

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
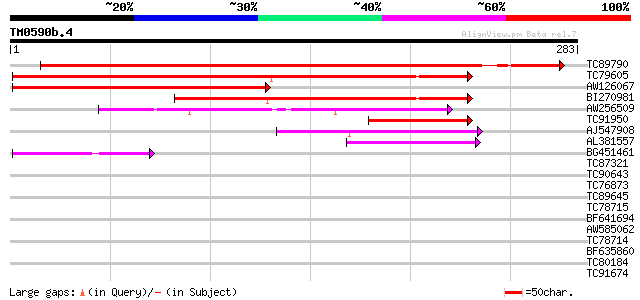
Query= TM0590b.4
(283 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89790 similar to PIR|T51519|T51519 cyclic nucleotide-gated cat... 322 1e-88
TC79605 similar to GP|14532660|gb|AAK64058.1 putative cyclic nuc... 204 3e-53
AW126067 similar to GP|7228242|emb cyclic nucleotide gated chann... 135 2e-32
BI270981 similar to PIR|T51354|T51 cyclic nucleotide-regulated i... 124 4e-29
AW256509 weakly similar to GP|8131898|gb| cyclic nucleotide-bind... 87 6e-18
TC91950 similar to PIR|T51354|T51354 cyclic nucleotide-regulated... 60 7e-10
AJ547908 weakly similar to GP|9294159|dbj| cyclic nucleotide and... 53 1e-07
AL381557 similar to GP|9294159|dbj| cyclic nucleotide and calmod... 52 3e-07
BG451461 weakly similar to GP|8131898|gb|A cyclic nucleotide-bin... 41 6e-04
TC87321 similar to PIR|C86148|C86148 hypothetical protein AAF784... 32 0.36
TC90643 weakly similar to GP|19386797|dbj|BAB86176. hypothetical... 31 0.61
TC76873 weakly similar to PIR|H96593|H96593 unknown protein 798... 30 0.80
TC89645 weakly similar to GP|21537016|gb|AAM61357.1 unknown {Ara... 30 0.80
TC78715 weakly similar to GP|15487382|gb|AAK98797.1 BONZAI1 {Ara... 30 1.0
BF641694 similar to GP|438387|gb|A alpha-amylase {Ostrinia nubil... 29 1.8
AW585062 28 4.0
TC78714 weakly similar to GP|15487382|gb|AAK98797.1 BONZAI1 {Ara... 27 6.8
BF635860 weakly similar to PIR|T01215|T012 protein kinase homolo... 27 6.8
TC80184 similar to PIR|C86148|C86148 hypothetical protein AAF784... 27 6.8
TC91674 27 8.9
>TC89790 similar to PIR|T51519|T51519 cyclic nucleotide-gated cation channel
- Arabidopsis thaliana, partial (33%)
Length = 1146
Score = 322 bits (824), Expect = 1e-88
Identities = 156/262 (59%), Positives = 197/262 (74%)
Frame = +1
Query: 16 RMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLD 75
R R++EWWMR+RQLP LRQRVR++ERQRW AM G DE ++I++LP+GLRRDIK HLCLD
Sbjct: 1 RCRDMEWWMRRRQLPSRLRQRVRHFERQRWAAMGGEDEMELIKDLPEGLRRDIKRHLCLD 180
Query: 76 LVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSSQDLRD 135
L+++VPLF ++DDL+L+NICDRVK LVF++ E I REGDPV RM+F+VRG ++ +Q L
Sbjct: 181 LIKKVPLFHNLDDLILDNICDRVKPLVFSRDEKIIREGDPVPRMVFIVRGRIKRNQSLSK 360
Query: 136 GVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQH 195
G+ + L PG F GDELLSWCLRRPFI+RLP SSAT V LE+TEAFGL+A+++RY+T H
Sbjct: 361 GIVATSVLEPGGFLGDELLSWCLRRPFIDRLPASSATFVCLESTEAFGLDAQNLRYITDH 540
Query: 196 FRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLSFIRPRRPASRCTSM 255
FRY F E++KR+ARYYS WRTWAAV IQLA+RRY+ R P P R
Sbjct: 541 FRYKFANERLKRTARYYSSNWRTWAAVNIQLAYRRYRQRTR-------GPVTPV-RDNGG 696
Query: 256 EEDRLRLYTALLTSPKPNDQEE 277
E RL Y A+ S +P+D E
Sbjct: 697 TERRLMQYAAMFMSIRPHDHLE 762
>TC79605 similar to GP|14532660|gb|AAK64058.1 putative cyclic nucleotide and
calmodulin-regulated ion channel protein {Arabidopsis
thaliana}, partial (65%)
Length = 1371
Score = 204 bits (519), Expect = 3e-53
Identities = 104/232 (44%), Positives = 150/232 (63%), Gaps = 2/232 (0%)
Frame = +2
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LP+ LR+RVR Y++ +W A RGVDE ++++LP
Sbjct: 188 YLQSLTLRLEEMRVKRRDSEQWMHHRLLPKELRERVRRYDQYKWLATRGVDEDILVQSLP 367
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL LVR+VPLF+ MD+ +L+ IC+R+K +FT+ I REGDPV MLF
Sbjct: 368 KDLRRDIKRHLCLALVRRVPLFESMDERLLDAICERLKPCLFTENTYIVREGDPVDEMLF 547
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L F G+ELL+W L LP S+ T+ L
Sbjct: 548 IIRGRLESVTTDGGRSGFFNRTYLKEAEFCGEELLTWALDPRSGSNLPTSTRTVKALTEV 727
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRY 231
E F L A+++++V FR ++ V+ + R+YS WRTWAA IQ AWRRY
Sbjct: 728 ETFALTADELKFVASQFRRLHSRQ-VQHTFRFYSQQWRTWAACFIQAAWRRY 880
>AW126067 similar to GP|7228242|emb cyclic nucleotide gated channel (CNGC4)
like protein {Arabidopsis thaliana}, partial (21%)
Length = 451
Score = 135 bits (339), Expect = 2e-32
Identities = 58/129 (44%), Positives = 96/129 (73%)
Frame = +2
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L +TT + + M+++ R+ E WM R LP ++QR+R YE+ +W RGV+E ++I NLP
Sbjct: 29 YLQSTTVRVEEMRIKRRDAEQWMCHRMLPDHMKQRIRRYEQYKWQENRGVEEEKLICNLP 208
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDI+ HLCLDL+++VP+F +MD +L+ +CD++K +++T+ + REGDPV MLF
Sbjct: 209 KDLRRDIRRHLCLDLLKKVPIFSNMDKQLLDAMCDKLKPVLYTEKSFVVREGDPVDEMLF 388
Query: 122 VVRGHLQSS 130
++RG L+++
Sbjct: 389 IMRGKLKTA 415
>BI270981 similar to PIR|T51354|T51 cyclic nucleotide-regulated ion channel 1
[validated] - Arabidopsis thaliana, partial (20%)
Length = 455
Score = 124 bits (311), Expect = 4e-29
Identities = 66/151 (43%), Positives = 94/151 (61%), Gaps = 2/151 (1%)
Frame = +2
Query: 83 FQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHL--QSSQDLRDGVKSY 140
F+ MD+ +L+ +CD +K +++TK I REGDPV MLF++RG L ++ R G +
Sbjct: 2 FEKMDEQLLDAVCDCLKPVLYTKESCIVREGDPVDEMLFIMRGKLLTVTTNGGRTGFFNS 181
Query: 141 CTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTF 200
L G+F G+ELL+W L LP S+ T+ TL EAF L+AED+++V FR
Sbjct: 182 EYLKAGDFCGEELLTWALDPRSASNLPISTRTVQTLSEVEAFALKAEDLKFVASQFRRLH 361
Query: 201 MKEKVKRSARYYSPGWRTWAAVAIQLAWRRY 231
K+ ++ + R+YS WRTWAA IQ AWRRY
Sbjct: 362 SKQ-LRHTFRFYSQQWRTWAACFIQAAWRRY 451
>AW256509 weakly similar to GP|8131898|gb| cyclic nucleotide-binding
transporter 1 {Arabidopsis thaliana}, partial (21%)
Length = 566
Score = 87.4 bits (215), Expect = 6e-18
Identities = 61/189 (32%), Positives = 95/189 (49%), Gaps = 12/189 (6%)
Frame = +2
Query: 45 WTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQVPLFQHMDD--LVLENICDRVKSLV 102
W A RGV E ++ NLP+ L+ DI+ HL V+ + F MD+ +L I +R+
Sbjct: 11 WAATRGVSEKMVLENLPEDLQTDIRRHL-FKFVKNIRFFSLMDEDEPILYAIRERLVQTT 187
Query: 103 FTKGETITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRP- 161
+ +G + +G +++M+F+VRG ++S +D + L G+ SG+ELL+W L +
Sbjct: 188 YIEGSIVFSQGGLIQKMVFIVRGKMESIG--KDEIP--VLLSEGDASGEELLTWYLEQTA 355
Query: 162 ---------FIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYY 212
R S T L EAF L+A+D+ VT F V++ RY
Sbjct: 356 *SKDGKKVKIRGRGLTSDRTGKCLTNVEAFSLDAKDIEEVTTLFSRFLRSPHVQQVIRYQ 535
Query: 213 SPGWRTWAA 221
SP WR+ AA
Sbjct: 536 SPYWRSLAA 562
>TC91950 similar to PIR|T51354|T51354 cyclic nucleotide-regulated ion
channel 1 [validated] - Arabidopsis thaliana, partial
(15%)
Length = 656
Score = 60.5 bits (145), Expect = 7e-10
Identities = 25/52 (48%), Positives = 36/52 (69%)
Frame = +1
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRY 231
EAF L+A+D+++V FR ++++R+ R YSP W+TW A IQ AWRRY
Sbjct: 1 EAFALKADDLKFVASQFRRLINSKQLQRTFRSYSPQWKTWGACFIQAAWRRY 156
>AJ547908 weakly similar to GP|9294159|dbj| cyclic nucleotide and
calmodulin-regulated ion channel protein-like
{Arabidopsis thaliana}, partial (12%)
Length = 523
Score = 53.1 bits (126), Expect = 1e-07
Identities = 38/113 (33%), Positives = 51/113 (44%), Gaps = 10/113 (8%)
Frame = -1
Query: 134 RDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPP----------SSATLVTLETTEAFG 183
R G S L G+ G+ELL W L + + S T+ L EAF
Sbjct: 502 RRGWNSRFPLSEGDACGEELLRWYLEQSSESKEGKKIKLHGQGLTSDRTVRCLTNVEAFS 323
Query: 184 LEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLS 236
L A+D+ VT + +V+ RY S WR+ AA I++AWR K RLS
Sbjct: 322 LRAKDIEEVTTLYARFLRSPRVRGVIRYESTYWRSLAANRIRVAWRYRKKRLS 164
>AL381557 similar to GP|9294159|dbj| cyclic nucleotide and
calmodulin-regulated ion channel protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 524
Score = 52.0 bits (123), Expect = 3e-07
Identities = 29/67 (43%), Positives = 36/67 (53%)
Frame = +2
Query: 169 SSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAW 228
S+ T++ L EAF L A D+ VT F V+ + RY SP WR AA IQ+AW
Sbjct: 23 SNRTVLCLTNVEAFSLRAADLEEVTSLFARFLRSPGVQGAIRYESPYWRCLAATRIQVAW 202
Query: 229 RRYKHRL 235
R K RL
Sbjct: 203 RYRKKRL 223
>BG451461 weakly similar to GP|8131898|gb|A cyclic nucleotide-binding
transporter 1 {Arabidopsis thaliana}, partial (6%)
Length = 671
Score = 40.8 bits (94), Expect = 6e-04
Identities = 22/71 (30%), Positives = 35/71 (48%)
Frame = +2
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
FL + + M R ++E WM +LP+ LR RV E W + E + NLP
Sbjct: 431 FLRTLSKRSLEMMHRGCDVEQWMNHHRLPEDLRMRVLQAE---WNSRHNAPERTXLENLP 601
Query: 62 QGLRRDIKYHL 72
+ L+ DI+ ++
Sbjct: 602 ENLQIDIRRYI 634
>TC87321 similar to PIR|C86148|C86148 hypothetical protein AAF78401.1
[imported] - Arabidopsis thaliana, partial (70%)
Length = 1413
Score = 31.6 bits (70), Expect = 0.36
Identities = 22/75 (29%), Positives = 32/75 (42%), Gaps = 8/75 (10%)
Frame = +2
Query: 80 VPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQ--------SSQ 131
VPL Q + L NI V + GE + REG+P + + F+ G + +
Sbjct: 167 VPLLQRLPRSSLRNISQLVIVKNYEPGEYVVREGEPGEGLYFIWEGEAEVIGSGLADDEK 346
Query: 132 DLRDGVKSYCTLGPG 146
DL +K Y G G
Sbjct: 347 DLEFQLKRYDYFGFG 391
>TC90643 weakly similar to GP|19386797|dbj|BAB86176. hypothetical
protein~similar to Arabidopsis thaliana protein
T1F15.13, partial (21%)
Length = 771
Score = 30.8 bits (68), Expect = 0.61
Identities = 19/65 (29%), Positives = 32/65 (49%)
Frame = +3
Query: 183 GLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLSF 242
G + +R V + ++T M +VK + +Y + A+ + WR+ KH+L LS
Sbjct: 75 GYMVDQLRKVPKQ-KWTEMWRQVKNISHHYEFQYPPKREDAVNMLWRQVKHKLPGVRLSI 251
Query: 243 IRPRR 247
R RR
Sbjct: 252 HRTRR 266
>TC76873 weakly similar to PIR|H96593|H96593 unknown protein 79801-80364
[imported] - Arabidopsis thaliana, partial (39%)
Length = 1445
Score = 30.4 bits (67), Expect = 0.80
Identities = 14/30 (46%), Positives = 20/30 (66%), Gaps = 2/30 (6%)
Frame = +1
Query: 187 EDVRYVTQHFRYTFM--KEKVKRSARYYSP 214
EDV++VT HF Y + KE++K + YY P
Sbjct: 1306 EDVQFVTFHFEYIHL*KKEELKSMSMYYLP 1395
>TC89645 weakly similar to GP|21537016|gb|AAM61357.1 unknown {Arabidopsis
thaliana}, partial (83%)
Length = 996
Score = 30.4 bits (67), Expect = 0.80
Identities = 14/44 (31%), Positives = 25/44 (56%)
Frame = +2
Query: 109 ITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDE 152
+ +E PVK+ ++ + ++ L D V +CT+G GNF+ E
Sbjct: 95 LVKEEIPVKQQPLLLSDNFKTGLVLVDLVNGFCTVGSGNFAPKE 226
>TC78715 weakly similar to GP|15487382|gb|AAK98797.1 BONZAI1 {Arabidopsis
thaliana}, partial (31%)
Length = 924
Score = 30.0 bits (66), Expect = 1.0
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -2
Query: 25 RKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRR 66
R + G + RNY R+RW ++ + R LP G RR
Sbjct: 314 RSERRDLGSHET*RNYRRRRWKCFPDSEDLRQFRRLPCGHRR 189
>BF641694 similar to GP|438387|gb|A alpha-amylase {Ostrinia nubilalis},
partial (28%)
Length = 617
Score = 29.3 bits (64), Expect = 1.8
Identities = 19/56 (33%), Positives = 25/56 (43%), Gaps = 9/56 (16%)
Frame = +1
Query: 135 DGVKSYCT--LGPGNFSG-------DELLSWCLRRPFIERLPPSSATLVTLETTEA 181
D + + C LGP F G + ++ W RP+ ER P S LVT EA
Sbjct: 139 DDIANECETFLGPRGFGGIQISPPNENVVLWSYNRPWWERYQPMSYKLVTRSGNEA 306
>AW585062
Length = 528
Score = 28.1 bits (61), Expect = 4.0
Identities = 20/51 (39%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Frame = -3
Query: 134 RDGVKSYCTLGPGNFSGDELLSWCLRRPFIER--LPPSSATLVTLETTEAF 182
R+G+ Y F GD+LL+ CL P + R LP TL +L TE+F
Sbjct: 214 REGLHRYLCR*LSGFYGDDLLTICLPFPALRRSVLP---FTLRSLRITESF 71
>TC78714 weakly similar to GP|15487382|gb|AAK98797.1 BONZAI1 {Arabidopsis
thaliana}, partial (30%)
Length = 776
Score = 27.3 bits (59), Expect = 6.8
Identities = 12/39 (30%), Positives = 18/39 (45%)
Frame = -2
Query: 25 RKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQG 63
R + G + RNY R+RW ++ + R LP G
Sbjct: 121 RSERRDLGSHET*RNYRRRRWKCFPDSEDLRQFRRLPCG 5
>BF635860 weakly similar to PIR|T01215|T012 protein kinase homolog F6N23.2 -
Arabidopsis thaliana, partial (40%)
Length = 535
Score = 27.3 bits (59), Expect = 6.8
Identities = 15/65 (23%), Positives = 29/65 (44%)
Frame = +2
Query: 75 DLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSSQDLR 134
+ + VPL Q + + I + V + +G+ + REG+P F+ G + +
Sbjct: 74 EFLGSVPLLQRLPSSSIRKISELVIVKHYERGDYVAREGEPGDGTYFIWDGEAKVVGSVS 253
Query: 135 DGVKS 139
D +S
Sbjct: 254 DNDES 268
>TC80184 similar to PIR|C86148|C86148 hypothetical protein AAF78401.1
[imported] - Arabidopsis thaliana, partial (26%)
Length = 843
Score = 27.3 bits (59), Expect = 6.8
Identities = 15/65 (23%), Positives = 29/65 (44%)
Frame = +3
Query: 75 DLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSSQDLR 134
+ + VPL Q + + I + V + +G+ + REG+P F+ G + +
Sbjct: 63 EFLGSVPLLQRLPSSSIRKISELVIVKHYERGDYVAREGEPGDGTYFIWDGEAKVVGSVS 242
Query: 135 DGVKS 139
D +S
Sbjct: 243 DNDES 257
>TC91674
Length = 747
Score = 26.9 bits (58), Expect = 8.9
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = -2
Query: 226 LAWRRYKHRLSLTSLSFIRPRRPASRCTSMEEDRLRLYTALLTSPKP 272
L + RY HR L +R RR R TS + LRL+ + P+P
Sbjct: 218 LRYDRYGHRRHL-----VRDRRRPGRSTSAADRALRLWR*TSSHPRP 93
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,180,091
Number of Sequences: 36976
Number of extensions: 129388
Number of successful extensions: 858
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 849
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 851
length of query: 283
length of database: 9,014,727
effective HSP length: 95
effective length of query: 188
effective length of database: 5,502,007
effective search space: 1034377316
effective search space used: 1034377316
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0590b.4


