

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.5
(191 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
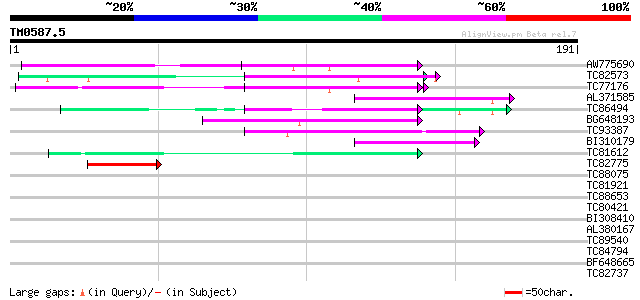
Score E
Sequences producing significant alignments: (bits) Value
AW775690 similar to GP|8567776|gb|A hypothetical protein {Arabid... 53 6e-08
TC82573 similar to PIR|T09602|T09602 probable zinc finger protei... 51 2e-07
TC77176 homologue to GP|7228329|emb|CAB77055.1 putative TFIIIA (... 50 4e-07
AL371585 similar to PIR|S69193|S691 probable finger protein Pszf... 49 1e-06
TC86494 homologue to GP|7228329|emb|CAB77055.1 putative TFIIIA (... 47 3e-06
BG648193 similar to GP|2346976|dbj| ZPT2-13 {Petunia x hybrida},... 47 5e-06
TC93387 similar to GP|1786136|dbj|BAA19111.1 PEThy;ZPT2-6 {Petun... 46 1e-05
BI310179 similar to PIR|T52381|T523 zinc finger protein Zat1 C2... 45 2e-05
TC81612 similar to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia x ... 44 3e-05
TC82775 similar to PIR|T52382|T52382 zinc finger protein ZPT4-4 ... 40 4e-04
TC88075 homologue to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia ... 39 0.001
TC81921 homologue to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia ... 39 0.002
TC88653 similar to GP|1786134|dbj|BAA19110.1 PEThy;ZPT2-5 {Petun... 39 0.002
TC80421 similar to PIR|D96707|D96707 probable zinc finger protei... 38 0.003
BI308410 similar to PIR|S55882|S558 CCHH finger protein 2 - Arab... 37 0.005
AL380167 homologue to PIR|S55887|S558 CCHH finger protein 7 - Ar... 34 0.040
TC89540 similar to PIR|B96686|B96686 probable C2H2-type zinc fin... 33 0.068
TC84794 similar to PIR|S54156|S54156 extensin-like protein - cow... 33 0.088
BF648665 weakly similar to GP|15294296|gb AT3g56820/T8M16_150 {A... 33 0.088
TC82737 homologue to PIR|B96686|B96686 probable C2H2-type zinc f... 31 0.34
>AW775690 similar to GP|8567776|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (20%)
Length = 631
Score = 53.1 bits (126), Expect = 6e-08
Identities = 39/135 (28%), Positives = 58/135 (42%)
Frame = +2
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRP 64
++H H + + S + +C C + F S +AL GH H + P K
Sbjct: 236 NTHVSNHRDHNNNKSSNFYLYECKTCNRCFPSFQALGGHRASHTK--------PNKANCN 391
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLR 124
Q Q T+ + ++ +L A A ++ + I + H+CSIC
Sbjct: 392 VAQQKQAVTTSSFVDDHYDPTMNTILSLQAFA-----ATPITTIPTTKKSKVHECSICGA 556
Query: 125 VFSSGQALGGHKRRH 139
FSSGQALGGH RRH
Sbjct: 557 EFSSGQALGGHMRRH 601
Score = 40.8 bits (94), Expect = 3e-04
Identities = 25/67 (37%), Positives = 38/67 (56%), Gaps = 6/67 (8%)
Frame = +2
Query: 79 QEEHEVAACLLLLANA-NAKGDTCSSSKS-----EISCSSSHHHHKCSICLRVFSSGQAL 132
+EE ++A CL+LLA N + D ++ S + SS+ + ++C C R F S QAL
Sbjct: 164 EEEEDLANCLILLARGHNHRHDYQNTHVSNHRDHNNNKSSNFYLYECKTCNRCFPSFQAL 343
Query: 133 GGHKRRH 139
GGH+ H
Sbjct: 344 GGHRASH 364
>TC82573 similar to PIR|T09602|T09602 probable zinc finger protein SCOF-1
cold-inducible - soybean, partial (65%)
Length = 857
Score = 51.2 bits (121), Expect = 2e-07
Identities = 40/147 (27%), Positives = 59/147 (39%), Gaps = 9/147 (6%)
Frame = +3
Query: 4 SHSHTQKH----VVPSPSSSPSGKWS-----QCSECGKKFWSEKALHGHMRCHPEREWRG 54
+ HT +H + P P++ + + +CS C K+F S +AL GH H + G
Sbjct: 189 ARGHTNRHDFNPLNPPPTTIDNNNNNTKLSYKCSVCNKEFSSYQALGGHKASHRKNSVGG 368
Query: 55 INPPPKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSH 114
++H + ++AN G S
Sbjct: 369 GG-----------------------DDHPSTSSAATTSSANTNGGGVRS----------- 446
Query: 115 HHHKCSICLRVFSSGQALGGHKRRHWE 141
H+CSIC R F +GQALGGHKR H+E
Sbjct: 447 --HECSICHRSFPTGQALGGHKRCHYE 521
Score = 41.6 bits (96), Expect = 2e-04
Identities = 25/75 (33%), Positives = 39/75 (51%), Gaps = 9/75 (12%)
Frame = +3
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHH-----HKCSICLRVFSSGQALGG 134
EE +A CL++LA + + + ++++ +KCS+C + FSS QALGG
Sbjct: 153 EEEYLALCLIMLARGHTNRHDFNPLNPPPTTIDNNNNNTKLSYKCSVCNKEFSSYQALGG 332
Query: 135 HKRRHWEK----GGD 145
HK H + GGD
Sbjct: 333 HKASHRKNSVGGGGD 377
Score = 31.6 bits (70), Expect = 0.20
Identities = 16/49 (32%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Frame = +3
Query: 3 NSHSHTQKHVVPSPSSSPSG--KWSQCSECGKKFWSEKALHGHMRCHPE 49
+ H T S +++ G + +CS C + F + +AL GH RCH E
Sbjct: 375 DDHPSTSSAATTSSANTNGGGVRSHECSICHRSFPTGQALGGHKRCHYE 521
>TC77176 homologue to GP|7228329|emb|CAB77055.1 putative TFIIIA (or
kruppel)-like zinc finger protein {Medicago sativa},
partial (96%)
Length = 1005
Score = 50.4 bits (119), Expect = 4e-07
Identities = 37/139 (26%), Positives = 57/139 (40%)
Frame = +2
Query: 3 NSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFR 62
N+ + T+ VP+P ++ +CS C K F S +AL GH H +
Sbjct: 302 NNDNKTESVPVPAPLTTVKLS-HKCSVCNKAFSSYQALGGHKASHRKAVM---------- 448
Query: 63 RPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSIC 122
T + + ++ + + +N K T H+CSIC
Sbjct: 449 ----------SATTVEDQTTTTSSAVTTSSASNGKNKT----------------HECSIC 550
Query: 123 LRVFSSGQALGGHKRRHWE 141
+ F +GQALGGHKR H+E
Sbjct: 551 HKSFPTGQALGGHKRCHYE 607
Score = 47.8 bits (112), Expect = 3e-06
Identities = 25/61 (40%), Positives = 34/61 (54%), Gaps = 1/61 (1%)
Frame = +2
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKS-EISCSSSHHHHKCSICLRVFSSGQALGGHKRR 138
EE +A CL++LA + D + S ++ HKCS+C + FSS QALGGHK
Sbjct: 251 EEEYLALCLIMLARSGNNNDNKTESVPVPAPLTTVKLSHKCSVCNKAFSSYQALGGHKAS 430
Query: 139 H 139
H
Sbjct: 431 H 433
>AL371585 similar to PIR|S69193|S691 probable finger protein Pszf1 - garden
pea, partial (19%)
Length = 497
Score = 48.9 bits (115), Expect = 1e-06
Identities = 27/56 (48%), Positives = 31/56 (55%), Gaps = 2/56 (3%)
Frame = +1
Query: 117 HKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAPAPV--LDLNLPPPE 170
H+CSIC F+SGQALGGH RRH + P P + LDLNLP PE
Sbjct: 70 HECSICGSEFTSGQALGGHMRRHRVSVANAAAVAAPDERVRPRNILQLDLNLPAPE 237
Score = 32.7 bits (73), Expect = 0.088
Identities = 15/37 (40%), Positives = 21/37 (56%)
Frame = +1
Query: 11 HVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCH 47
H + + + K +CS CG +F S +AL GHMR H
Sbjct: 28 HGLKAAINGNKAKIHECSICGSEFTSGQALGGHMRRH 138
>TC86494 homologue to GP|7228329|emb|CAB77055.1 putative TFIIIA (or
kruppel)-like zinc finger protein {Medicago sativa},
partial (39%)
Length = 1092
Score = 47.4 bits (111), Expect = 3e-06
Identities = 50/165 (30%), Positives = 64/165 (38%), Gaps = 13/165 (7%)
Frame = +2
Query: 18 SSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVI 77
SSP +CS C K F S +AL GH H R S + Q TTV
Sbjct: 287 SSPLKLNHKCSVCNKAFPSYQALGGHKASH---------------RKSSSENQ--STTV- 412
Query: 78 TQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKR 137
+T S S S + H+CSIC + F +GQALGGHKR
Sbjct: 413 --------------------NETISVSVS------TSKMHECSICHKSFPTGQALGGHKR 514
Query: 138 RHWEKGGDLIHHH---------LPPPAPAPAPV----LDLNLPPP 169
H+E + H+H + A + + DLNLP P
Sbjct: 515 CHYEGVINNNHNHNNSNSSGITVSDAGAASSSISHRGFDLNLPAP 649
Score = 47.4 bits (111), Expect = 3e-06
Identities = 24/60 (40%), Positives = 34/60 (56%)
Frame = +2
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
EE +A CL++L+ +N ++I S +HKCS+C + F S QALGGHK H
Sbjct: 227 EEEYLALCLIMLSQSN----------NQIQSSPLKLNHKCSVCNKAFPSYQALGGHKASH 376
Score = 33.5 bits (75), Expect = 0.052
Identities = 20/50 (40%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Frame = +2
Query: 2 RNSHSHTQKHVVPSPSSSP--SGKWSQCSECGKKFWSEKALHGHMRCHPE 49
R S S Q V S + K +CS C K F + +AL GH RCH E
Sbjct: 377 RKSSSENQSTTVNETISVSVSTSKMHECSICHKSFPTGQALGGHKRCHYE 526
>BG648193 similar to GP|2346976|dbj| ZPT2-13 {Petunia x hybrida}, partial
(43%)
Length = 629
Score = 47.0 bits (110), Expect = 5e-06
Identities = 28/87 (32%), Positives = 38/87 (43%), Gaps = 13/87 (14%)
Frame = +2
Query: 66 QPQPQPPQTTVITQEEHEVAACLLLLANANA-------------KGDTCSSSKSEISCSS 112
QPQ P E+E C + A +GD ++ + +S +
Sbjct: 173 QPQNNKPNQKSFAPVEYECKTCNKKFPSFQALGGHRASHKRSKLEGDELLTNSTSLSLGN 352
Query: 113 SHHHHKCSICLRVFSSGQALGGHKRRH 139
H+CSIC + FS GQALGGH RRH
Sbjct: 353 KPKMHECSICGQNFSLGQALGGHMRRH 433
Score = 30.0 bits (66), Expect = 0.57
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = +2
Query: 23 KWSQCSECGKKFWSEKALHGHMRCH 47
K +CS CG+ F +AL GHMR H
Sbjct: 359 KMHECSICGQNFSLGQALGGHMRRH 433
>TC93387 similar to GP|1786136|dbj|BAA19111.1 PEThy;ZPT2-6 {Petunia x
hybrida}, partial (17%)
Length = 650
Score = 45.8 bits (107), Expect = 1e-05
Identities = 30/95 (31%), Positives = 41/95 (42%), Gaps = 14/95 (14%)
Frame = +1
Query: 80 EEHEVAACLLLLA--------------NANAKGDTCSSSKSEISCSSSHHHHKCSICLRV 125
EE ++A CL+LLA N KG + E S + ++C C R
Sbjct: 319 EEEDLAKCLILLAQGGNHREDGGVVDENKRVKGSHGNKKIGETSTKLGLYIYECKTCNRT 498
Query: 126 FSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAPAP 160
F S QALGGH+ H +K + PP P+ P
Sbjct: 499 FPSFQALGGHRASH-KKPKIMAEEKKPPSPPSQQP 600
Score = 29.6 bits (65), Expect = 0.75
Identities = 17/60 (28%), Positives = 28/60 (46%)
Frame = +1
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C C + F S +AL GH H + + P P QP+P ++ +Q ++ VA
Sbjct: 475 ECKTCNRTFPSFQALGGHRASHKKPKIMAEEKKPP--SPPSQQPRPQSSSHDSQSDNLVA 648
>BI310179 similar to PIR|T52381|T523 zinc finger protein Zat1 C2H2-type
[imported] - Arabidopsis thaliana, partial (16%)
Length = 704
Score = 45.1 bits (105), Expect = 2e-05
Identities = 20/42 (47%), Positives = 23/42 (54%)
Frame = +2
Query: 117 HKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAP 158
HKC C+R FS+G+ALGGH R H PPP AP
Sbjct: 77 HKCKFCMRSFSNGRALGGHMRSHMMNLPKQTESPTPPPPRAP 202
>TC81612 similar to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia x hybrida},
partial (50%)
Length = 612
Score = 44.3 bits (103), Expect = 3e-05
Identities = 33/126 (26%), Positives = 45/126 (35%)
Frame = +3
Query: 14 PSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQ 73
P S +G++ +C C KKF S +AL GH H +
Sbjct: 162 PQQKSYENGEY-ECKTCNKKFSSFQALGGHRASHKRMKL--------------------- 275
Query: 74 TTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALG 133
A+G+ +S + H+CSIC FS GQALG
Sbjct: 276 ----------------------AEGEELKEQAKSLSLWNKPKMHECSICGMGFSLGQALG 389
Query: 134 GHKRRH 139
GH R+H
Sbjct: 390 GHMRKH 407
Score = 33.9 bits (76), Expect = 0.040
Identities = 21/62 (33%), Positives = 30/62 (47%)
Frame = +3
Query: 78 TQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKR 137
+ E + A CL+LL+ C KS + ++C C + FSS QALGGH+
Sbjct: 114 SNEGIDYANCLMLLS--------CPQQKSY-----ENGEYECKTCNKKFSSFQALGGHRA 254
Query: 138 RH 139
H
Sbjct: 255 SH 260
>TC82775 similar to PIR|T52382|T52382 zinc finger protein ZPT4-4 C2H2-type
[imported] - garden petunia, partial (17%)
Length = 1152
Score = 40.4 bits (93), Expect = 4e-04
Identities = 16/25 (64%), Positives = 19/25 (76%)
Frame = +3
Query: 27 CSECGKKFWSEKALHGHMRCHPERE 51
C ECGK F S +AL GHM+CH E+E
Sbjct: 408 CKECGKCFHSWQALFGHMKCHSEKE 482
Score = 31.6 bits (70), Expect = 0.20
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +3
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
H C C + F G++LGGH R H
Sbjct: 150 HACKFCSKSFPCGRSLGGHMRSH 218
Score = 31.6 bits (70), Expect = 0.20
Identities = 29/119 (24%), Positives = 48/119 (39%), Gaps = 3/119 (2%)
Frame = +3
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVAA 86
C C K F ++L GHMR H + + + Q T++ + +
Sbjct: 156 CKFCSKSFPCGRSLGGHMRSHINN-----------LSAEKAEHKEKQLIFSTKKNDSIGS 302
Query: 87 CLLLLANANAKGDTCSSSKSEISCSS--SHHHHK-CSICLRVFSSGQALGGHKRRHWEK 142
+ + + + + I+ SS S+ + K C C + F S QAL GH + H EK
Sbjct: 303 EAATNGSYGLRENPKKTWRKRIADSSEDSYVYDKFCKECGKCFHSWQALFGHMKCHSEK 479
>TC88075 homologue to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia x hybrida},
partial (38%)
Length = 674
Score = 39.3 bits (90), Expect = 0.001
Identities = 31/114 (27%), Positives = 42/114 (36%)
Frame = +3
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C C +KF S +AL GH H +
Sbjct: 147 ECKTCNRKFSSFQALGGHRASHKK------------------------------------ 218
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
L LL + A ++K ++ H+CSIC + F GQALGGH RRH
Sbjct: 219 --LKLLDSEEAHKVNIHNNKPKM--------HQCSICGQEFKLGQALGGHMRRH 350
Score = 32.0 bits (71), Expect = 0.15
Identities = 21/61 (34%), Positives = 31/61 (50%)
Frame = +3
Query: 79 QEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRR 138
+E ++A L+LL SS ++ + + S +C C R FSS QALGGH+
Sbjct: 60 KESIDLATTLMLL----------SSRITQQNKTYSPVEFECKTCNRKFSSFQALGGHRAS 209
Query: 139 H 139
H
Sbjct: 210 H 212
Score = 32.0 bits (71), Expect = 0.15
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = +3
Query: 9 QKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCH 47
+ H V ++ P K QCS CG++F +AL GHMR H
Sbjct: 240 EAHKVNIHNNKP--KMHQCSICGQEFKLGQALGGHMRRH 350
>TC81921 homologue to GP|2346978|dbj|BAA21923.1 ZPT2-14 {Petunia x hybrida},
partial (41%)
Length = 839
Score = 38.5 bits (88), Expect = 0.002
Identities = 16/23 (69%), Positives = 18/23 (77%)
Frame = +3
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
H+CSIC F+ GQALGGH RRH
Sbjct: 381 HECSICGLEFAIGQALGGHMRRH 449
Score = 37.4 bits (85), Expect = 0.004
Identities = 26/80 (32%), Positives = 39/80 (48%), Gaps = 7/80 (8%)
Frame = +3
Query: 84 VAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEK- 142
+A CL+LL+ + + + SS + S++ +C C R F S QALGGH+ H +
Sbjct: 159 MANCLMLLSRGSDQFEATYSSTT-----SNNRVFECKTCNRQFPSFQALGGHRASHKKPR 323
Query: 143 ------GGDLIHHHLPPPAP 156
G L+H PP P
Sbjct: 324 LMGENIDGQLLH---TPPKP 374
Score = 31.2 bits (69), Expect = 0.26
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +3
Query: 20 PSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRG 54
P K +CS CG +F +AL GHMR H G
Sbjct: 366 PKPKTHECSICGLEFAIGQALGGHMRRHRAANMNG 470
>TC88653 similar to GP|1786134|dbj|BAA19110.1 PEThy;ZPT2-5 {Petunia x
hybrida}, partial (59%)
Length = 866
Score = 38.5 bits (88), Expect = 0.002
Identities = 34/123 (27%), Positives = 47/123 (37%)
Frame = +1
Query: 17 SSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTV 76
SS+ + + +C C ++F S +AL GH R S +P+ + T
Sbjct: 127 SSNFNNRVFECKTCKRQFSSFQALGGH-------------------RASHKKPRLMEMTS 249
Query: 77 ITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHK 136
+ H +L + AK H CSIC F GQALGGH
Sbjct: 250 DGDDHHGS----ILTSTTKAKT------------------HACSICGLEFGIGQALGGHM 363
Query: 137 RRH 139
RRH
Sbjct: 364 RRH 372
Score = 32.3 bits (72), Expect = 0.12
Identities = 15/35 (42%), Positives = 23/35 (64%), Gaps = 2/35 (5%)
Frame = +1
Query: 107 EISCSSSHHHH--KCSICLRVFSSGQALGGHKRRH 139
+++ SS+ ++ +C C R FSS QALGGH+ H
Sbjct: 115 QVNYSSNFNNRVFECKTCKRQFSSFQALGGHRASH 219
Score = 30.8 bits (68), Expect = 0.34
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +1
Query: 17 SSSPSGKWSQCSECGKKFWSEKALHGHMRCHPERE 51
+S+ K CS CG +F +AL GHMR H E
Sbjct: 280 TSTTKAKTHACSICGLEFGIGQALGGHMRRHRRTE 384
>TC80421 similar to PIR|D96707|D96707 probable zinc finger protein T22E19.1
[imported] - Arabidopsis thaliana, partial (37%)
Length = 1217
Score = 37.7 bits (86), Expect = 0.003
Identities = 27/92 (29%), Positives = 39/92 (42%), Gaps = 9/92 (9%)
Frame = +2
Query: 93 NANAKGDTCSSS----KSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDLIH 148
N + G T S S S I+ +S ++C C R F++ QALGGH+ H ++ L
Sbjct: 164 NTSPSGSTESVSGDGGASSINPTSGDRKYECQYCCREFANSQALGGHQNAHKKERQQLKR 343
Query: 149 HHLPPPAPAPA-----PVLDLNLPPPESGSDP 175
L A P++ PP S P
Sbjct: 344 AQLQASRNAAVSFVRNPIISAFAPPHHLLSSP 439
>BI308410 similar to PIR|S55882|S558 CCHH finger protein 2 - Arabidopsis
thaliana, partial (46%)
Length = 492
Score = 37.0 bits (84), Expect = 0.005
Identities = 18/38 (47%), Positives = 20/38 (52%)
Frame = +3
Query: 102 SSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
SSS S S S C+ C R F S QALGGH+ H
Sbjct: 57 SSSSSSSSSSMEQRIFSCNYCQRKFYSSQALGGHQNAH 170
Score = 33.1 bits (74), Expect = 0.068
Identities = 18/48 (37%), Positives = 23/48 (47%), Gaps = 7/48 (14%)
Frame = +3
Query: 7 HTQKHVVPSPSSSPSGKWSQ-------CSECGKKFWSEKALHGHMRCH 47
H +V PSSS S S C+ C +KF+S +AL GH H
Sbjct: 27 HLNLDLVLEPSSSSSSSSSSMEQRIFSCNYCQRKFYSSQALGGHQNAH 170
>AL380167 homologue to PIR|S55887|S558 CCHH finger protein 7 - Arabidopsis
thaliana, partial (11%)
Length = 476
Score = 33.9 bits (76), Expect = 0.040
Identities = 15/48 (31%), Positives = 24/48 (49%)
Frame = +2
Query: 92 ANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
++ N++ + + +S + H C C R F S QALGGH+ H
Sbjct: 332 SDTNSEVLGADQAHNTVSAPTIHRVFSCHYCRRKFFSSQALGGHQNAH 475
Score = 27.7 bits (60), Expect = 2.8
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +2
Query: 27 CSECGKKFWSEKALHGHMRCH 47
C C +KF+S +AL GH H
Sbjct: 413 CHYCRRKFFSSQALGGHQNAH 475
>TC89540 similar to PIR|B96686|B96686 probable C2H2-type zinc finger protein
F15E12.19 [imported] - Arabidopsis thaliana, partial
(28%)
Length = 1639
Score = 33.1 bits (74), Expect = 0.068
Identities = 22/73 (30%), Positives = 35/73 (47%)
Frame = +1
Query: 67 PQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVF 126
PQ P T +T A L+ ++A+++ +++ S S + C+ C R F
Sbjct: 436 PQDLDPMTLDLTLNFSSSEAELMGSSDASSE---VGAAEVHASPSVTPRIFSCNYCKRKF 606
Query: 127 SSGQALGGHKRRH 139
S QALGGH+ H
Sbjct: 607 YSSQALGGHQNAH 645
Score = 30.8 bits (68), Expect = 0.34
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = +1
Query: 15 SPSSSPSGKWSQCSECGKKFWSEKALHGHMRCH 47
SPS +P + C+ C +KF+S +AL GH H
Sbjct: 553 SPSVTP--RIFSCNYCKRKFYSSQALGGHQNAH 645
>TC84794 similar to PIR|S54156|S54156 extensin-like protein - cowpea
(fragment), partial (49%)
Length = 478
Score = 32.7 bits (73), Expect = 0.088
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P+P P + PPP S S P
Sbjct: 379 PPPSPSPPPPYEYKSPPPPSSSPP 450
>BF648665 weakly similar to GP|15294296|gb AT3g56820/T8M16_150 {Arabidopsis
thaliana}, partial (48%)
Length = 647
Score = 32.7 bits (73), Expect = 0.088
Identities = 20/52 (38%), Positives = 24/52 (45%)
Frame = +2
Query: 138 RHWEKGGDLIHHHLPPPAPAPAPVLDLNLPPPESGSDPITVPLDFAALDLSL 189
+H E +HH PPP P P LPP +GS P + F L LSL
Sbjct: 56 QHREDQSKSAYHHPPPPPPPP-------LPPCSNGSAPPSSLHSFKDLTLSL 190
>TC82737 homologue to PIR|B96686|B96686 probable C2H2-type zinc finger
protein F15E12.19 [imported] - Arabidopsis thaliana,
partial (18%)
Length = 1006
Score = 30.8 bits (68), Expect = 0.34
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = +2
Query: 119 CSICLRVFSSGQALGGHKRRH 139
C+ C R F S QALGGH+ H
Sbjct: 419 CNYCRRKFYSSQALGGHQNAH 481
Score = 28.9 bits (63), Expect = 1.3
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +2
Query: 27 CSECGKKFWSEKALHGHMRCH 47
C+ C +KF+S +AL GH H
Sbjct: 419 CNYCRRKFYSSQALGGHQNAH 481
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,807,788
Number of Sequences: 36976
Number of extensions: 278016
Number of successful extensions: 5265
Number of sequences better than 10.0: 225
Number of HSP's better than 10.0 without gapping: 3047
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4934
length of query: 191
length of database: 9,014,727
effective HSP length: 91
effective length of query: 100
effective length of database: 5,649,911
effective search space: 564991100
effective search space used: 564991100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0587.5


