPASAGAT 496
+L+ A+ A +KP A+P A PK K+V A+ AAAT+P A S AS T
Sbjct: 478 KLSSATKPAAKVKAKPAAKPKAKAVVKPKTKSVTAKPKAAAATKPKPAATKSKAASKAKT 657
Query: 497 AAFA----------AESSIGATAASAGVNATK-AAASSDTPIGDKEKENETPKSPPRQ 543
AA A A +S G A+A A K AAA P+ + + +T KSP ++
Sbjct: 658 AAKAKPNAKVAKTTARTSPGKKVAAAKPAAKKVAAAKKKAPVKSVKPKTKTVKSPAKK 831
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 40.4 bits (93), Expect = 0.003
Identities = 29/97 (29%), Positives = 39/97 (39%), Gaps = 16/97 (16%)
Frame = +3
Query: 482 PTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP---- 537
P++ P S+PA+ A S AT +S + T +A TP TP
Sbjct: 24 PSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISP 203
Query: 538 ------------KSPPRQDAPPSPPSTHDAGPMPSPP 562
++PP D PSPPS+ P PSPP
Sbjct: 204 PSGGGTTPSPPSRTPPSSDDSPSPPSS-KTPPPPSPP 311
Score = 35.0 bits (79), Expect = 0.13
Identities = 23/89 (25%), Positives = 32/89 (35%)
Frame = +3
Query: 516 ATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATT 575
+T A TP TP SPP P +PP + + PSPP S P T
Sbjct: 42 STPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAISPPSGGGT 221
Query: 576 SEAAQIEQAPAPEVGSSSYYNMLPNAIEP 604
+ + P+ + S + P P
Sbjct: 222 TPSPPSRTPPSSDDSPSPPSSKTPPPPSP 308
Score = 31.6 bits (70), Expect = 1.5
Identities = 24/72 (33%), Positives = 28/72 (38%), Gaps = 3/72 (4%)
Frame = +3
Query: 519 AAASSDTPIGDKEKENETPKSPPRQD-APPS--PPSTHDAGPMPSPPHQGEKSCPGAATT 575
A SS P TP +PP A PS PPST + P PS P S P
Sbjct: 18 ATPSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNSPPSPPTTPAI 197
Query: 576 SEAAQIEQAPAP 587
S + P+P
Sbjct: 198 SPPSGGGTTPSP 233
Score = 31.6 bits (70), Expect = 1.5
Identities = 17/55 (30%), Positives = 22/55 (39%)
Frame = +3
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
S P P SPP + A P P P S P ++ + AP P S+S
Sbjct: 3 SSPPPATPSSPPPSTPANPPPVTPSAPPPSTPATPSSPPPSTTPSAPPPSTPSNS 167
Score = 30.4 bits (67), Expect = 3.2
Identities = 19/56 (33%), Positives = 25/56 (43%)
Frame = +3
Query: 538 KSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
K PP PP PPS H A P P P S G+ + ++ P+P + SS
Sbjct: 561 KPPPPAPIPPRPPS-HVAPPPPPPAFIS--SSGGSGSNYSGGELLPPPSPGIAFSS 719
>BQ751262 similar to GP|6715100|gb|A versicolorin B synthase {Aspergillus
parasiticus}, partial (11%)
Length = 710
Score = 40.0 bits (92), Expect = 0.004
Identities = 44/188 (23%), Positives = 76/188 (40%), Gaps = 2/188 (1%)
Frame = +3
Query: 385 SRTPKYLEDMNFTNAELLKARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRL- 443
+R+ + L T+ L RR + P+ + + + R R+
Sbjct: 162 TRSGRTLSATTLTHGTLSSPTSRRASSSLPRVLRDSPTPAPSTTQMPSPPPAGLLRSRMP 341
Query: 444 TKASSDAGTSKPGAQPTTAAAAPKGK-NVAEASVAAATEPTAVPASSPASAGATAAFAAE 502
T S A S P +T +A+P+ + + AE+S A +T PT P +++ A + +
Sbjct: 342 TMPSRSAHISSP---LSTRSASPRPRTSTAESSWARSTAPTPSPQRRKSASRARRRSSPK 512
Query: 503 SSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPP 562
S T+ S + + ++S+ T K SPP +PP+P + A SPP
Sbjct: 513 PSAARTSRSTPSPSPRRSSSTAT------KRPPASSSPPTPASPPTP--SRRARRSSSPP 668
Query: 563 HQGEKSCP 570
E P
Sbjct: 669 APSEPPAP 692
>TC85722 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (61%)
Length = 897
Score = 40.0 bits (92), Expect = 0.004
Identities = 34/121 (28%), Positives = 57/121 (47%), Gaps = 3/121 (2%)
Frame = +3
Query: 428 AENQAESTKAPKRRRLTKASSDAGTSKP---GAQPTTAAAAPKGKNVAEASVAAATEPTA 484
A+ +A+ PK + + K + + T+KP A+P +AA PK V +A AA +P A
Sbjct: 471 AKVKAKPAAKPKAKAVVKPRTKSVTAKPKAAAAKPKSAATKPKAA-VGKAKTAAKVKPNA 647
Query: 485 VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD 544
A + +S G A+A K AA TP+ + + + +T +SP ++
Sbjct: 648 KVAKT----------TTRTSPGKKVATA-----KPAAKKKTPVKNVKPKTKTVRSPAKKV 782
Query: 545 A 545
A
Sbjct: 783 A 785
>TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2G14_40
{Arabidopsis thaliana}, partial (11%)
Length = 1320
Score = 39.7 bits (91), Expect = 0.005
Identities = 40/146 (27%), Positives = 55/146 (37%), Gaps = 3/146 (2%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEPTA-VPASSPASAGATAAFAAESSIGATAASAGVNAT 517
P A+ P + A A V A P PA +P A A + T A A +
Sbjct: 537 PLPPASKPPKVSPAPAKVPKAPAPAKEAPAPAPTHKKKAPAPAPDKE---TPAPAPTHKK 707
Query: 518 KAAASSDTPIGDKEKENETPKSPPRQDAPPS--PPSTHDAGPMPSPPHQGEKSCPGAATT 575
K A S +P+ +P P D P S PS D P P PPH+ ++ +
Sbjct: 708 KKAPKS-SPVPSPAILPPSPAPTPAIDTPSSAPAPSPEDDAPEPPPPHKHKRRKHKHSKH 884
Query: 576 SEAAQIEQAPAPEVGSSSYYNMLPNA 601
+ + AP P SS+ P A
Sbjct: 885 KKHHALALAPEPTSSSSTIIRRSPPA 962
Score = 32.7 bits (73), Expect = 0.65
Identities = 27/125 (21%), Positives = 42/125 (33%), Gaps = 8/125 (6%)
Frame = +2
Query: 473 EASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEK 532
++ V AT PT+ P P AT A + + K T
Sbjct: 86 KSPVPKATTPTSAPVKPPVPKVATPTVAPAKPLVPKVTTPTAAPVKPIVPKTTTPTSAPL 265
Query: 533 ENETPKSPPRQDAPPSPP--------STHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQA 584
+ PK+P + +P PP + P+P PP P T A +
Sbjct: 266 KPPVPKAPAPKPSPVKPPVPKSKPPTTAPVKSPVPDPPKLAPVKLPVPKVT--PALSPKT 439
Query: 585 PAPEV 589
P+P++
Sbjct: 440 PSPKI 454
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 39.7 bits (91), Expect = 0.005
Identities = 57/237 (24%), Positives = 72/237 (30%), Gaps = 25/237 (10%)
Frame = -1
Query: 376 ATVLEGADLSRTPKYLEDM--NFTNAELLKARERRMARFSPKFNTEGDAKKRGGAENQAE 433
A VL + R P+ L + T A +L R AR P+ +
Sbjct: 995 ARVLRARTVRRLPRRLPLLYRRATAAAVLSCSARSRARDRPRADVR-------------- 858
Query: 434 STKAPKRRRLTKASSDAGTSK-PGAQPTTAAAAPKGKNVAEASVAA-----ATEPTAVPA 487
+ P R RL A S A + P A P +A P G A A + A P P
Sbjct: 857 ALPRPPRSRLYAARSCARVLRAPSALPAPPSATPPGAPCARAPHSIPRAHPAATPLLPPP 678
Query: 488 SSPASAGATAAFAAESSIGATAASAGVNATKAAA-------------SSDTPIGDKEKEN 534
P IG A V + A P + E
Sbjct: 677 RPPPPPPEWERVPLAKKIGGQPRPALVPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEA 498
Query: 535 ETPKSPP----RQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
P PP R AP PPS D P P QG PG + + + P P
Sbjct: 497 APPPPPPPRGARPRAPLRPPSAPDPAGHPRAPRQGVAVVPGPRSGNSPRAAHRGPEP 327
Score = 34.7 bits (78), Expect = 0.17
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = -2
Query: 537 PKSPPRQDAPPSPPSTHDAGPMPSPPHQ 564
P PP++ PP PP TH P SPP +
Sbjct: 148 PPPPPQRPPPPPPPHTHQPEPPTSPPRR 65
Score = 31.6 bits (70), Expect = 1.5
Identities = 39/172 (22%), Positives = 52/172 (29%), Gaps = 22/172 (12%)
Frame = -1
Query: 438 PKRRRLTKASSDAGTSKPGAQPTTAAAA-------------PKGKNVAEASVAAATEPTA 484
P+ R+ A G +P P A P A AA P
Sbjct: 659 PEWERVPLAKKIGGQPRPALVPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEAAPPPPP 480
Query: 485 VPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD 544
P + A A + + A GV S ++P P+ PR
Sbjct: 479 PPRGARPRAPLRPPSAPDPAGHPRAPRQGVAVVPGPRSGNSPRA--AHRGPEPRPLPRPP 306
Query: 545 APPSPPSTHDAGPMPSP-------PHQGEKSCPGAATTSEAAQIE--QAPAP 587
P PP+ GP P+P P E P A A ++ APAP
Sbjct: 305 PPTRPPAAPGPGPPPAPGPTPLSKPPAAEGIPPATAPPPRARRLRPPSAPAP 150
Score = 31.2 bits (69), Expect = 1.9
Identities = 31/123 (25%), Positives = 41/123 (33%), Gaps = 1/123 (0%)
Frame = -1
Query: 442 RLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAA 501
R A AG + Q P+ N A+ EP +P P + A
Sbjct: 446 RPPSAPDPAGHPRAPRQGVAVVPGPRSGNSPRAA-HRGPEPRPLPRPPPPTRPPAAP--- 279
Query: 502 ESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPP-SPPSTHDAGPMPS 560
G A +K A+ P PP APP +PP+ A P P+
Sbjct: 278 --GPGPPPAPGPTPLSKPPAAEGIPPATAPPPRARRLRPPSAPAPPPTPPAA--APPAPT 111
Query: 561 PPH 563
PPH
Sbjct: 110 PPH 102
Score = 29.3 bits (64), Expect = 7.2
Identities = 17/49 (34%), Positives = 20/49 (40%)
Frame = -2
Query: 539 SPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
+PPR PP PP P P PPH + P + Q PAP
Sbjct: 172 APPRPPPPPPPPPQRP--PPPPPPHTHQPEPPTSPPR------RQPPAP 50
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 39.3 bits (90), Expect = 0.007
Identities = 30/110 (27%), Positives = 48/110 (43%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATK 518
PT+ +++P A + V+ T P PAS+PAS+ + + +S A++ +
Sbjct: 84 PTSPSSSPS----ASSPVSTTTSP---PASTPASSPVSTTTSPPASTPASSPVPTTTSPP 242
Query: 519 AAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKS 568
A + +P+ S P A PSP ST P PS G +S
Sbjct: 243 APTPASSPVSTNSPTASPAGSLPAA-ATPSPSSTTAGSPSPSSTPPGTRS 389
Score = 38.5 bits (88), Expect = 0.012
Identities = 39/129 (30%), Positives = 56/129 (43%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATK 518
P A P G +A S PT+ P+SSP+++ + S +T AS+ V+ T
Sbjct: 18 PPGVAQTPSGNGIAP-SPYQFNNPTS-PSSSPSASSPVST--TTSPPASTPASSPVSTTT 185
Query: 519 AAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEA 578
+ +S TP T SPP SP ST+ SP S P AAT S +
Sbjct: 186 SPPAS-TPASSPVP---TTTSPPAPTPASSPVSTN------SPTASPAGSLPAAATPSPS 335
Query: 579 AQIEQAPAP 587
+ +P+P
Sbjct: 336 STTAGSPSP 362
Score = 37.4 bits (85), Expect = 0.026
Identities = 29/132 (21%), Positives = 54/132 (39%)
Frame = +3
Query: 426 GGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAV 485
G A+ + + AP + +S + +S + P + +P A + V+ T P
Sbjct: 24 GVAQTPSGNGIAPSPYQFNNPTSPS-SSPSASSPVSTTTSPPASTPASSPVSTTTSP--- 191
Query: 486 PASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDA 545
PAS+PAS+ + + A++ + + T + A S + T SP
Sbjct: 192 PASTPASSPVPTTTSPPAPTPASSPVSTNSPTASPAGSLPAAATPSPSSTTAGSPSPSST 371
Query: 546 PPSPPSTHDAGP 557
PP S+++ P
Sbjct: 372 PPGTRSSNETTP 407
>TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens culinaris},
partial (86%)
Length = 1215
Score = 38.9 bits (89), Expect = 0.009
Identities = 46/167 (27%), Positives = 70/167 (41%), Gaps = 24/167 (14%)
Frame = +2
Query: 404 ARERRMARFSPKFNTEGDAKKRGGAENQAESTKAP-----KRRRLTKASSDAGTSKPGAQ 458
A++ A+ PK + K A+++A++ A K + + A +KP A+
Sbjct: 485 AKKPAAAKPKPKPKAKAPVAKAPAAKSKAKAAPAKAKAKAKAKAAPAKAKPAAKAKPAAK 664
Query: 459 PTTAA-------AAPKGK-NVAEAS---VAAATEPT--------AVPASSPASAGATAAF 499
AA A PK K VA+A+ VAA +P A PA+ PA A T
Sbjct: 665 AKPAAKAKPVAKAKPKAKAPVAKANAVPVAAKAKPAAKAKPAAKAKPAARPAKASRT--- 835
Query: 500 AAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAP 546
+ +S G AA+A KAA + K+ K+ P + AP
Sbjct: 836 STRTSPGKRAAAAKPAVKKAATPVKKAVPAKKAATPVKKAAPARKAP 976
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 38.5 bits (88), Expect = 0.012
Identities = 51/222 (22%), Positives = 68/222 (29%), Gaps = 34/222 (15%)
Frame = +3
Query: 403 KARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRL-TKASSDAGTSKPGAQPTT 461
K R+ + P+ T A G NQ +T PK++ T + P P T
Sbjct: 99 KKSPRKKEKPPPRPATPPPAPAPPGPPNQTTNTTNPKKQHTPTTPTHPPRPHTPPTPPPT 278
Query: 462 AAAAPKGKNVAEASVAAATEPTA-----VPASSPASAGATAAFA----------AESSIG 506
P + + T PTA PA P G
Sbjct: 279 TQTTPPREAPTDPGTTRGTPPTADDAQTSPAHHPGEGGGKGGAPPPPPRPPPRHGRGGHQ 458
Query: 507 ATAASAGVNATKAAASSDTPIGDKEKENETP----------KSPPRQDAPPSPPS---TH 553
A T+ D P D K+ P K P +D PP P+
Sbjct: 459 NQTREAKRTTTEPQGKRDRPRTDPRKKKHPPPHQKEGGNTQKEPRGEDPPPHTPAGGGGG 638
Query: 554 DAGPMPSPPHQ----GEKSCP-GAATTSEAAQIEQAPAPEVG 590
GP P PP + GE+ P GA E + +Q A G
Sbjct: 639 PGGPRPPPPKKTARGGEERPPAGARARREKTRGDQQEAAPAG 764
Score = 33.9 bits (76), Expect = 0.29
Identities = 32/143 (22%), Positives = 46/143 (31%)
Frame = +1
Query: 410 ARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGK 469
AR K NT KK+G ++ + P T +PG A AP +
Sbjct: 520 ARTPGKKNTPPHTKKKGETHKKSPGGRTPPP-----------TPRPGGGGARGAPAPPPQ 666
Query: 470 NVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGD 529
+ + P P AT+ GA + +A +
Sbjct: 667 K--KQREGGKSGPPRAPGHEEKKPAATSR--KRPPRGAREKNPAAHAREREKKKTPKKNP 834
Query: 530 KEKENETPKSPPRQDAPPSPPST 552
+ N PK+PP Q P PP T
Sbjct: 835 QNHNNNPPKTPPPQKKHPPPPKT 903
Score = 33.1 bits (74), Expect = 0.50
Identities = 20/76 (26%), Positives = 29/76 (37%), Gaps = 1/76 (1%)
Frame = +1
Query: 530 KEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEK-SCPGAATTSEAAQIEQAPAPE 588
+++ E K+PPR PP PP A P P Q K + P + P P+
Sbjct: 97 QKRAPEKKKNPPRGQPPPPPPPPPQAPPTKPPTRQTPKNNTPPPPPHTPPDPTPPQPPPQ 276
Query: 589 VGSSSYYNMLPNAIEP 604
++ P EP
Sbjct: 277 PPKPPHHEKRPRTPEP 324
Score = 31.6 bits (70), Expect = 1.5
Identities = 18/48 (37%), Positives = 24/48 (49%), Gaps = 1/48 (2%)
Frame = +3
Query: 518 KAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPM-PSPPHQ 564
K S P G K+ T KSP +++ PP P+T P P PP+Q
Sbjct: 48 KGHPSQKPPTGRSHKK--TKKSPRKKEKPPPRPATPPPAPAPPGPPNQ 185
Score = 31.6 bits (70), Expect = 1.5
Identities = 41/162 (25%), Positives = 57/162 (34%), Gaps = 25/162 (15%)
Frame = +1
Query: 427 GAENQAESTKAPKRRRLTKASSDAGTSK---PGAQPTTAAAAPKGKNVAEASVAAATEPT 483
G + T+ K+R + + GT PG + T KG+ + S T P
Sbjct: 436 GTGGEDTKTRPGKQREPQQNPRERGTGHARTPGKKNTPPHTKKKGET-HKKSPGGRTPP- 609
Query: 484 AVPASSPASAGATAAFAA-------ESSIGATAASAGVNATKAAASSDT--PIGDKEK-- 532
P P GA A A E + G K AA+S P G +EK
Sbjct: 610 --PTPRPGGGGARGAPAPPPQKKQREGGKSGPPRAPGHEEKKPAATSRKRPPRGAREKNP 783
Query: 533 --------ENETPKSPPRQ---DAPPSPPSTHDAGPMPSPPH 563
+ +TPK P+ + P +PP P P PH
Sbjct: 784 AAHAREREKKKTPKKNPQNHNNNPPKTPPPQKKHPPPPKTPH 909
Score = 30.4 bits (67), Expect = 3.2
Identities = 34/140 (24%), Positives = 43/140 (30%)
Frame = +3
Query: 423 KKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEP 482
KK+ +Q E K R G P P P G A
Sbjct: 534 KKKHPPPHQKEGGNTQKEPR--------GEDPPPHTPAGGGGGPGGPRPPPPKKTARGGE 689
Query: 483 TAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPR 542
PA + A T E AA AG K + + K + + PK P+
Sbjct: 690 ERPPAGARARREKTRGDQQE------AAPAGRPRKKPRGAREREGKKKNPQKKPPK--PQ 845
Query: 543 QDAPPSPPSTHDAGPMPSPP 562
Q P +PP T P P P
Sbjct: 846 QQPPKNPPPTKKTPPPPKNP 905
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,459,355
Number of Sequences: 36976
Number of extensions: 433365
Number of successful extensions: 6803
Number of sequences better than 10.0: 275
Number of HSP's better than 10.0 without gapping: 3716
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5503
length of query: 919
length of database: 9,014,727
effective HSP length: 105
effective length of query: 814
effective length of database: 5,132,247
effective search space: 4177649058
effective search space used: 4177649058
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0590c.8


