

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
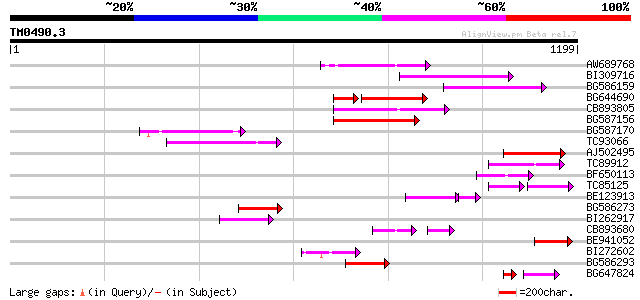
Query= TM0490.3
(1199 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 199 4e-51
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 182 5e-46
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 159 6e-39
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 127 9e-39
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 153 4e-37
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 149 6e-36
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 140 4e-33
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 134 2e-31
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 126 6e-29
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 107 3e-23
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 100 3e-21
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 64 4e-21
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 79 1e-20
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 94 3e-19
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 86 1e-16
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 64 1e-14
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 76 7e-14
BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%) 73 8e-13
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 72 2e-12
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 55 3e-12
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 199 bits (507), Expect = 4e-51
Identities = 105/232 (45%), Positives = 138/232 (59%)
Frame = +1
Query: 658 TRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWEL 717
TR+ G KPK V + +P + P LSDP W AM++E+ ALI + TW+L
Sbjct: 4 TRTKTGKLKPK------VFLTEPCPQDCENLP---LSDPRWLQAMKTEYKALIDNKTWDL 156
Query: 718 VPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIR 777
VP P I C W++R K+ +G ++KARLV G SQ G D ETFSPV+KP TIR
Sbjct: 157 VPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVTIR 336
Query: 778 TVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQ 837
+L+IA++ W I Q+D+ NAFL+G L E +YM P GF + + VC+L KSLYGLKQ
Sbjct: 337 LILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGF-EAANKSLVCKLNKSLYGLKQ 513
Query: 838 APRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASS 889
APRAWY+ GF S D SL I+ Q Y+ +YVDDI++ SS
Sbjct: 514 APRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYLXIYVDDILITGSS 669
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 182 bits (463), Expect = 5e-46
Identities = 99/240 (41%), Positives = 135/240 (56%)
Frame = +2
Query: 825 VCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDII 884
VC L+KS+YGLKQA R WY + ++ + S G+ S+SD SLF + S +L+YVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 885 LVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGM 944
L + + L F +KDLG L YFLG+ V R G+ L+Q Y E++ +G
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSGN 379
Query: 945 ASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHA 1004
+ + TP D KL +S D + YR L G L YLT TRPDIS+AVQQ+ +
Sbjct: 380 LAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVSK 559
Query: 1005 PHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLG 1064
P H A RVL+Y++ GL + L S+ D+DW CP TR+S +GY VFLG
Sbjct: 560 PQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVFLG 739
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 159 bits (402), Expect = 6e-39
Identities = 82/217 (37%), Positives = 124/217 (56%)
Frame = +1
Query: 918 IAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSL 977
+ V ++ G+++ Q Y T+++ R GM N S P+ + KL DA+ Y+ +
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 978 DGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVET 1037
G L YL TRPD+ Y + + M+ P HM A+KRVLRY+ GT+ G+ + E
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 1038 LVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSES 1097
L +YTD+D+ G D R+STSGY L +SWSSK+QP ++ S+ +AE+ A +S
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 1098 CWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHH 1134
W+R +L +L + S + ++CDN S I LS NPV H
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 127 bits (319), Expect(2) = 9e-39
Identities = 59/139 (42%), Positives = 94/139 (67%)
Frame = -2
Query: 745 RYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDL 804
R K++LV G +Q G+D DE FSPV + IR +++ A + ++Q+DV++AF++GDL
Sbjct: 418 RNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFINGDL 239
Query: 805 HETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSL 864
E +++ P GF D P++V RL K+LYGLKQAPRAWY+R + ++ GF+ D++L
Sbjct: 238 KEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTL 59
Query: 865 FIFRQGSDIAYILLYVDDI 883
F+ ++ ++ I +YVDDI
Sbjct: 58 FLLKRE*ELLIIQVYVDDI 2
Score = 52.8 bits (125), Expect(2) = 9e-39
Identities = 24/51 (47%), Positives = 31/51 (60%)
Frame = -3
Query: 686 PHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHK 736
P N K+AL D +W ++MQ E + RS W LVPRP VI W+FR+K
Sbjct: 585 PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 153 bits (386), Expect = 4e-37
Identities = 88/248 (35%), Positives = 135/248 (53%), Gaps = 4/248 (1%)
Frame = +3
Query: 686 PHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFER 745
P ++A+ W+++M +E A R+NTWEL I WIF+ K NG E+
Sbjct: 42 PTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNENGEIEK 221
Query: 746 YKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWP---IHQLDVQNAFLHG 802
YKARLV G SQ GVD E F+PV + TIR V+++A I + ++ +
Sbjct: 222 YKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQIKRDGVCIS*M*KAHSCMEN 401
Query: 803 DLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDH 862
+ + + ++H + R R+K++LYGLKQAPRAWY R Y + GF +H
Sbjct: 402 *MRKFLLINHRVM*RRV----IS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEH 569
Query: 863 SLFI-FRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVT 921
+LF+ +G I I LYVDD+I + + ++ + F + EF M DLG + YFLG+ VT
Sbjct: 570 TLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVT 749
Query: 922 RHAGGLFL 929
++ G+++
Sbjct: 750 QNEKGIYI 773
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 149 bits (376), Expect = 6e-36
Identities = 75/181 (41%), Positives = 111/181 (60%)
Frame = -1
Query: 685 LPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFE 744
+P + ++A+ D WK ++ +E A+I+++TW P + WIF K +++G E
Sbjct: 561 IPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRWIFTIKYKADGSIE 382
Query: 745 RYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDL 804
R K RLV G + G D ETF+PV K TIR VLS+A++ W + Q+DV+NAFL G+L
Sbjct: 381 RKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLWQMDVKNAFLQGEL 202
Query: 805 HETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSL 864
+ +YM+ P G V RLKK++YGLKQ+PRAWY + + ++ GFR S DH+L
Sbjct: 201 EDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTL 22
Query: 865 F 865
F
Sbjct: 21 F 19
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 140 bits (352), Expect = 4e-33
Identities = 76/235 (32%), Positives = 125/235 (52%), Gaps = 11/235 (4%)
Frame = -3
Query: 275 GDLYPVTRSSPFAGLASS-----------VWHNRLGHPASSALNHLRNNKLIFCEPSRSS 323
GDLY + + P + S +WH RLGHP ALN L ++F +
Sbjct: 671 GDLYMLEKLDPVSNYKCSFTSSSSLNKDALWHARLGHPHGRALN-LMLPGVVF-----EN 510
Query: 324 SVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLW 383
C++C+LGKH + F ++ + FD++++DLWT+P LS H+Y+V F+D+ + + W
Sbjct: 509 KNCEACILGKHCKNVFPRTSTVYENCFDLIYTDLWTAPSLSRDNHKYFVTFIDEKSKYTW 330
Query: 384 TFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSC 443
I K +V + F + + A IK L+ DNG EY + +F + D +G++ + SC
Sbjct: 329 LTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFKSHLDHHGILHQTSC 150
Query: 444 PHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQ 498
P+T QNG A+RK + + + R+L+ ++V A YL+N +P K L+
Sbjct: 149 PYTPQQNGVAKRKNKHLMEVARSLMFQANV---------STACYLINWIPTKVLR 12
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 134 bits (338), Expect = 2e-31
Identities = 80/248 (32%), Positives = 124/248 (49%), Gaps = 5/248 (2%)
Frame = +1
Query: 332 GKHVRLPFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKK 390
G ++ FS++ T D +HSDLW S V S G RY + +DD +W + + K
Sbjct: 43 GNRKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYK 222
Query: 391 SQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQN 450
++ + TF L++TQ N+K L DN E+ + F+ +C +G+ + P QN
Sbjct: 223 NETFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQN 402
Query: 451 GKAERKIRTINNMIRTLLAHSSV--PPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYH 508
G AER IRT+ R +L+++ + W A A +L+N P L P +
Sbjct: 403 GVAERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSG 582
Query: 509 RDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYK--CYDLSHRKILI 566
YS+LR+FGC P + KL PR+ C+FL Y +GY+ C D +K+++
Sbjct: 583 NLVDYSNLRIFGC---PAYALVNDGKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLIL 753
Query: 567 SRHVIFDE 574
SR V F+E
Sbjct: 754 SRDVTFNE 777
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 126 bits (316), Expect = 6e-29
Identities = 59/132 (44%), Positives = 86/132 (64%)
Frame = +2
Query: 1044 ADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNL 1103
+DW G +TR+STSGY LG ISWSSK+QP ++ S+AEAEY + +++ W+R +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1104 LLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPS 1163
L +H + T ++CDN SAI LS NPV H R+KHI++ H +RE + + + + P+
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1164 RHQIADIFTKGL 1175
+IADIFTK L
Sbjct: 362 EEKIADIFTKPL 397
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 107 bits (267), Expect = 3e-23
Identities = 58/164 (35%), Positives = 94/164 (56%), Gaps = 2/164 (1%)
Frame = +1
Query: 1012 ALKRVLRYVRGTLTYGLHLYPSPVE--TLVSYTDADWGGCPDTRRSTSGYCVFLGDNLIS 1069
ALK VL+Y+ +L L + E L Y DAD+ G DTR+S SG+ L IS
Sbjct: 4 ALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTIS 183
Query: 1070 WSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSG 1129
W + +Q ++ S+ +AEY V ++ W++ ++ EL + +HCD+ SAI+L+
Sbjct: 184 WKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGIT-QEYVKIHCDSQSAIHLAN 360
Query: 1130 NPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTK 1173
+ V+H+RTKHI++ +HF+R+ + + V + S AD+FTK
Sbjct: 361 HQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 100 bits (249), Expect = 3e-21
Identities = 55/124 (44%), Positives = 74/124 (59%), Gaps = 4/124 (3%)
Frame = +1
Query: 988 RPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYP----SPVETLVSYTD 1043
RPDI Y+V + MH P H++A R+LRYVRGT+ YGL L+P S V L+ Y+D
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGL-LFPYGAKSEVYELICYSD 297
Query: 1044 ADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNL 1103
+DW G RRSTSGY D ISW +K+QP + SS EAEY ++ W+ ++
Sbjct: 298 SDWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSV 468
Query: 1104 LLEL 1107
+ EL
Sbjct: 469 IKEL 480
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 64.3 bits (155), Expect(2) = 4e-21
Identities = 36/96 (37%), Positives = 53/96 (54%)
Frame = +3
Query: 1096 ESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQ 1155
E+ W++ L+ EL Q T V+CD+ SA++++ NP H RTKHI + HFVRE V G
Sbjct: 249 EAIWMQRLMEELGHKQEQIT-VYCDSQSALHIARNPAFHSRTKHIGIQYHFVREVVEEGS 425
Query: 1156 ARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGE 1191
+ + + +AD TK + F RSS + E
Sbjct: 426 VDMQKIHTNDNLADAMTKSINTDKFIWCRSSYGLLE 533
Score = 56.6 bits (135), Expect(2) = 4e-21
Identities = 29/75 (38%), Positives = 45/75 (59%)
Frame = +1
Query: 1013 LKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSS 1072
+KR++RY++GT + S + T+ Y D+D+ G D R+ST+GY L +SW S
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSEL-TVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLS 177
Query: 1073 KRQPTLSRSSAEAEY 1087
K Q ++ S+ EAEY
Sbjct: 178 KLQTVVALSTTEAEY 222
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 78.6 bits (192), Expect(2) = 1e-20
Identities = 44/119 (36%), Positives = 61/119 (50%), Gaps = 2/119 (1%)
Frame = +1
Query: 837 QAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDI--AYILLYVDDIILVASSHDLRK 894
Q+PR W+ RF V G+ +DH++FI + S + A +++YVDDI L K
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFI-KHSSTVKKAILIVYVDDIFLTGDHGK*IK 177
Query: 895 SFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATP 953
LLA EF +KDLG L YFLG+ V R G +SQ Y +++ M C P
Sbjct: 178 RLKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 40.8 bits (94), Expect(2) = 1e-20
Identities = 21/47 (44%), Positives = 26/47 (54%)
Frame = +2
Query: 949 PSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAV 995
PS TP+D KL + D Y+ L G L YL+ TRPDIS+ V
Sbjct: 341 PSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVV 481
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 94.0 bits (232), Expect = 3e-19
Identities = 42/92 (45%), Positives = 61/92 (65%)
Frame = -2
Query: 485 ATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVF 544
A YL+N +P + L++ +P ++L R PS +++RVFGCLC+ L P NKL+ RS +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 545 LGYPMNHRGYKCYDLSHRKILISRHVIFDETR 576
+GY +GYKCYD R++L+SR V F E R
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEER 429
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 85.5 bits (210), Expect = 1e-16
Identities = 42/114 (36%), Positives = 62/114 (53%)
Frame = +1
Query: 445 HTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQ 504
HT QNG AER RT+ R +L + + SFW A++ A Y++N P + +P +
Sbjct: 82 HTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPME 261
Query: 505 LLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYD 558
+ + YS L VFGC + ++ S KL P+S C+FLGY N +GY +D
Sbjct: 262 MWKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 64.3 bits (155), Expect(2) = 1e-14
Identities = 42/94 (44%), Positives = 50/94 (52%)
Frame = -2
Query: 767 FSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVC 826
F P+VK TI +LSI + + LDV+ AFL GDL E IYMH P GF V
Sbjct: 554 FVPIVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVEDIYMHQPEGF-S*EVGKMVG 378
Query: 827 RLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTS 860
+LKKS+YGLKQ PR + GFR S
Sbjct: 377 KLKKSMYGLKQGPRQCI*SLKALCTRKGFRSEIS 276
Score = 34.7 bits (78), Expect(2) = 1e-14
Identities = 20/59 (33%), Positives = 31/59 (51%), Gaps = 2/59 (3%)
Frame = -3
Query: 883 IILVASSHDLRKSFMALLASEFAMKDLGPLSYFLG--IAVTRHAGGLFLSQSTYATEII 939
+++V S+ D K+ + E MKDLGP +G I + + G L LSQ Y T ++
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVL 95
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 76.3 bits (186), Expect = 7e-14
Identities = 37/80 (46%), Positives = 53/80 (66%)
Frame = +2
Query: 1110 PLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIAD 1169
P+SQ+ L+ CD +SA YL+ NPV+H R KHI +DIHFVR+ V +G+ +V HV + Q+AD
Sbjct: 2 PISQSGLLRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQLAD 181
Query: 1170 IFTKGLPRLLFDDFRSSLSV 1189
TK L + R+ + V
Sbjct: 182 CLTKPLSKSRHQLLRNKIGV 241
>BI272602 GP|11602812|db avenaIII {Gallus gallus}, partial (3%)
Length = 488
Score = 72.8 bits (177), Expect = 8e-13
Identities = 52/133 (39%), Positives = 69/133 (51%), Gaps = 7/133 (5%)
Frame = +1
Query: 617 VQPSSPTDSSPPLSALVSPTASPSPLPLPPVPPAPPVRTMTT-------RSMHGISKPKK 669
+Q SSP +SP + L + T SP+P P PP P A P TT RS + I K K
Sbjct: 61 IQISSP--ASPTSNELANITTSPTPPP-PPPPTARPHNNHTTNVGGIITRSQNNIVKTIK 231
Query: 670 PFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRC 729
+L V P P N QAL D +W+S MQ EF+AL +NTW+LV R N++ C
Sbjct: 232 KLNLHVR---PLCPIEPSNITQALRDHDWRSIMQDEFDALQNNNTWDLVSRSFAQNLVGC 402
Query: 730 MWIFRHKKQSNGL 742
F ++ +S L
Sbjct: 403 KLGFSYQTKSGWL 441
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 71.6 bits (174), Expect = 2e-12
Identities = 37/92 (40%), Positives = 56/92 (60%)
Frame = +2
Query: 711 RSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPV 770
+ T +LV +P V I WI++ K+ +G +YKARLV G + G+D DE F+PV
Sbjct: 47 QKQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPV 226
Query: 771 VKPATIRTVLSIALSRSWPIHQLDVQNAFLHG 802
V+ TI +L++A + IH +DV+ AFL+G
Sbjct: 227 VRIETI*LLLALAATNGC*IHHIDVKIAFLNG 322
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 55.1 bits (131), Expect(2) = 3e-12
Identities = 28/78 (35%), Positives = 45/78 (56%), Gaps = 1/78 (1%)
Frame = -3
Query: 1087 YRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAI-YLSGNPVHHQRTKHIEMDIH 1145
YR + + + E W+ LL +L F + +++CDN SA +++ N +RTKHIE+D H
Sbjct: 374 YRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAARHIAANSSFLERTKHIELDCH 195
Query: 1146 FVREKVVRGQARVLHVPS 1163
VR K+ +LH+ S
Sbjct: 194 IVRVKLQLKLFHILHILS 141
Score = 35.8 bits (81), Expect(2) = 3e-12
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = -2
Query: 1045 DWGGCPDTRRSTSGYCVFLGDNLISWSS 1072
D C DT +S S +C+FLGD+LI W S
Sbjct: 486 D*SSCLDT*KSISYFCIFLGDSLICWKS 403
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,876,076
Number of Sequences: 36976
Number of extensions: 947364
Number of successful extensions: 15424
Number of sequences better than 10.0: 666
Number of HSP's better than 10.0 without gapping: 7599
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11310
length of query: 1199
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1092
effective length of database: 5,058,295
effective search space: 5523658140
effective search space used: 5523658140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0490.3


