

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0545.3
(2087 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
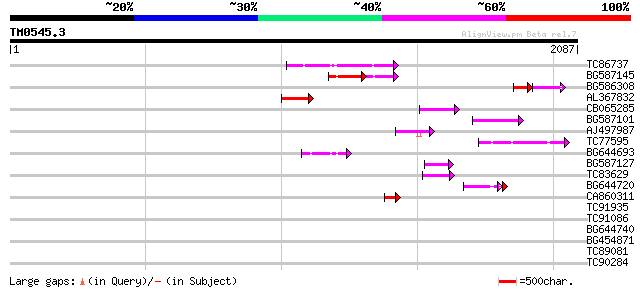
Score E
Sequences producing significant alignments: (bits) Value
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 184 3e-46
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 126 1e-28
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 67 4e-22
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 100 5e-21
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 100 6e-21
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 92 2e-18
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 92 2e-18
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 89 2e-17
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 83 1e-15
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 70 9e-12
TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNa... 55 3e-07
BG644720 47 3e-07
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 48 3e-05
TC91935 41 0.006
TC91086 39 0.028
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 36 0.14
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 34 0.52
TC89081 similar to GP|15982743|gb|AAL09712.1 AT5g17920/MPI7_60 {... 32 2.0
TC90284 homologue to GP|18483235|gb|AAL73979.1 methionine syntha... 32 3.4
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 184 bits (467), Expect = 3e-46
Identities = 134/420 (31%), Positives = 214/420 (50%), Gaps = 8/420 (1%)
Frame = +1
Query: 1020 ALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVELKLPIKEDKK------PVKQLP 1073
A +DP K +L+EF D F + +R L++ +P+ DK P L
Sbjct: 307 APVDPR-KHVPAGVLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLY 483
Query: 1074 RRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATP 1133
++LV +K+ +E LL FI+ + A V+ V K G +R C+D+R LNA T
Sbjct: 484 GMSRQELLV-LKKTLEDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAITK 657
Query: 1134 KDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVM 1193
KD Y +P+ + AG + + LD + ++++ I ED KTAFR G +EW+V
Sbjct: 658 KDRYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRT--RYGLFEWIVC 831
Query: 1194 PFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVV-KSPSRDGHLLHLRKSFER 1252
PF GL A AT+QR +N H+F++ F+ YIDD+++ + S+ H +R+ R
Sbjct: 832 PF-----GLTGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRR 996
Query: 1253 MRKYGLKMNPLKCAFGVIAGDFLGFVVHK-KGIEINKNKAKAILDTSPPTSKKQLQSLLG 1311
+ GL ++P KC F V ++GF++ KG+ + K AI D PP S K +S LG
Sbjct: 997 LADAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLG 1176
Query: 1312 KINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEAEHQKAFDELKKYLSNPPVMIPPIKG 1371
N+ + FI SE T+ + L R K+ FRW AE + AF +LK+ + PV+
Sbjct: 1177 FCNYYKDFIPGYSEITEPLTRLTR--KDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPE 1350
Query: 1372 RPMKLYISATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVK 1431
+ + +G +L QED G + + S+ L+ AE Y + +K L ++ +C++
Sbjct: 1351 AVTTVETDCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAVW-ACLR 1527
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 126 bits (316), Expect = 1e-28
Identities = 65/141 (46%), Positives = 96/141 (67%)
Frame = +2
Query: 1172 EDVSKTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDD 1231
+D+ KTAF GTY + VMPFGLKNAG +TYQR++N +F D + M+VYIDD
Sbjct: 11 DDLEKTAFITDR--GTYCYKVMPFGLKNAG-----STYQRLVNRMFADKLGNTMEVYIDD 169
Query: 1232 IVVKSPSRDGHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKA 1291
++VKS HL HL++ F+ + +Y +K+NP KC FGV +G+FLG++V ++GIE+N +
Sbjct: 170 MLVKSLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQI 349
Query: 1292 KAILDTSPPTSKKQLQSLLGK 1312
AILD P + +++Q L G+
Sbjct: 350 TAILDLPSPKNSREVQRLTGE 412
Score = 81.6 bits (200), Expect = 3e-15
Identities = 50/145 (34%), Positives = 78/145 (53%), Gaps = 7/145 (4%)
Frame = +3
Query: 1294 ILDTSPPTSKKQLQSLL-------GKINFLRRFITNLSEKTKSFSSLLRLKKEDVFRWEA 1346
IL SPP +Q + G+I L RFI+ ++K F LL K F W+
Sbjct: 336 ILSRSPPY*TSLVQRIAERSSDSRGRIAALNRFISRSTDKCLPFYKLLCGNKR--FVWDE 509
Query: 1347 EHQKAFDELKKYLSNPPVMIPPIKGRPMKLYISATDETIGSMLAQEDEDGNERAIFYLSR 1406
+ ++AF++LK+YL+ PPV+ P G + LYI+ + + S+L +ED G ++ IFY S+
Sbjct: 510 KCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIAISSTAVSSVLIREDR-GEQKPIFYTSK 686
Query: 1407 VLNGAETRYTMIEKLCLCLYFSCVK 1431
+ ETRY +EK+ + S K
Sbjct: 687 RMTDPETRYPTLEKMAFAVITSARK 761
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 67.4 bits (163), Expect(2) = 4e-22
Identities = 32/70 (45%), Positives = 43/70 (60%)
Frame = -2
Query: 1853 GLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVG 1912
GLP I TD G+ F+ K F E W I+L T++P Y Q+NGQ EA+NKI+I +KK +
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 1913 RKPKRWHESL 1922
K W + L
Sbjct: 502 LKKGCWADEL 473
Score = 57.8 bits (138), Expect(2) = 4e-22
Identities = 39/121 (32%), Positives = 63/121 (51%)
Frame = -3
Query: 1925 VLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYLQSCRIQRQEEIPSEVYWNMMLDEMVN 1984
VLW++R +P AT +TPF +A+ EA+ PAEV + S R + E+ + + + +
Sbjct: 465 VLWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQ-NIELNNDRLFNALET 289
Query: 1985 LDEERLLALDVLTRQKDRIAKAYNKKVRDRSFITGDYVWKVILPMDKKDRVYGKWTPNWE 2044
++E R AL + + +I YNK V+ + GD V + + K+ GK NWE
Sbjct: 288 IEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFE-NTKELNAGKLGTNWE 112
Query: 2045 G 2045
G
Sbjct: 111 G 109
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 100 bits (250), Expect = 5e-21
Identities = 46/119 (38%), Positives = 79/119 (65%)
Frame = -1
Query: 999 QDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLNRELVEL 1058
Q+ +E ++LG KR + A ++ +K ++ +LL+E+ D FA Y++MPGL+ ++VE
Sbjct: 357 QEEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEH 178
Query: 1059 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNG 1117
++P K + PV+ RR HPD+ +KIK E+++ + F+ T Y +W+AN+VPV KK+G
Sbjct: 177 RIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 100 bits (249), Expect = 6e-21
Identities = 57/147 (38%), Positives = 83/147 (55%)
Frame = -2
Query: 1510 WKLYFDGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGA 1569
W L FDG+ + G GIG I+SPQG F RI C+NN EYEA I G+E + +
Sbjct: 534 WGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAIDMRI 355
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
K++ I GDS LV+ Q+ E++ +L Y A LL F ++ + H+PR ENQ A+ L
Sbjct: 354 KHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADAL 175
Query: 1630 AQIASGYMVDKLKLTELIKIKEKLSPS 1656
A ++S + V+ +IK++ PS
Sbjct: 174 ATLSSMFRVNHWNDVPIIKVQRLERPS 94
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 92.4 bits (228), Expect = 2e-18
Identities = 53/188 (28%), Positives = 91/188 (48%)
Frame = +2
Query: 1704 LFKKNINGTLLKCLSENDAFMAVSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIE 1763
L+K+ + ++C++E + + H H A +K + + G +WPT+ KD
Sbjct: 41 LYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDAHS 220
Query: 1764 YARGCQDCQKHSGIQHVPASELHSIIKPWPFRGWAIDLIGEIHPASSKQHKYIIVAVDYF 1823
+ C CQ+ I + I++ F W ID +G SS +KYI+VAVDY
Sbjct: 221 FISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPF--PSSYNNKYILVAVDYV 394
Query: 1824 TKWVEAIPLQNVTQETVIEFIQNHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLL 1883
+KWVEAI V++ ++ I RFG+P + +D G+ F+ + + G++
Sbjct: 395 SKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKKNGVRHK 574
Query: 1884 TSTPYYAQ 1891
+T Y+ Q
Sbjct: 575 VATAYHPQ 598
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 92.4 bits (228), Expect = 2e-18
Identities = 54/167 (32%), Positives = 83/167 (49%), Gaps = 25/167 (14%)
Frame = -2
Query: 1420 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1479
K C L ++ +L++Y+ + S D IK++ KP L RI +W + L+EY + Y
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1480 LKAIKGQAVADFLADHTV----------PKEIVTYVGIQP---------------WKLYF 1514
KAIKG +AD LA + P E + Y+ ++ W L F
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1515 DGSSHKNGTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGL 1561
DG+ + G GIG +++P+GA F R+ +C+NN EYEA I G+
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 89.0 bits (219), Expect = 2e-17
Identities = 84/347 (24%), Positives = 142/347 (40%), Gaps = 14/347 (4%)
Frame = +2
Query: 1726 VSAAHDGLCGAHQAGAKMKWILFRQGLYWPTIMKDCIEYARGCQDCQKHSGIQHVPASEL 1785
V +HD H I+ R+ +WP + + R C C G H+
Sbjct: 182 VQESHDSTAAGHPGRNGTLEIVSRK-FFWPGQSQTVRRFVRNCDVC----GGIHIWRQAK 346
Query: 1786 HSIIKPWPFRG-----WAIDLIGEIHPASSKQHKYIIVAVDYFTKWVEAIPLQNVTQETV 1840
+KP P ++D I + P + +Y+ V VD +K V + + E
Sbjct: 347 RGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAEAC 526
Query: 1841 IE-FIQNHIVYRF-GLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEA 1898
+ F+ H YRF G+P+SI +D+G+ +VGR F G+ L ST Y+ Q +G E
Sbjct: 527 AQRFLSCH--YRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 1899 ANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEVYL 1958
N+ + ++++ +V W + L V A RN + G TPF + +G V P
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 1959 QSCRIQRQEEIPSEVYWNMMLDEMVNLDEERLLALDVLTRQKDRIAKAYNKKVRDRSFIT 2018
+ + + E +++ M D + +++ Q+ A A ++ +
Sbjct: 878 DTGGVVSEGEAAAQLLVKRM------KDVTGFIQAEIVAAQQRSEASANKRRCPADRYQV 1039
Query: 2019 GDYVW------KVILPMDKKDRVYGKW-TPNWEGPFTVEKTLPNNAY 2058
GD VW K P K D ++ K+ + P VE +P Y
Sbjct: 1040 GDKVWLNVSNYKSPRPSKKLDWLHHKYEVTRFVTPHVVELNVPGTVY 1180
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 82.8 bits (203), Expect = 1e-15
Identities = 56/184 (30%), Positives = 102/184 (55%)
Frame = +2
Query: 1075 RFHPDVLVKIKEEIERLLKCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPK 1134
R +P L +K +++ LL+ FI+ + Y + V+ + KK+G +R+ ID+ LN K
Sbjct: 92 RINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRMSIDYPQLNNVNIK 268
Query: 1135 DEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVSKTAFRCPGALGTYEWVVMP 1194
+Y +P+ + + D+ G ++ +D G +Q + EDV KTAFR G YE +VM
Sbjct: 269 IKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRI--RYGHYEILVMS 442
Query: 1195 FGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKSFERMR 1254
FG N + + +MN +F D++++ + V+ +DI++ S + + H HLR + + ++
Sbjct: 443 FG*TNPPM-----AFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRLALKVLK 607
Query: 1255 KYGL 1258
GL
Sbjct: 608 DIGL 619
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 70.1 bits (170), Expect = 9e-12
Identities = 39/109 (35%), Positives = 58/109 (52%)
Frame = +3
Query: 1526 GMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGAKNVVIKGDSELVVKQL 1585
G+ + SP K FR+E + SNNE YEALI+G+ + L +N+ DS+LV Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1586 TKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANELAQIAS 1634
+ EY+ E + Y L K + + +PR EN +A+ LA +AS
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALAS 329
>TC83629 weakly similar to GP|6692261|gb|AAF24611.1| putative RNase H
{Arabidopsis thaliana}, partial (21%)
Length = 1071
Score = 55.1 bits (131), Expect = 3e-07
Identities = 38/122 (31%), Positives = 57/122 (46%), Gaps = 5/122 (4%)
Frame = +2
Query: 1519 HKNGTGIGMFIMSPQGAPTKFKFRIE-GNCSNNEVEYEALISGLEILLALGAKNVVIKGD 1577
++ G G +++ G+ + FR G+ + EY AL+ GL+ G K V KGD
Sbjct: 587 NRGPAGAGALLLAEDGS-LLYGFRQGLGHQTKESAEYRALLLGLKHASMKGFKYVTAKGD 763
Query: 1578 SELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIEN----QEANELAQIA 1633
SELV+ Q+ +K EHL K +A L F+ I H+ R N + AN +
Sbjct: 764 SELVINQILDPWKIKDEHLKKLCAEALELSDNFHSFRIQHISRERNYGAVRNANRAINLT 943
Query: 1634 SG 1635
G
Sbjct: 944 DG 949
>BG644720
Length = 678
Score = 47.4 bits (111), Expect(2) = 3e-07
Identities = 44/150 (29%), Positives = 62/150 (41%), Gaps = 7/150 (4%)
Frame = -3
Query: 1671 WRKPIVEY------LQNPVGSTDRKVKYRALSYTILGNELFKKNINGTLLKCLSENDAFM 1724
WR+PI+ Y L+NP ST ++ L+ G LL CL E +
Sbjct: 463 WRQPIIIYMCYGIFLENPRRST--YIRRHTPHLH*YNETLYI*LFEGVLL*CLGEEETIQ 290
Query: 1725 AVSAAHDGLCGAHQAGAKMKWILFRQGLYWPT-IMKDCIEYARGCQDCQKHSGIQHVPAS 1783
A AH +CG+H+ +K+ + + R G YW T IM A Q P
Sbjct: 289 AFQ*AHSRVCGSHK--SKLYFHIKRMGYYWNT*IMLKGALLANFIQQ----------PPE 146
Query: 1784 ELHSIIKPWPFRGWAIDLIGEIHPASSKQH 1813
LH I F W +++ G + P SS H
Sbjct: 145 VLHPTITS*LFESWELNVFG-LFPKSSGGH 59
Score = 26.9 bits (58), Expect(2) = 3e-07
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = -1
Query: 1815 YIIVAVDYFTKWVEAIPL 1832
YI+ AV YF KWVE + L
Sbjct: 54 YILAAVYYF*KWVEVVAL 1
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 48.1 bits (113), Expect = 3e-05
Identities = 22/57 (38%), Positives = 36/57 (62%)
Frame = +1
Query: 1380 ATDETIGSMLAQEDEDGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKYYI 1436
A D IG+ L+Q+DE+ +E I Y SR+L AE YT++E+ CL ++ ++Y+
Sbjct: 1 APDWGIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYL 171
>TC91935
Length = 1144
Score = 40.8 bits (94), Expect = 0.006
Identities = 17/40 (42%), Positives = 25/40 (62%)
Frame = +3
Query: 1395 DGNERAIFYLSRVLNGAETRYTMIEKLCLCLYFSCVKLKY 1434
DG + A +YLS +L T+YT I+KLCL ++C + Y
Sbjct: 984 DGTKIAAYYLSWILKDVGTKYTHIDKLCLSFLYACNNISY 1103
>TC91086
Length = 697
Score = 38.5 bits (88), Expect = 0.028
Identities = 13/22 (59%), Positives = 21/22 (95%)
Frame = -1
Query: 1096 FIRTARYVDWLANVVPVIKKNG 1117
FI+T+RY+ WL+N++P+IKK+G
Sbjct: 676 FIQTSRYIKWLSNILPIIKKHG 611
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 36.2 bits (82), Expect = 0.14
Identities = 15/36 (41%), Positives = 23/36 (63%)
Frame = -1
Query: 1112 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVD 1147
V KK+G R+CID+R N T K++Y +P + + D
Sbjct: 223 VRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 34.3 bits (77), Expect = 0.52
Identities = 19/71 (26%), Positives = 34/71 (47%)
Frame = +2
Query: 1879 GIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHESLSQVLWAYRNSPTEATG 1938
G L S+ Y+ ++GQ EA NK ++ + P +W ++ + Y S +
Sbjct: 65 GTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEYWYNTSYNISAA 244
Query: 1939 TTPFRLAYGQE 1949
TPF+ YG++
Sbjct: 245 MTPFKALYGRD 277
>TC89081 similar to GP|15982743|gb|AAL09712.1 AT5g17920/MPI7_60 {Arabidopsis
thaliana}, partial (40%)
Length = 1219
Score = 32.3 bits (72), Expect = 2.0
Identities = 20/83 (24%), Positives = 37/83 (44%)
Frame = +1
Query: 1180 RCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1239
R G +W V F + N G+++ + + + F+D I + + + D I +++
Sbjct: 472 RKSGQADYLDWAVHSFRITNVGVQDTTQIHTHMCYSHFNDIIHSIIDMDADVITIENSRS 651
Query: 1240 DGHLLHLRKSFERMRKYGLKMNP 1262
D LL + F KYG + P
Sbjct: 652 DEKLLRV---FRDGVKYGAGIGP 711
>TC90284 homologue to GP|18483235|gb|AAL73979.1 methionine synthase protein
{Sorghum bicolor}, partial (51%)
Length = 1377
Score = 31.6 bits (70), Expect = 3.4
Identities = 18/73 (24%), Positives = 34/73 (45%)
Frame = +2
Query: 1190 WVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKS 1249
W V F + N G+++ + + + F+D I + + + D I +++ D LL +
Sbjct: 755 WAVHSFRITNCGVQDTTQIHTHMCYSNFNDIIHSIIDMDADVITIENSRSDEKLLSV--- 925
Query: 1250 FERMRKYGLKMNP 1262
F KYG + P
Sbjct: 926 FREGVKYGAGIGP 964
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.143 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,728,157
Number of Sequences: 36976
Number of extensions: 913944
Number of successful extensions: 6804
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 3724
Number of HSP's successfully gapped in prelim test: 317
Number of HSP's that attempted gapping in prelim test: 2891
Number of HSP's gapped (non-prelim): 4356
length of query: 2087
length of database: 9,014,727
effective HSP length: 111
effective length of query: 1976
effective length of database: 4,910,391
effective search space: 9702932616
effective search space used: 9702932616
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0545.3


