

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
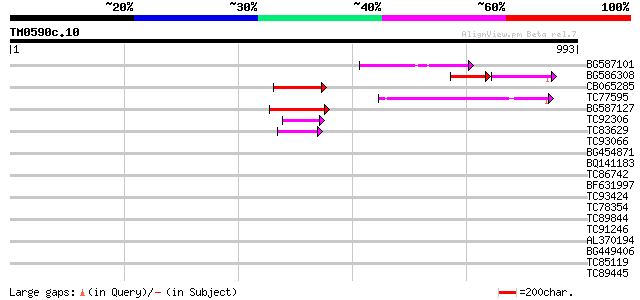
Query= TM0590c.10
(993 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 119 4e-27
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 77 2e-26
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 83 5e-16
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 83 6e-16
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 77 4e-14
TC92306 45 2e-04
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 44 4e-04
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 38 0.022
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 36 0.064
BQ141183 weakly similar to GP|12654765|gb| Similar to erythrobla... 32 0.92
TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG00... 32 1.2
BF631997 weakly similar to GP|18542925|gb Putative pol polyprote... 30 3.5
TC93424 30 4.6
TC78354 similar to GP|13272425|gb|AAK17151.1 unknown protein {Ar... 30 4.6
TC89844 similar to GP|3258570|gb|AAC24380.1| Unknown protein {Ar... 30 6.0
TC91246 similar to GP|13374872|emb|CAC34506. mannosyltransferase... 30 6.0
AL370194 homologue to GP|21952816|db putative chromodomain-helic... 29 7.8
BG449406 similar to PIR|T04629|T04 hypothetical protein F10N7.30... 29 7.8
TC85119 similar to PIR|G84590|G84590 probable heat shock protein... 29 7.8
TC89445 similar to GP|7417006|gb|AAF62403.1| harpin inducing pro... 29 7.8
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 119 bits (299), Expect = 4e-27
Identities = 62/202 (30%), Positives = 105/202 (51%), Gaps = 3/202 (1%)
Frame = +2
Query: 613 REASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAG 672
++ Y + LY++ ++CV E+ I+ H A H K+ +AG
Sbjct: 5 KDVRRYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAG 184
Query: 673 FYWPTLRKDCMDFVKQCKECQVFADLS---KAPPKELVTMSAPWPFAMWGVDLVGPFPTA 729
F+WPT+ KD F+ +C CQ ++S + P ++ + F +WG+D +GPFP++
Sbjct: 185 FWWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEV---FDVWGIDFMGPFPSS 355
Query: 730 RAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSS 789
K+ILVAVDY +KW+EA + +V + I RFG+PR ++SD G+ F +
Sbjct: 356 YNN-KYILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINK 532
Query: 790 QTREFCKEMGIQMRFASVEHPQ 811
+ K+ G++ + A+ HPQ
Sbjct: 533 VFEKLLKKNGVRHKVATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 686
Score = 77.4 bits (189), Expect(2) = 2e-26
Identities = 34/70 (48%), Positives = 51/70 (72%)
Frame = -2
Query: 773 GIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLA 832
G+P IV+DNG+ F S++ REFC+ I++ AS +PQ+NGQ E++N++I+ GL++RL
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 833 EAKGAWLDEL 842
KG W DEL
Sbjct: 502 LKKGCWADEL 473
Score = 61.2 bits (147), Expect(2) = 2e-26
Identities = 38/119 (31%), Positives = 64/119 (52%), Gaps = 6/119 (5%)
Frame = -3
Query: 845 VLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTW-RTR-PGFEEENQANMAVELDLL 902
VLWS+ T + T+ TPF M + V+AM P E++ + R+R P E N + L+ +
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNNDRLFNALETI 286
Query: 903 SETRDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVL----KWRSGAPGNKLTPNWEG 957
E RD+A +R + ++ + +N V+ + +++GD+VL + KL NWEG
Sbjct: 285 EERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNWEG 109
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 83.2 bits (204), Expect = 5e-16
Identities = 44/94 (46%), Positives = 62/94 (65%)
Frame = -2
Query: 462 AMVAIKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYL 521
A + + TNN AEYEA I G++ A +++I+ L I DS LV NQ+KG ++ NLI Y
Sbjct: 441 ARILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYR 262
Query: 522 ERVRYLMTLFQEVVVEYVPRAENQRADALAKLAS 555
+ R L+T F +V + ++PR ENQ ADALA L+S
Sbjct: 261 DYARRLLTYFTKVELHHIPRDENQMADALATLSS 160
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 82.8 bits (203), Expect = 6e-16
Identities = 78/315 (24%), Positives = 130/315 (40%), Gaps = 9/315 (2%)
Frame = +2
Query: 646 IMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKE 705
++ E H+ A H GR+ +++ F+WP + FV+ C C +A
Sbjct: 179 LVQESHDSTAAGH-PGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGF 355
Query: 706 LVTMSAPWPF-AMWGVDLVGPFPTARAQ-MKFILVAVDYFTKWIEAEPLAKITSAKIVNF 763
L + P + +D + P R + +++ V VD +K + E + + +
Sbjct: 356 LKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEACAQR 535
Query: 764 YWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVI 823
+ G+P++IVSD G+ + REFC+ G+ ++ HPQT+G E N+ I
Sbjct: 536 FLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEI 715
Query: 824 LRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYG--VDAMLPVEIDNFTW 881
LR + ++ W D LP V + S+ TPF + +G VD + VE
Sbjct: 716 QAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYHVDPIPTVE------ 877
Query: 882 RTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNS-RVRVREMQVGDL-- 938
G E +A + + + + A +QR A N R QVGD
Sbjct: 878 -DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADRYQVGDKVW 1054
Query: 939 --VLKWRSGAPGNKL 951
V ++S P KL
Sbjct: 1055LNVSNYKSPRPSKKL 1099
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 76.6 bits (187), Expect = 4e-14
Identities = 44/105 (41%), Positives = 65/105 (61%)
Frame = +3
Query: 455 EIVKASGAMVAIKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKD 514
EI+K S + A+NN+ YEALIAG++LA +KIR++ DSQLV +Q G ++ +D
Sbjct: 30 EILKQSFRL-EFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQFSGEYEARD 206
Query: 515 PNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKP 559
+ YL+ V+ L + +PR+EN +ADALA LAS+ P
Sbjct: 207 ELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASSSDP 341
>TC92306
Length = 521
Score = 44.7 bits (104), Expect = 2e-04
Identities = 28/73 (38%), Positives = 37/73 (50%)
Frame = +2
Query: 479 IAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEY 538
I GL AR + +R D Q V Q G+++V +PNL L + F+ V VE+
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 539 VPRAENQRADALA 551
VPR N ADA A
Sbjct: 182 VPRGXNXAADAQA 220
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 43.5 bits (101), Expect = 4e-04
Identities = 25/79 (31%), Positives = 39/79 (48%)
Frame = +2
Query: 469 TNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLM 528
T AEY AL+ GLK A + + + DS+LV NQ+ +++KD +L K L
Sbjct: 674 TKESAEYRALLLGLKHASMKGFKYVTAKGDSELVINQILDPWKIKDEHLKKLCAEALELS 853
Query: 529 TLFQEVVVEYVPRAENQRA 547
F ++++ R N A
Sbjct: 854 DNFHSFRIQHISRERNYGA 910
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 37.7 bits (86), Expect = 0.022
Identities = 27/107 (25%), Positives = 42/107 (39%), Gaps = 24/107 (22%)
Frame = +1
Query: 776 RAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEA- 834
+ +++DN +F SS EFC GI +PQ NG E R +L R L+ A
Sbjct: 289 KKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGVAERMIRTLLERARCMLSNAG 468
Query: 835 ----KGAWLD-------------------ELPAVLWSYNTTEQSTTR 858
+ W++ ++P +WS N + S R
Sbjct: 469 L*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGNLVDYSNLR 609
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 36.2 bits (82), Expect = 0.064
Identities = 28/113 (24%), Positives = 49/113 (42%)
Frame = +2
Query: 792 REFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNT 851
++ K G + +S HP ++GQ E+ N+ LR + W P + YNT
Sbjct: 44 KQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEYWYNT 223
Query: 852 TEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSE 904
+ + TPF+ YG D + + + T + Q+ +A +LLS+
Sbjct: 224 SYNISAAMTPFKALYGRDLSMLIRSKGSSKDT-----ADLQSQLAQREELLSQ 367
>BQ141183 weakly similar to GP|12654765|gb| Similar to erythroblast
macrophage protein {Homo sapiens}, partial (10%)
Length = 640
Score = 32.3 bits (72), Expect = 0.92
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = -2
Query: 292 ARWERCVLVPALIFSTRRTTLSKESARSWKRATR 325
A W C + P L+ +TR + S RSW+RA+R
Sbjct: 405 ALWLPCAVEPPLLAATRACASGRASWRSWRRASR 304
>TC86742 similar to PIR|T01541|T01541 hypothetical protein A_IG005I10.16 -
Arabidopsis thaliana, partial (19%)
Length = 2073
Score = 32.0 bits (71), Expect = 1.2
Identities = 22/85 (25%), Positives = 43/85 (49%)
Frame = -1
Query: 468 ATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYL 527
AT +AE+ A + ++ A E+ + ++ + TDS V N + +++ +R+
Sbjct: 411 ATPLEAEFCACMIAIEKAMELGLNNICLETDSLKVVNAFHKIVGIPWQMRVRWHNCIRFC 232
Query: 528 MTLFQEVVVEYVPRAENQRADALAK 552
++ V ++PR N ADALA+
Sbjct: 231 HSI--ACVCVHIPREGNLVADALAR 163
>BF631997 weakly similar to GP|18542925|gb Putative pol polyprotein {Oryza
sativa}, partial (6%)
Length = 650
Score = 30.4 bits (67), Expect = 3.5
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = -2
Query: 781 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESAN 820
+NG + S++ E CKE I ++ PQ NG E N
Sbjct: 544 NNGLEICSAEFNELCKEEHITRQYTVRNTPQKNGVAERMN 425
>TC93424
Length = 786
Score = 30.0 bits (66), Expect = 4.6
Identities = 14/37 (37%), Positives = 23/37 (61%), Gaps = 3/37 (8%)
Frame = -1
Query: 354 SNRPRNSC---IHGAMIMNNVGLGNLSLIASGRKLIW 387
+N P+ +C I+G ++ +V +GNL LI KL+W
Sbjct: 543 TNAPQRTCCFGIYGLILKQSVVIGNLKLIIGLWKLVW 433
>TC78354 similar to GP|13272425|gb|AAK17151.1 unknown protein {Arabidopsis
thaliana}, partial (92%)
Length = 1637
Score = 30.0 bits (66), Expect = 4.6
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = +2
Query: 37 IVWTWILRYERQEYSRLLLFHHHH 60
+V TW L+ + LLL HHHH
Sbjct: 326 LVVTWFLQQHHYQQQHLLLHHHHH 397
>TC89844 similar to GP|3258570|gb|AAC24380.1| Unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 680
Score = 29.6 bits (65), Expect = 6.0
Identities = 19/75 (25%), Positives = 32/75 (42%)
Frame = +1
Query: 689 CKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIE 748
C+ + AD PP+ + AP P + + GP P + K ++V + Y K+
Sbjct: 187 CQAITLVADPRALPPQPASSPHAPPPPSPYNHAPPGPLPNVHGRKKAVIVGISY--KFSR 360
Query: 749 AEPLAKITSAKIVNF 763
E I AK + +
Sbjct: 361 HELKGCINDAKCMRY 405
>TC91246 similar to GP|13374872|emb|CAC34506. mannosyltransferase-like
protein {Arabidopsis thaliana}, partial (19%)
Length = 678
Score = 29.6 bits (65), Expect = 6.0
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = +1
Query: 178 LRRQMR*SAGCFHRRSNRRRWHG 200
LRRQ + C HRR N +WHG
Sbjct: 193 LRRQKLLRSFCHHRRRNSHQWHG 261
>AL370194 homologue to GP|21952816|db putative
chromodomain-helicase-DNA-binding protein {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 418
Score = 29.3 bits (64), Expect = 7.8
Identities = 26/87 (29%), Positives = 41/87 (46%), Gaps = 2/87 (2%)
Frame = +1
Query: 454 KEIVKASGAMVAIKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVK 513
+E+VK MV I +T NQ E + R+ R D Q+ + Q G +
Sbjct: 109 EEMVK----MVVIASTTNQEEMVTID-----------RTHAKRGDIQVTKCQEMGRILLS 243
Query: 514 DPN--LIKYLERVRYLMTLFQEVVVEY 538
P LIK + RV++ + +FQ +VV +
Sbjct: 244 PPKSQLIKKISRVQFCVQIFQLLVVNF 324
>BG449406 similar to PIR|T04629|T04 hypothetical protein F10N7.30 -
Arabidopsis thaliana, partial (19%)
Length = 684
Score = 29.3 bits (64), Expect = 7.8
Identities = 12/38 (31%), Positives = 18/38 (46%)
Frame = +2
Query: 289 SRRARWERCVLVPALIFSTRRTTLSKESARSWKRATRR 326
+ R W CV++P+ +F + A SW TRR
Sbjct: 311 NHRTGWSYCVIIPSWVFVPKSKNSDPIVASSWYTITRR 424
>TC85119 similar to PIR|G84590|G84590 probable heat shock protein [imported]
- Arabidopsis thaliana, partial (72%)
Length = 796
Score = 29.3 bits (64), Expect = 7.8
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 2/39 (5%)
Frame = -2
Query: 34 GGLIVWTWILRYERQEYSR--LLLFHHHHLLHRLHR*DL 70
G L++ I Y Q + L+ +HHHHL H H+ DL
Sbjct: 207 GFLLLLQKIQTYHHQTSPQRTLISYHHHHLFHPCHQMDL 91
>TC89445 similar to GP|7417006|gb|AAF62403.1| harpin inducing protein
{Nicotiana tabacum}, partial (36%)
Length = 651
Score = 29.3 bits (64), Expect = 7.8
Identities = 21/53 (39%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Frame = +3
Query: 40 TWILRYERQEYSRLLLFHHHHLLHRLHR*DLWSAHQRIHLLRNNS---RR*HR 89
+W+LR Q + R L +HHH HR HR R+HLL + S R+ HR
Sbjct: 186 SWLLRLHFQLHLRFNLQNHHH-NHRHHR------DPRVHLLAHCST*RRQIHR 323
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.336 0.143 0.468
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,938,209
Number of Sequences: 36976
Number of extensions: 473621
Number of successful extensions: 4621
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 2594
Number of HSP's successfully gapped in prelim test: 173
Number of HSP's that attempted gapping in prelim test: 1701
Number of HSP's gapped (non-prelim): 3065
length of query: 993
length of database: 9,014,727
effective HSP length: 106
effective length of query: 887
effective length of database: 5,095,271
effective search space: 4519505377
effective search space used: 4519505377
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0590c.10


