

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0546.13
(1118 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
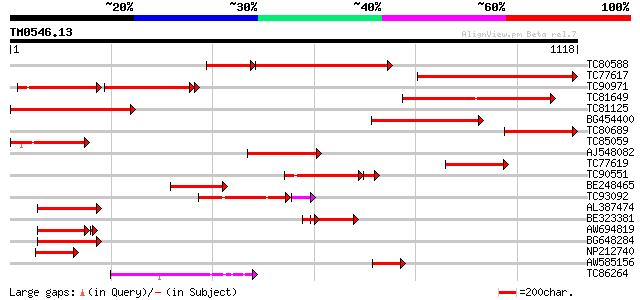
Score E
Sequences producing significant alignments: (bits) Value
TC80588 similar to GP|23617092|dbj|BAC20775. putative ubiquitin ... 499 0.0
TC77617 similar to GP|6671947|gb|AAF23207.1| putative ubiquitin ... 608 e-174
TC90971 similar to GP|10998129|dbj|BAB17021. ubiquitin carboxyl-... 348 e-160
TC81649 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-... 482 e-136
TC81125 similar to GP|17381108|gb|AAL36366.1 putative ubiquitin ... 473 e-133
BG454400 similar to GP|6671947|gb putative ubiquitin carboxyl-te... 429 e-120
TC80689 similar to GP|6671947|gb|AAF23207.1| putative ubiquitin ... 279 4e-75
TC85059 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-... 271 8e-73
AJ548082 similar to GP|10178116|d ubiquitin carboxyl-terminal hy... 252 5e-67
TC77619 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-... 232 5e-61
TC90551 similar to GP|23617092|dbj|BAC20775. putative ubiquitin ... 195 1e-60
BE248465 homologue to GP|10178116|d ubiquitin carboxyl-terminal ... 223 4e-58
TC93092 similar to GP|16944687|emb|CAD11412. conserved hypotheti... 169 5e-45
AL387474 143 3e-34
BE323381 similar to GP|11993471|g ubiquitin-specific protease 12... 133 3e-31
AW694819 117 1e-27
BG648284 110 4e-24
NP212740 NP212740|Y18251.1|CAA77095.1 NtN2 109 7e-24
AW585156 similar to GP|10178116|db ubiquitin carboxyl-terminal h... 99 7e-21
TC86264 similar to GP|11993486|gb|AAG42761.1 ubiquitin-specific ... 93 5e-19
>TC80588 similar to GP|23617092|dbj|BAC20775. putative ubiquitin
carboxyl-terminal hydrolase {Oryza sativa (japonica
cultivar-group)}, partial (30%)
Length = 1110
Score = 499 bits (1284), Expect(2) = 0.0
Identities = 243/271 (89%), Positives = 256/271 (93%)
Frame = +2
Query: 485 LEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAEHLRE 544
LEEQYGGEEELPQTNPGFNN PFKFTKYSNAYMLVYIRE+DKDK+ICNVDEKDIAEHLRE
Sbjct: 293 LEEQYGGEEELPQTNPGFNNPPFKFTKYSNAYMLVYIRESDKDKIICNVDEKDIAEHLRE 472
Query: 545 RLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRVQKQM 604
RLKKEQEEKEHKKKEKAEAHLYTIIKVAR+ED++ Q+GKDIYFDLVDHDKVRSFRVQKQ
Sbjct: 473 RLKKEQEEKEHKKKEKAEAHLYTIIKVARDEDIEGQMGKDIYFDLVDHDKVRSFRVQKQT 652
Query: 605 SFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSVGQVREVSNKVHN 664
FN+FKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLT EE Q+VGQ+REVSNKVHN
Sbjct: 653 PFNVFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTQIEETQTVGQLREVSNKVHN 832
Query: 665 AELKLFLEVELGPDLRPIAPSDKTKDDILLFFKLYDPEKEELRYVGRLFVKSTGKPSEIL 724
AELKLFLEVE G DL PIAP DKTKDDILLFFKLYDPEKEELRYVGRL VK TGKPSEIL
Sbjct: 833 AELKLFLEVEKGMDLCPIAPPDKTKDDILLFFKLYDPEKEELRYVGRLSVKCTGKPSEIL 1012
Query: 725 TRLNEMAGYDPDEEIGLYEEIKFEPNVMCEP 755
TRLNEMAGY P+++I LY+EIKFEPNVMCEP
Sbjct: 1013TRLNEMAGYVPEKDIVLYDEIKFEPNVMCEP 1105
Score = 193 bits (491), Expect(2) = 0.0
Identities = 90/97 (92%), Positives = 95/97 (97%)
Frame = +1
Query: 388 LQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVH 447
LQLQLKRFEYDFMRDTMVKINDRYEFP +LDLDR++GKYLSPDADR+VRNLYTLHSVLVH
Sbjct: 1 LQLQLKRFEYDFMRDTMVKINDRYEFPSQLDLDRENGKYLSPDADRSVRNLYTLHSVLVH 180
Query: 448 SGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRA 484
SGGVHGGHYYAFIRPTLS+QWYKFDDERVTKED KRA
Sbjct: 181 SGGVHGGHYYAFIRPTLSEQWYKFDDERVTKEDNKRA 291
>TC77617 similar to GP|6671947|gb|AAF23207.1| putative ubiquitin
carboxyl-terminal hydrolase {Arabidopsis thaliana},
partial (27%)
Length = 1473
Score = 608 bits (1569), Expect = e-174
Identities = 291/315 (92%), Positives = 305/315 (96%)
Frame = +1
Query: 804 VVHFRSLDKPKEDDFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQP 863
VVHFRSLD+PKEDDF LEMSRLYTYDDVVE+VAQQL LDDPSKIRLTPHNCYSQQPKPQP
Sbjct: 1 VVHFRSLDRPKEDDFSLEMSRLYTYDDVVERVAQQLGLDDPSKIRLTPHNCYSQQPKPQP 180
Query: 864 IKYRGVDHLSDMLVHYNQTSDILYYEILDIPLPELQGLKTLKVAFYHATKDEVVSHTIRL 923
IK+RGVDHLSDMLVHYNQTSDILYYE+LDIPLPELQGLKTLKVAF+HA KDEVVSHTIRL
Sbjct: 181 IKHRGVDHLSDMLVHYNQTSDILYYEVLDIPLPELQGLKTLKVAFHHAIKDEVVSHTIRL 360
Query: 924 PKQSTVGDVLDDLKTKVELSHPNAELRLLEVFYHKIYKVFPPNEKIETINDQYWTLRAEE 983
PKQSTVGDVLDDLKTKVELSHP+AELRLLEVFYHKIYKVFP NEKIE INDQYWTLRAEE
Sbjct: 361 PKQSTVGDVLDDLKTKVELSHPDAELRLLEVFYHKIYKVFPSNEKIENINDQYWTLRAEE 540
Query: 984 VPEEEKNLGPHDRLIHVYHFTKDTSQNQMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQ 1043
+PEEEKN+GP DRLIHVYHFTKDT+QNQMQIQNFG+PFFLVI EGE L+EIKVRIQKKLQ
Sbjct: 541 IPEEEKNIGPQDRLIHVYHFTKDTAQNQMQIQNFGDPFFLVIHEGEALSEIKVRIQKKLQ 720
Query: 1044 VPDDEFEKWKFAFFALGRPEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAV 1103
VPD+EF KWKFAF +LGRPEYLQDSDIVS+RFQRRDVYGAWEQYLGLEHTDNAPKRSYAV
Sbjct: 721 VPDEEFSKWKFAFLSLGRPEYLQDSDIVSSRFQRRDVYGAWEQYLGLEHTDNAPKRSYAV 900
Query: 1104 NQNRHTFEKPVKIYN 1118
NQNRHTFEKPVKIYN
Sbjct: 901 NQNRHTFEKPVKIYN 945
>TC90971 similar to GP|10998129|dbj|BAB17021. ubiquitin carboxyl-terminal
hydrolase-like protein {Arabidopsis thaliana}, partial
(50%)
Length = 1299
Score = 348 bits (893), Expect(3) = e-160
Identities = 165/177 (93%), Positives = 172/177 (96%)
Frame = +2
Query: 188 WTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQ 247
W +DSKKETG VGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMP+GSIPLALQ
Sbjct: 737 WAHDSKKETGCVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPSGSIPLALQ 916
Query: 248 SLFYKLQYSDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQ 307
SLFYKLQY+D+SV+TKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQ
Sbjct: 917 SLFYKLQYNDSSVSTKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGTVVEGTIQ 1096
Query: 308 KLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNK 364
KLFEGHHMNYIEC+NVDYKSTRKESFYDLQLDVKGC DVYASFDKYVEV LEGDN+
Sbjct: 1097KLFEGHHMNYIECMNVDYKSTRKESFYDLQLDVKGCTDVYASFDKYVEVXQLEGDNQ 1267
Score = 231 bits (589), Expect(3) = e-160
Identities = 111/167 (66%), Positives = 138/167 (82%), Gaps = 1/167 (0%)
Frame = +3
Query: 16 EMLVPHTD-LPENNHQPMEVVAQPEAAPTVESQPVEEPPQSRFTWRIDNFSRMNVKKLYS 74
EMLVP++D + QPMEVV Q E TV++ VE+PP RFTW IDNFSR+ KK YS
Sbjct: 222 EMLVPNSDAVVVEGPQPMEVV-QAENTSTVDAVAVEDPPTGRFTWTIDNFSRLP-KKHYS 395
Query: 75 EVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWSRYAQFSLAVVNQIQNKYTV 134
+VF VGGYKWR+LIFPKGNN ++LSMY+DVAD+ ++PYGW+R+AQFSL VVNQ+ +KY+V
Sbjct: 396 DVFTVGGYKWRILIFPKGNNAEHLSMYIDVADAGSMPYGWTRFAQFSLTVVNQVHSKYSV 575
Query: 135 RKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEVLV 181
RK+TQHQFNARESDWGFT+FMPL ELYDPSRGY++ D ++EA+V V
Sbjct: 576 RKETQHQFNARESDWGFTNFMPLAELYDPSRGYVVEDRCILEADVNV 716
Score = 26.9 bits (58), Expect(3) = e-160
Identities = 10/11 (90%), Positives = 11/11 (99%)
Frame = +3
Query: 364 KYHAEQYGLQD 374
KYHAEQYGLQ+
Sbjct: 1266 KYHAEQYGLQE 1298
>TC81649 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-terminal
hydrolase {Arabidopsis thaliana}, partial (24%)
Length = 905
Score = 482 bits (1240), Expect = e-136
Identities = 229/302 (75%), Positives = 268/302 (87%), Gaps = 1/302 (0%)
Frame = +1
Query: 775 FQKAP-AMDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDDVVE 833
FQK+P A D ++ YPDVPS+ EYV NRQVV FR L+KPKED+F LE+S+L+TYDDVVE
Sbjct: 4 FQKSPPAGDGQQQYCYPDVPSFFEYVQNRQVVRFRFLEKPKEDEFSLELSKLHTYDDVVE 183
Query: 834 KVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEILDI 893
+V+Q L L+DPSKIRLT HNCYSQQPKPQPIKYRGVDHLS+MLVHYNQ SDILYYE+LDI
Sbjct: 184 RVSQHLGLNDPSKIRLTSHNCYSQQPKPQPIKYRGVDHLSEMLVHYNQASDILYYEVLDI 363
Query: 894 PLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHPNAELRLLE 953
PLPELQ LKTLK+AF+H KDEV+ IRL K STV DV++DLK+KV+LSHP+AELRL+E
Sbjct: 364 PLPELQNLKTLKIAFHHDAKDEVM--IIRLQKHSTVADVINDLKSKVDLSHPDAELRLVE 537
Query: 954 VFYHKIYKVFPPNEKIETINDQYWTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQNQMQ 1013
VF HKIYK+F NEKIE IND YWTLRAEE+PEEEK+LGPHDR+IHVYHF KDT+QNQM
Sbjct: 538 VFNHKIYKIFHVNEKIENINDHYWTLRAEEIPEEEKSLGPHDRMIHVYHFLKDTAQNQMH 717
Query: 1014 IQNFGEPFFLVIREGETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPEYLQDSDIVSN 1073
+QNFG+PFFLVIREGETL ++K+R+QKKLQVP++EF KWKFAF +LGRPEYLQDSDI+S+
Sbjct: 718 VQNFGDPFFLVIREGETLADVKLRVQKKLQVPNEEFLKWKFAFVSLGRPEYLQDSDIISS 897
Query: 1074 RF 1075
RF
Sbjct: 898 RF 903
>TC81125 similar to GP|17381108|gb|AAL36366.1 putative ubiquitin
carboxyl-terminal hydrolase {Arabidopsis thaliana},
partial (34%)
Length = 820
Score = 473 bits (1216), Expect = e-133
Identities = 224/248 (90%), Positives = 237/248 (95%), Gaps = 1/248 (0%)
Frame = +1
Query: 1 MTVMTPAPIDQQEDEEMLVPHTDLPENN-HQPMEVVAQPEAAPTVESQPVEEPPQSRFTW 59
MTVM APIDQQEDEE+LVPH DLPENN HQPMEVVAQPE A TVESQPV +PPQ+RFTW
Sbjct: 76 MTVMMSAPIDQQEDEEVLVPHADLPENNNHQPMEVVAQPETANTVESQPVPDPPQTRFTW 255
Query: 60 RIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWSRYAQ 119
RIDNF+R+N KKLYSEVFVVG YKWRVLIFPKGNNVDYLSMYLDVADST+LPYGWSRYAQ
Sbjct: 256 RIDNFTRLNTKKLYSEVFVVGAYKWRVLIFPKGNNVDYLSMYLDVADSTSLPYGWSRYAQ 435
Query: 120 FSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEV 179
FSLA+VNQI NK+TVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYL+NDTL++EAEV
Sbjct: 436 FSLAIVNQIHNKFTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLVNDTLIIEAEV 615
Query: 180 LVRRIVDYWTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPA 239
LVR+IVDYW YDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMP+
Sbjct: 616 LVRKIVDYWNYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPS 795
Query: 240 GSIPLALQ 247
GSIPLALQ
Sbjct: 796 GSIPLALQ 819
>BG454400 similar to GP|6671947|gb putative ubiquitin carboxyl-terminal
hydrolase {Arabidopsis thaliana}, partial (18%)
Length = 664
Score = 429 bits (1103), Expect = e-120
Identities = 204/220 (92%), Positives = 215/220 (97%)
Frame = +1
Query: 714 VKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDII 773
V +TGKPSEIL RLN+MAGYDP+EEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDII
Sbjct: 1 VNNTGKPSEILARLNKMAGYDPEEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDII 180
Query: 774 CFQKAPAMDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDDVVE 833
CFQKAPA D+EEH+RYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRL+TYDDVVE
Sbjct: 181 CFQKAPATDNEEHIRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLFTYDDVVE 360
Query: 834 KVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEILDI 893
+VA+QL LDDPSKIRLTPHNCYSQQPKPQPIKYRGV+HLSDMLVHYNQTSDILYYE+LDI
Sbjct: 361 RVAEQLGLDDPSKIRLTPHNCYSQQPKPQPIKYRGVEHLSDMLVHYNQTSDILYYEVLDI 540
Query: 894 PLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVL 933
PLPELQGLKTLKVAF+HATKDEVV HTIRLPKQSTVGDVL
Sbjct: 541 PLPELQGLKTLKVAFHHATKDEVVIHTIRLPKQSTVGDVL 660
>TC80689 similar to GP|6671947|gb|AAF23207.1| putative ubiquitin
carboxyl-terminal hydrolase {Arabidopsis thaliana},
partial (12%)
Length = 1037
Score = 279 bits (714), Expect = 4e-75
Identities = 129/142 (90%), Positives = 137/142 (95%)
Frame = +1
Query: 977 WTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQNQMQIQNFGEPFFLVIREGETLTEIKV 1036
WTLRAEE+PEEEKNLGPHDRLIHVYHFTKDT+QNQMQIQNFGEPFFLVI E ETL EI++
Sbjct: 4 WTLRAEEIPEEEKNLGPHDRLIHVYHFTKDTTQNQMQIQNFGEPFFLVIHECETLAEIRL 183
Query: 1037 RIQKKLQVPDDEFEKWKFAFFALGRPEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNA 1096
RIQKKLQVPDDEF KWKFAFF+LGRPEYL+DS++VSNRFQRRDVYGAWEQYLGLEHTDNA
Sbjct: 184 RIQKKLQVPDDEFGKWKFAFFSLGRPEYLEDSEVVSNRFQRRDVYGAWEQYLGLEHTDNA 363
Query: 1097 PKRSYAVNQNRHTFEKPVKIYN 1118
PKRSYA NQNRHTFEKPVKIYN
Sbjct: 364 PKRSYAANQNRHTFEKPVKIYN 429
>TC85059 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-terminal
hydrolase {Arabidopsis thaliana}, partial (10%)
Length = 561
Score = 271 bits (694), Expect = 8e-73
Identities = 138/164 (84%), Positives = 145/164 (88%), Gaps = 7/164 (4%)
Frame = +2
Query: 1 MTVMTPAPIDQQ--EDEEMLVPHT----DLPENN-HQPMEVVAQPEAAPTVESQPVEEPP 53
MT+MTPAPIDQQ EDEEMLVPH DL ENN HQPM+VVAQPE A TVE PVE+P
Sbjct: 74 MTIMTPAPIDQQQQEDEEMLVPHAVPHADLAENNNHQPMDVVAQPETANTVE--PVEDPS 247
Query: 54 QSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYG 113
SRFTWRIDNFSR+N+KKLYS+VFVVG YKWRVLIFPKGNNVDYLSMYLDVADST+LPYG
Sbjct: 248 PSRFTWRIDNFSRVNLKKLYSDVFVVGSYKWRVLIFPKGNNVDYLSMYLDVADSTSLPYG 427
Query: 114 WSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPL 157
WSRYAQ SLAVVNQI NKYT RKDTQHQFNARESDWGFTSFMPL
Sbjct: 428 WSRYAQVSLAVVNQIHNKYTGRKDTQHQFNARESDWGFTSFMPL 559
>AJ548082 similar to GP|10178116|d ubiquitin carboxyl-terminal hydrolase
{Arabidopsis thaliana}, partial (11%)
Length = 474
Score = 252 bits (644), Expect = 5e-67
Identities = 126/146 (86%), Positives = 131/146 (89%)
Frame = -2
Query: 470 KFDDERVTKEDTKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKV 529
KFDDERVTKED KRA+EEQYGGEEELPQTNPGFN TPFKFTKYSNAYMLVYIREADK+KV
Sbjct: 473 KFDDERVTKEDPKRAMEEQYGGEEELPQTNPGFNYTPFKFTKYSNAYMLVYIREADKEKV 294
Query: 530 ICNVDEKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDL 589
ICNVDEKDIAEHLRERLKKEQE KEHKKKEKAEAHLYTIIKVAR+EDL EQIGKDIYFDL
Sbjct: 293 ICNVDEKDIAEHLRERLKKEQEVKEHKKKEKAEAHLYTIIKVARDEDLGEQIGKDIYFDL 114
Query: 590 VDHDKVRSFRVQKQMSFNLFKEEVAK 615
V+HDKVRSFRV F F + K
Sbjct: 113 VEHDKVRSFRVLFFFFFFFFCHTIIK 36
>TC77619 similar to GP|10178116|dbj|BAB11409. ubiquitin carboxyl-terminal
hydrolase {Arabidopsis thaliana}, partial (11%)
Length = 822
Score = 232 bits (592), Expect = 5e-61
Identities = 113/124 (91%), Positives = 118/124 (95%)
Frame = -1
Query: 860 KPQPIKYRGVDHLSDMLVHYNQTSDILYYEILDIPLPELQGLKTLKVAFYHATKDEVVSH 919
KPQPIK+RGVDHLSDMLVHYNQTSDILYYE+LDIPLPELQGLKTLKVAF+HA KDEVVSH
Sbjct: 372 KPQPIKHRGVDHLSDMLVHYNQTSDILYYEVLDIPLPELQGLKTLKVAFHHAIKDEVVSH 193
Query: 920 TIRLPKQSTVGDVLDDLKTKVELSHPNAELRLLEVFYHKIYKVFPPNEKIETINDQYWTL 979
TIRLPKQSTVGDVLDDLKTKVELSHP+AELRLLE YHKIYKVFP +EKIE IND YWTL
Sbjct: 192 TIRLPKQSTVGDVLDDLKTKVELSHPDAELRLLESLYHKIYKVFPIHEKIENINDPYWTL 13
Query: 980 RAEE 983
RAEE
Sbjct: 12 RAEE 1
>TC90551 similar to GP|23617092|dbj|BAC20775. putative ubiquitin
carboxyl-terminal hydrolase {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 557
Score = 195 bits (496), Expect(2) = 1e-60
Identities = 101/157 (64%), Positives = 121/157 (76%)
Frame = +1
Query: 542 LRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRVQ 601
++ER +++ +KE K + L+ + + +EDL EQIGKDIYFDLV+HDKVRSFRVQ
Sbjct: 1 VKERTGRKRAQKEGKGRGSP---LHLL*RWHWDEDLGEQIGKDIYFDLVEHDKVRSFRVQ 171
Query: 602 KQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSVGQVREVSNK 661
KQ FN+FKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLT EEAQSVGQ+RE+SNK
Sbjct: 172 KQTPFNVFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTQIEEAQSVGQLREISNK 351
Query: 662 VHNAELKLFLEVELGPDLRPIAPSDKTKDDILLFFKL 698
VHNAELKLFLEVEL + + K + FF++
Sbjct: 352 VHNAELKLFLEVELWTGFMSHCTT*EDKG*YITFFQV 462
Score = 57.8 bits (138), Expect(2) = 1e-60
Identities = 26/31 (83%), Positives = 29/31 (92%)
Frame = +3
Query: 699 YDPEKEELRYVGRLFVKSTGKPSEILTRLNE 729
YDPEKEELR+VGRLFV +TGKPSEIL RLN+
Sbjct: 465 YDPEKEELRFVGRLFVNNTGKPSEILXRLNK 557
>BE248465 homologue to GP|10178116|d ubiquitin carboxyl-terminal hydrolase
{Arabidopsis thaliana}, partial (9%)
Length = 336
Score = 223 bits (567), Expect = 4e-58
Identities = 107/111 (96%), Positives = 109/111 (97%)
Frame = +2
Query: 318 IECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQYGLQDAKK 377
IECINVDYKSTRKESFYDLQLDVKGC DVYASFDKYVEVE LEGDNKYHAEQYGLQDAKK
Sbjct: 2 IECINVDYKSTRKESFYDLQLDVKGCPDVYASFDKYVEVERLEGDNKYHAEQYGLQDAKK 181
Query: 378 GVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLS 428
GVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPL+LDLDRD+GKYLS
Sbjct: 182 GVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLQLDLDRDNGKYLS 334
>TC93092 similar to GP|16944687|emb|CAD11412. conserved hypothetical protein
{Neurospora crassa}, partial (17%)
Length = 714
Score = 169 bits (428), Expect(2) = 5e-45
Identities = 91/186 (48%), Positives = 118/186 (62%), Gaps = 3/186 (1%)
Frame = +1
Query: 372 LQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDA 431
L DA KGV+F FP VL LQLKRFEYD RD M+KINDRYEFP D +LS DA
Sbjct: 4 LLDANKGVIFQSFPNVLHLQLKRFEYDIQRDMMMKINDRYEFPEFFDA----APFLSEDA 171
Query: 432 DRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQYGG 491
D++ Y LH VLVHSG ++ GHYYAF++P +YK+DD++VTK + LEE +GG
Sbjct: 172 DKSEPWTYQLHGVLVHSGDLNAGHYYAFLKPEKDGWFYKYDDDKVTKATMREVLEENFGG 351
Query: 492 EEELPQTN---PGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAEHLRERLKK 548
E + P + P TP + ++AYMLVYIR+ D +IC V ++DI HLR R ++
Sbjct: 352 EYKTPANHLRAPLQKKTP--VMRQNSAYMLVYIRQTRLDDIICPVTKEDIPIHLRTRFEE 525
Query: 549 EQEEKE 554
E +E
Sbjct: 526 ETALRE 543
Score = 31.6 bits (70), Expect(2) = 5e-45
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 4/52 (7%)
Frame = +2
Query: 556 KKKEKAEAHLYTIIKVARNEDLKEQIGKDIYF---DLVDHDKV-RSFRVQKQ 603
K+KEK E HLY +KV + K+ G D+ D D + RS+RV +Q
Sbjct: 548 KRKEKEEQHLYIGVKVITGDTFKKHGGTDLTSFEPDQADTEAAPRSYRVLRQ 703
>AL387474
Length = 494
Score = 143 bits (361), Expect = 3e-34
Identities = 62/128 (48%), Positives = 92/128 (71%), Gaps = 2/128 (1%)
Frame = +2
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNN--VDYLSMYLDVADSTNLPYG 113
+FTW+I+NFSR+NV KLYSE +V+ GY WR+ +FPKG++ VD L ++L+ + N+ G
Sbjct: 80 KFTWKIENFSRLNVDKLYSEPYVLSGYPWRIALFPKGSSSAVDQLGIFLEAMKTANMSEG 259
Query: 114 WSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTL 173
W R A+F AV NQ+++ T+ K+T +F+A E +WG+ SFM L L DP RG+++NDT
Sbjct: 260 WKRDAKFKFAVFNQVEDNRTITKETSQEFSASEDEWGYFSFMTLAALRDPGRGFIVNDTC 439
Query: 174 VVEAEVLV 181
+V AE+ V
Sbjct: 440 IVGAEIFV 463
>BE323381 similar to GP|11993471|g ubiquitin-specific protease 12
{Arabidopsis thaliana}, partial (9%)
Length = 356
Score = 133 bits (335), Expect = 3e-31
Identities = 62/95 (65%), Positives = 81/95 (85%), Gaps = 1/95 (1%)
Frame = +1
Query: 594 KVRSFRVQKQMSFNLF-KEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSV 652
+V +F+ + + F+ F +EEVAKEFGIPV++QRFW+WAKRQNHT+RP+RP+T EEAQ+V
Sbjct: 67 EVFAFKNRCHLPFSSFDQEEVAKEFGIPVEYQRFWMWAKRQNHTFRPSRPVTAQEEAQAV 246
Query: 653 GQVREVSNKVHNAELKLFLEVELGPDLRPIAPSDK 687
GQ+REVSNK +N ELKLFLE+E+G DLRPI P +K
Sbjct: 247 GQLREVSNKANNGELKLFLEIEMGQDLRPIPPPEK 351
Score = 62.8 bits (151), Expect = 7e-10
Identities = 28/34 (82%), Positives = 31/34 (90%)
Frame = +3
Query: 577 LKEQIGKDIYFDLVDHDKVRSFRVQKQMSFNLFK 610
L EQIGKDI+FDLVDHDKVRSFR+QKQM F +FK
Sbjct: 9 LHEQIGKDIFFDLVDHDKVRSFRIQKQMPFTIFK 110
>AW694819
Length = 502
Score = 117 bits (293), Expect(2) = 1e-27
Identities = 50/104 (48%), Positives = 75/104 (72%), Gaps = 2/104 (1%)
Frame = +1
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNN--VDYLSMYLDVADSTNLPYG 113
+FTW+I+NFSR+NV KLYSE +V+ GY WR+ +FPKG++ VD L ++L+ + N+ G
Sbjct: 124 KFTWKIENFSRLNVDKLYSEPYVLSGYPWRIALFPKGSSSAVDQLGIFLEAMKTANMSEG 303
Query: 114 WSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPL 157
W R A+F AV NQ+++ T+ K+T +F+A E +WG+ SFM L
Sbjct: 304 WKRDAKFKFAVFNQVEDNRTITKETSQEFSASEDEWGYFSFMTL 435
Score = 25.0 bits (53), Expect(2) = 1e-27
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = +2
Query: 160 LYDPSRGYLLNDTL 173
L DP RG+++NDTL
Sbjct: 443 LRDPGRGFIVNDTL 484
>BG648284
Length = 778
Score = 110 bits (274), Expect = 4e-24
Identities = 52/126 (41%), Positives = 77/126 (60%)
Frame = +3
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWS 115
+FTWRI++FS+ N KL S+ F + GY W++L+ P +VD+ ++YL VADS PYGW
Sbjct: 93 KFTWRIEDFSKKNCMKLKSKDFKIRGYTWKILVHPLRRDVDHFTLYLMVADSLP-PYGWD 269
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
R F L +VNQ+ + K+TQ +FN WG F+ + D +GYL+ DT ++
Sbjct: 270 RNTFFKLVLVNQLDRNKNIVKETQQKFNGGYRSWG-AFFLNFNKFIDHKQGYLVRDTCII 446
Query: 176 EAEVLV 181
EA + V
Sbjct: 447 EAHICV 464
>NP212740 NP212740|Y18251.1|CAA77095.1 NtN2
Length = 400
Score = 109 bits (272), Expect = 7e-24
Identities = 49/85 (57%), Positives = 61/85 (71%)
Frame = +1
Query: 52 PPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLP 111
P +TWR + FSR+ LYS+VF GGYKWR +I P+GNN DYLS+YL ADS +LP
Sbjct: 43 PGIQSYTWRTERFSRVRATVLYSDVFEAGGYKWRAIIHPRGNNTDYLSIYLCTADSASLP 222
Query: 112 YGWSRYAQFSLAVVNQIQNKYTVRK 136
GWS Y +F+L VVNQI+ KY+V K
Sbjct: 223 DGWSSYVEFTLKVVNQIEYKYSVTK 297
>AW585156 similar to GP|10178116|db ubiquitin carboxyl-terminal hydrolase
{Arabidopsis thaliana}, partial (5%)
Length = 217
Score = 99.4 bits (246), Expect = 7e-21
Identities = 44/66 (66%), Positives = 56/66 (84%)
Frame = +3
Query: 715 KSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDIIC 774
+ + KP +IL +LNEMAG+DPDEEI L+EEIKF+P +MCE +D+K TFR + LEDGDIIC
Sbjct: 6 RGSRKPVDILKKLNEMAGFDPDEEIDLFEEIKFDPKIMCEHVDQKSTFRDNLLEDGDIIC 185
Query: 775 FQKAPA 780
FQK+PA
Sbjct: 186 FQKSPA 203
>TC86264 similar to GP|11993486|gb|AAG42761.1 ubiquitin-specific protease 23
{Arabidopsis thaliana}, partial (42%)
Length = 3030
Score = 93.2 bits (230), Expect = 5e-19
Identities = 82/305 (26%), Positives = 138/305 (44%), Gaps = 16/305 (5%)
Frame = +3
Query: 200 GLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPL-ALQS-LFYKLQYSD 257
GL N G TC++NS+LQ L + + + AG L A+Q+ + LQ +
Sbjct: 393 GLWNLGNTCFLNSVLQCLTYTEPLAAYLQSGKHKSSCHIAGFCALCAIQNHVSCALQATG 572
Query: 258 TSVATKELTKSFGWDT--YDSFMQHDVQELNRVLCEKLE---------DKMKGTVVEGTI 306
V+ +++ + + + Q D E L E + + G + +
Sbjct: 573 RIVSPQDMVGNLRCISRNFRKARQEDAHEYMVNLLESMHKCCLPSGVPSESPGAFEKSLV 752
Query: 307 QKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNK-Y 365
K+F G + ++C Y S + + F DL L++ + + + E L+G K Y
Sbjct: 753 HKIFGGRLRSQVKCRQCSYSSNKFDPFLDLSLEILKADSLQKALANFTAAELLDGGEKQY 932
Query: 366 HAEQYGLQ-DAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDG 424
H ++ + A K + P VL + LKRF + D +KI + F L+L
Sbjct: 933 HCQRCKQKVQAVKQLTIHKAPYVLAIHLKRF---YAHDPNIKIKKKVRFDSALNLK---- 1091
Query: 425 KYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQWYKFDDERVTKEDTKR 483
++S +D +V+ Y+L+ VLVHSG H GHYY ++R T ++ WY DD RV+ +
Sbjct: 1092PFVSGSSDGDVK--YSLYGVLVHSGFSTHSGHYYCYVR-TSNNMWYTLDDTRVSHVGEQE 1262
Query: 484 ALEEQ 488
L +Q
Sbjct: 1263VLNQQ 1277
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,122,234
Number of Sequences: 36976
Number of extensions: 455977
Number of successful extensions: 2763
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 2448
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2699
length of query: 1118
length of database: 9,014,727
effective HSP length: 106
effective length of query: 1012
effective length of database: 5,095,271
effective search space: 5156414252
effective search space used: 5156414252
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0546.13


