

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.12
(583 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
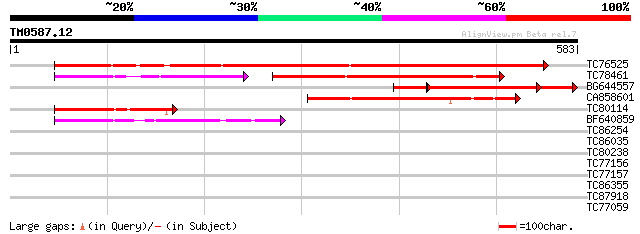
Sequences producing significant alignments: (bits) Value
TC76525 similar to GP|15450864|gb|AAK96703.1 Unknown protein {Ar... 612 e-175
TC78461 weakly similar to GP|22136978|gb|AAM91718.1 unknown prot... 270 e-113
BG644557 similar to GP|21321777|gb OSJNBa0031O09.02 {Oryza sativ... 200 3e-79
CA858601 weakly similar to GP|22136978|gb unknown protein {Arabi... 216 3e-56
TC80114 weakly similar to GP|6721151|gb|AAF26779.1| unknown prot... 190 1e-48
BF640859 weakly similar to GP|15450387|gb At2g44230/F4I1.4 {Arab... 165 5e-41
TC86254 similar to GP|14970841|emb|CAC44501. beta-galactosidase ... 32 0.52
TC86035 similar to PIR|E71435|E71435 probable triacylglycerol li... 31 1.5
TC80238 weakly similar to GP|14587270|dbj|BAB61188. putative rec... 30 2.0
TC77156 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-li... 30 3.4
TC77157 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-li... 28 7.5
TC86355 weakly similar to GP|9757805|dbj|BAB08323.1 CCR4-associa... 28 7.5
TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown prot... 28 9.8
TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 9.8
>TC76525 similar to GP|15450864|gb|AAK96703.1 Unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 2266
Score = 612 bits (1577), Expect = e-175
Identities = 294/509 (57%), Positives = 377/509 (73%), Gaps = 1/509 (0%)
Frame = +1
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G GFA G I+LG++EV V FE VW FY+P+ IPDGF L +YC S
Sbjct: 763 GGGFAGGRISLGKIEVTKVNKFEKVWRCTNSNGKALGFTFYRPLEIPDGFSCLGYYCH-S 939
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYF 166
++PL G VLVARE + S +++ PAL+KPL+Y+L+WC ++ + + YF
Sbjct: 940 NDQPLRGHVLVARETTSKSQADCS---ESESPALKKPLNYSLIWCMDSHDECV-----YF 1095
Query: 167 WLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFSVW 226
WLP PP+GYKA+G +VT PD+P +E+ CVR DLT+ CE +L + S+ + F VW
Sbjct: 1096 WLPNPPKGYKAVGIVVTTNPDEPKAEEVRCVRTDLTEVCETSDLLLTIKSK--KNSFQVW 1269
Query: 227 SLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVCLKNLNPVLPAMPCLQQIHALIEHYG 286
+ +PCDRGM+ +GVSVGTFFC + + + + VVCLKNL+ +L AMP L QIHALIEHYG
Sbjct: 1270 NTQPCDRGMLARGVSVGTFFCGTYFDSEQVVDVVCLKNLDSLLHAMPNLNQIHALIEHYG 1449
Query: 287 PTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEAIDAGGSNLPIGGTNDGEFWIDLPC 346
PTVYFHP+E Y+PSSV WFF N A+L+ G + G+AID G+NLP GG NDG FWIDLP
Sbjct: 1450 PTVYFHPDEKYMPSSVSWFFKNGAILYTAGNAKGKAIDYHGTNLPGGGYNDGAFWIDLPT 1629
Query: 347 DDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAFSKV 406
D+D R +K G+++ A+LYVHVKPALGG FTD+AMWVFCPFNGP+TLK + +I +K+
Sbjct: 1630 DED-ARSNLKKGNIESAELYVHVKPALGGAFTDIAMWVFCPFNGPATLKVSLMNIEMNKI 1806
Query: 407 GEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASF 466
GEHVGDWEHFTLR+ NF+GELWS++FS+HSGGKWV+++DLE+I NK IVYSS+ GHAS+
Sbjct: 1807 GEHVGDWEHFTLRVSNFTGELWSVFFSEHSGGKWVNAFDLEFIKENKPIVYSSRHGHASY 1986
Query: 467 PHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGD-VVREPQWLQYMREWG 525
PH GTY+QGSSKLGIG+RNDA SN +DSS +Y+IVAAEYLGD V+ EP WLQYMREWG
Sbjct: 1987 PHAGTYLQGSSKLGIGVRNDAAKSNFILDSSFRYKIVAAEYLGDGVITEPCWLQYMREWG 2166
Query: 526 PKIVYGSKTELDKIINALPFRLRISFVNL 554
P IVY S++E++KII+ LP +R S NL
Sbjct: 2167 PTIVYDSRSEIEKIIDMLPILVRFSVENL 2253
>TC78461 weakly similar to GP|22136978|gb|AAM91718.1 unknown protein
{Arabidopsis thaliana}, partial (65%)
Length = 1464
Score = 270 bits (689), Expect(2) = e-113
Identities = 129/240 (53%), Positives = 170/240 (70%), Gaps = 2/240 (0%)
Frame = +3
Query: 271 AMPCLQQIHALIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEA-IDAGGSN 329
+MP L QI AL + + P +Y HP E + P S+ W+F+N A+L++KG + ID GSN
Sbjct: 753 SMPNLPQIKALAQAHSPFMYLHPNEKFQPCSIKWYFTNGALLYKKGEESNPMEIDPLGSN 932
Query: 330 LPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFN 389
LP GG+NDG +W+DLP D D RE VK GD K + Y HVKP GGTFTDLAMW+F PFN
Sbjct: 933 LPQGGSNDGAYWLDLP-KDKDNRERVKKGDFKNCQGYFHVKPIFGGTFTDLAMWIFYPFN 1109
Query: 390 GPSTLKFG-ITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEY 448
GP TLK G I +I+ K+GEH+GDWEH TLR+ NF+GEL +YFSQHS G+WVDS ++E+
Sbjct: 1110GPGTLKVGLIDNISLGKIGEHIGDWEHVTLRVSNFNGELKRVYFSQHSNGQWVDSSEVEF 1289
Query: 449 INGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYL 508
+GNK + YSS +GHA +P G +QG+S IGI+N+ S+L +D + +EIV+ EYL
Sbjct: 1290QSGNKPVAYSSLNGHAIYPKAGLVLQGTS--DIGIKNETKKSDLVIDFGVNFEIVSGEYL 1463
Score = 157 bits (396), Expect(2) = e-113
Identities = 85/199 (42%), Positives = 116/199 (57%)
Frame = +1
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G GF +GII+LGEL+V +++F +W + + F++P GIP GF L HY QP+
Sbjct: 124 GGGFGNGIIDLGELKVSQISTFNKIWTTLEGGQGDLGATFFEPTGIPQGFFTLGHYSQPN 303
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYF 166
NKPL+G+VLVA++ AL+KP+DYTLVW S + K Y
Sbjct: 304 -NKPLFGWVLVAKD--------------ESNGALKKPIDYTLVWSSKS-EKIKQDKDGYI 435
Query: 167 WLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFSVW 226
WLP P GY LG++VT P+KP+LD + CVR+DLTD+CE I +I E F+V
Sbjct: 436 WLPIAPNGYSPLGHIVTTTPEKPSLDRIQCVRSDLTDQCEINSWIWGKDKKIDEKGFNVH 615
Query: 227 SLRPCDRGMMGKGVSVGTF 245
+RP +RG+ V VGTF
Sbjct: 616 YVRPINRGIEAPSVLVGTF 672
>BG644557 similar to GP|21321777|gb OSJNBa0031O09.02 {Oryza sativa (japonica
cultivar-group)}, partial (22%)
Length = 693
Score = 200 bits (508), Expect(3) = 3e-79
Identities = 93/117 (79%), Positives = 106/117 (90%)
Frame = -2
Query: 430 IYFSQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRNDACS 489
I + HSGGKWVD+Y+LEYI+GNKAIVY+SK+GHAS+PHPGTYIQGSSKLGIGIRNDA
Sbjct: 581 ILLTGHSGGKWVDAYELEYIDGNKAIVYASKNGHASYPHPGTYIQGSSKLGIGIRNDARR 402
Query: 490 SNLYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVYGSKTELDKIINALPFR 546
SNL VDSS+ YEIVAA YLGDVV+EPQWLQYMR+WGPKIVY SKTELDKI+N+ F+
Sbjct: 401 SNLRVDSSVHYEIVAAAYLGDVVKEPQWLQYMRQWGPKIVYDSKTELDKILNSSSFK 231
Score = 81.3 bits (199), Expect(3) = 3e-79
Identities = 36/41 (87%), Positives = 37/41 (89%)
Frame = -3
Query: 543 LPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDERW 583
LP RLR SF NL +KLPVELYGEEGPTGPKEKNNWIGDERW
Sbjct: 241 LPLRLRSSFGNLFRKLPVELYGEEGPTGPKEKNNWIGDERW 119
Score = 54.3 bits (129), Expect(3) = 3e-79
Identities = 24/39 (61%), Positives = 30/39 (76%)
Frame = -1
Query: 395 KFGITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYFS 433
KFG+ S+AFSKVGEH+GDWEHFTLRI E + +Y +
Sbjct: 690 KFGMKSMAFSKVGEHIGDWEHFTLRIR*LHLENFGVYIT 574
>CA858601 weakly similar to GP|22136978|gb unknown protein {Arabidopsis
thaliana}, partial (33%)
Length = 695
Score = 216 bits (549), Expect = 3e-56
Identities = 108/224 (48%), Positives = 143/224 (63%), Gaps = 5/224 (2%)
Frame = +2
Query: 307 SNRAMLFRKGVSTGEA-IDAGGSNLPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKL 365
SN A+L++KG + I G+NLP DG +WIDLP D + +E VK G+L+ A+
Sbjct: 2 SNGALLYKKGEESNPVPIAQNGTNLPQDPNTDGAYWIDLPADAAN-KERVKKGNLQSAES 178
Query: 366 YVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAFSKVGEHVGDWEHFTLRICNFSG 425
YVHVKP GGTFTD+AMWVF PFNGP K ++ K+GEHVGDWEH TLR+ N G
Sbjct: 179 YVHVKPMYGGTFTDIAMWVFYPFNGPGRAKVEFLNVKLGKIGEHVGDWEHVTLRVSNLDG 358
Query: 426 ELWSIYFSQHSGGKWVDSYDLEYING----NKAIVYSSKSGHASFPHPGTYIQGSSKLGI 481
+LW +YFSQH+ G W+DS LE+ +G + +VY+S GHAS+PH G + G K GI
Sbjct: 359 QLWHVYFSQHNLGSWIDSSQLEFQDGIGATKRPVVYASLHGHASYPHAGLVLLG--KNGI 532
Query: 482 GIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWG 525
G R+D +D ++ +V+AEYLG V+EP WL Y + G
Sbjct: 533 GARDDTNKGKNVMDMG-KFVLVSAEYLGSEVKEPAWLNYFKGMG 661
>TC80114 weakly similar to GP|6721151|gb|AAF26779.1| unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 765
Score = 190 bits (483), Expect = 1e-48
Identities = 90/129 (69%), Positives = 104/129 (79%), Gaps = 3/129 (2%)
Frame = +3
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
GQ FASGI+NLGE+EVC +T FE VWNSNVMVE RKA+ FYKPVGIPDGFH+L HYCQPS
Sbjct: 387 GQDFASGIVNLGEIEVCKITKFETVWNSNVMVEPRKAITFYKPVGIPDGFHILGHYCQPS 566
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEI---VTSS 163
Y KPLWGFVL A++V S I +QNKLPALR PLD+ L+WC+N+G K+I V S+
Sbjct: 567 Y-KPLWGFVLAAKQV--ADSSYNNICNQNKLPALRNPLDFALIWCTNSGRKKIAMPVDSA 737
Query: 164 AYFWLPQPP 172
AYFWLPQPP
Sbjct: 738 AYFWLPQPP 764
>BF640859 weakly similar to GP|15450387|gb At2g44230/F4I1.4 {Arabidopsis
thaliana}, partial (23%)
Length = 655
Score = 165 bits (417), Expect = 5e-41
Identities = 96/239 (40%), Positives = 139/239 (57%), Gaps = 2/239 (0%)
Frame = +2
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G GFA+GII+LG L+V V++F VW + + ++P GIP GF +L Y QP+
Sbjct: 2 GSGFANGIIDLGGLQVSQVSTFNKVWATYDGGPDNQGATIFEPTGIPQGFSMLGCYSQPN 181
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYF 166
NKPL+G+VLVA++V + ++ L+KP+DYTLV + + K ++ Y
Sbjct: 182 -NKPLFGYVLVAKDVSSSTTNS----------TLKKPIDYTLV-LNTSTIKVNQDTTCYI 325
Query: 167 WLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFSVW 226
WLP P+GYKALG++VT DKP+LD++ CVR+DLTD+CE I + F+ +
Sbjct: 326 WLPIAPDGYKALGHVVTATQDKPSLDKIMCVRSDLTDQCESSSWIWGSND------FNFY 487
Query: 227 SLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLP--VVCLKNLNPVLPAMPCLQQIHALIE 283
+RP + G GV VG F N G +P + CLKNLN + MP L QI A+++
Sbjct: 488 DVRPINXGTQAPGVHVGAFVAQ---NGGTNIPPSISCLKNLNSISKIMPNLXQIDAILK 655
>TC86254 similar to GP|14970841|emb|CAC44501. beta-galactosidase {Fragaria x
ananassa}, partial (87%)
Length = 2581
Score = 32.3 bits (72), Expect = 0.52
Identities = 25/75 (33%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = -1
Query: 330 LPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKLYVH--VKPALGGTFTDLAMWVFCP 387
LP G ND W ++ CD Q F++H D L +H V+PA T+ + CP
Sbjct: 2155 LPFGFKNDRGTWYNV-CDGFPQ--FLRHLDEV*GPL*LHESVQPAFEATYVGQYRPMLCP 1985
Query: 388 FNGPSTLKFGITSIA 402
F S + SIA
Sbjct: 1984 FTQASPFPIPVKSIA 1940
>TC86035 similar to PIR|E71435|E71435 probable triacylglycerol lipase -
Arabidopsis thaliana, partial (57%)
Length = 1783
Score = 30.8 bits (68), Expect = 1.5
Identities = 18/44 (40%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Frame = +3
Query: 334 GTNDGEFWIDLPCDDDDQREFVKHGDL-KGAKLYVHVKPALGGT 376
G+ND + +D P DD+ +RE V++GDL + A H PA+ T
Sbjct: 546 GSNDWKGMLD-PLDDNLRREVVRYGDLVQAAYQAFHADPAMSTT 674
>TC80238 weakly similar to GP|14587270|dbj|BAB61188. putative receptor
serine/threonine kinase {Oryza sativa (japonica
cultivar-group)}, partial (16%)
Length = 1130
Score = 30.4 bits (67), Expect = 2.0
Identities = 23/80 (28%), Positives = 35/80 (43%), Gaps = 7/80 (8%)
Frame = +2
Query: 342 IDLPCDDDDQREFVKHGDLKGAKLYV-HVKPALGGTFTDLAMWVFCPF------NGPSTL 394
I+ P D + V D + +LY+ H + L F +F PF N PS++
Sbjct: 176 IEFPADPVPVKLLVTDIDCQSQQLYLSHPQNCLSSIFVTHNFSLFYPFSLASLFNRPSSV 355
Query: 395 KFGITSIAFSKVGEHVGDWE 414
+T S VG+H+ WE
Sbjct: 356 N-NVTFFNCSSVGQHLRSWE 412
>TC77156 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-like protein
{Arabidopsis thaliana}, partial (66%)
Length = 1096
Score = 29.6 bits (65), Expect = 3.4
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 3/66 (4%)
Frame = +1
Query: 492 LYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVYGSKTELDKIINA---LPFRLR 548
L+ DSS ++V+ E+ +V EP+W EW K + +D +N LPF++
Sbjct: 898 LHTDSSCSEQVVSPEFASEVQSEPKW----NEW-DKNLEVPYNYIDATLNTGFWLPFQIL 1062
Query: 549 ISFVNL 554
I N+
Sbjct: 1063IHCKNV 1080
>TC77157 homologue to GP|13877717|gb|AAK43936.1 OsNAC6 protein-like protein
{Arabidopsis thaliana}, partial (66%)
Length = 1998
Score = 28.5 bits (62), Expect = 7.5
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = +3
Query: 492 LYVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVY 530
L+ DSS ++V+ E+ +V EP+W EW + +
Sbjct: 1074 LHTDSSCSEQVVSPEFASEVQSEPKW----NEWDKNLEF 1178
>TC86355 weakly similar to GP|9757805|dbj|BAB08323.1 CCR4-associated
factor-like protein {Arabidopsis thaliana}, partial
(84%)
Length = 1041
Score = 28.5 bits (62), Expect = 7.5
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +3
Query: 315 KGVSTGEAIDAGGSNLPIGGTNDGEFWIDLPCDDDDQRE 353
K + G + G NLP GTN+ W CD D +R+
Sbjct: 375 KLIQVGITLSDGNGNLPHFGTNNSYIWEFNFCDFDFERD 491
>TC87918 weakly similar to GP|15810397|gb|AAL07086.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1746
Score = 28.1 bits (61), Expect = 9.8
Identities = 22/65 (33%), Positives = 29/65 (43%), Gaps = 2/65 (3%)
Frame = -1
Query: 11 CQFHSHSPSTVFPSFSTTPMASRSVYSVIV--HVCVITGQGFASGIINLGELEVCSVTSF 68
C F S PS FPSF+ S S +S I T +GF S ++ L + + S
Sbjct: 978 CSFSS--PSASFPSFTPLFSLSLSSFSDIF*SLSSAATSEGFCSSVLVTSLLSLGLLISA 805
Query: 69 ELVWN 73
L WN
Sbjct: 804 ILFWN 790
>TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (96%)
Length = 1582
Score = 28.1 bits (61), Expect = 9.8
Identities = 30/120 (25%), Positives = 49/120 (40%), Gaps = 10/120 (8%)
Frame = +2
Query: 469 PGTYIQGSSKLGIGIRNDACSSNLYV----------DSSIQYEIVAAEYLGDVVREPQWL 518
PGT + SS+L I N++C+S + ++ ++AAEY+G V
Sbjct: 8 PGTSLSLSSRLSDQISNESCASEAMAALILHAPNNGNKNVYKTLIAAEYVGVQV------ 169
Query: 519 QYMREWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWI 578
+ P G + + IN +N + K+PV E P GP ++N I
Sbjct: 170 ----QLTPDFQMGVSNKTPQFIN----------MNPIGKVPV----LETPDGPVFESNAI 295
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,211,656
Number of Sequences: 36976
Number of extensions: 360533
Number of successful extensions: 1609
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1574
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1583
length of query: 583
length of database: 9,014,727
effective HSP length: 101
effective length of query: 482
effective length of database: 5,280,151
effective search space: 2545032782
effective search space used: 2545032782
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0587.12


