

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
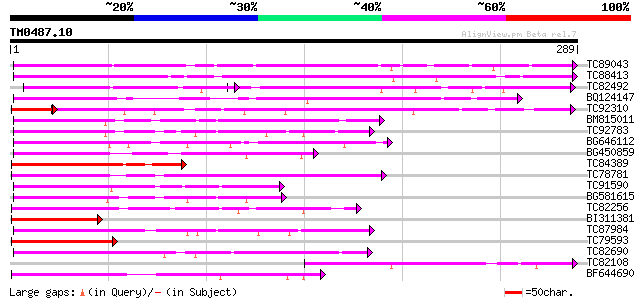
Query= TM0487.10
(289 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89043 similar to GP|10178241|dbj|BAB11673. gene_id:MDJ22.9~unk... 147 4e-36
TC88413 weakly similar to GP|10176786|dbj|BAB09900. gene_id:MIK1... 124 3e-29
TC82492 similar to GP|14596145|gb|AAK68800.1 Unknown protein {Ar... 80 2e-26
BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.... 103 6e-23
TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein... 96 4e-22
BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.... 90 1e-18
TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.... 77 8e-15
BG646112 weakly similar to GP|10177368|dbj contains similarity t... 76 2e-14
BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis ... 76 2e-14
TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 75 4e-14
TC78781 weakly similar to PIR|T02291|T02291 hypothetical protein... 74 9e-14
TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ2... 70 9e-13
BG581615 similar to GP|10177368|dbj contains similarity to heat ... 70 1e-12
TC82256 similar to GP|20042913|gb|AAM08741.1 Unknown protein {Or... 67 1e-11
BI311381 weakly similar to PIR|T46128|T461 hypothetical protein ... 66 2e-11
TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 65 2e-11
TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity... 65 4e-11
TC82690 weakly similar to GP|9802755|gb|AAF99824.1| Unknown prot... 64 7e-11
TC82108 similar to GP|21537320|gb|AAM61661.1 unknown {Arabidopsi... 63 2e-10
BF644690 weakly similar to GP|20042923|gb Putative copia-type po... 62 3e-10
>TC89043 similar to GP|10178241|dbj|BAB11673. gene_id:MDJ22.9~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 1413
Score = 147 bits (372), Expect = 4e-36
Identities = 112/306 (36%), Positives = 169/306 (54%), Gaps = 19/306 (6%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS LPD+ILC ILSFLPTK A T++LSKRW PL++ +T+L FD+ + + RF
Sbjct: 62 DRISHLPDDILCRILSFLPTKLAFTTTVLSKRWTPLYKLLTSLSFDDESV-LDEDTFLRF 238
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDIC-IYSLS 121
+ V V + D ++ + S +H N+++W+ A +R E+ ++ +
Sbjct: 239 CRFVDTVTFSTDLIKTLHLNCGSPNWKH----FNLDLWIGTA-KRHPVENFNLVGTWRSI 403
Query: 122 PLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPV 181
PL SIF +LVVLKL+ L + +VDLP LK+LHL + L + ++L GCPV
Sbjct: 404 PLR-PSIFRFPSLVVLKLKTLKIIVGNITVDLPLLKILHLDRVYLKNKTNFNKILYGCPV 580
Query: 182 LEDLKAKAIYFK-------------KYKETGFEYKTLSKLVRADISGTSTDLFRMEVINN 228
LEDL A IY+K K TG E+K L KL+R I + D I+N
Sbjct: 581 LEDLIAN-IYYKEPTPEPDEVFTLSKATATG-EFKILPKLIRVQI---NADEVPFRAIHN 745
Query: 229 VDFLRIHEIYIRIPHDD-----QFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQV 283
V+FL + + R+P + + +F NL ++L Y+++ +W V+E L +CP +QV
Sbjct: 746 VEFLAL-TMRSRLPDPEINSYNILSPIFRNLILLQLCMYNFH-HWDHVMEVLQHCPNIQV 919
Query: 284 LVINQV 289
L IN++
Sbjct: 920 LRINKL 937
>TC88413 weakly similar to GP|10176786|dbj|BAB09900.
gene_id:MIK19.29~pir||T01344~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1288
Score = 124 bits (312), Expect = 3e-29
Identities = 92/294 (31%), Positives = 154/294 (52%), Gaps = 7/294 (2%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS LPDE+LC ILS LP K+ + TS+LS+RW+ L +T ++ D+ ++ + + R+
Sbjct: 247 DRISILPDELLCRILSLLPAKQIMVTSLLSRRWRSLRPRMTEINIDDTSYIHDRDAYDRY 426
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSP 122
+ + PI K ++ +S S + + +W+N +H+D+ +
Sbjct: 427 YHVIALFLFEMKIHHPIMK-TVTILSA-SPFFNPLPLWLNCL----KVQHLDVTSSATLC 588
Query: 123 LNLS-SIFSCKTLVVLKLEGLDLN-TTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCP 180
L + + + LVVLKL L ++ SS +LP LK+LHL + + + L ++L+ P
Sbjct: 589 LCVPYKVLTSTALVVLKLNALTIDYVHRSSTNLPSLKILHLTQVHFLKLKFLIKILSMSP 768
Query: 181 VLEDLKAKAIYFKK---YKETGFEYKTLSKLVRADISGT--STDLFRMEVINNVDFLRIH 235
+LEDL K + ++ K KL+RADIS + S L +++ NV FLR
Sbjct: 769 LLEDLLLKDLQVTDNTLAQDDAAALKPFPKLLRADISDSCISPLLLPLKLFYNVHFLRSQ 948
Query: 236 EIYIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVINQV 289
+ D QF +LTH++L +D+ WI +++F+ CP LQ L I ++
Sbjct: 949 LQTLEEQQDTQFL----SLTHLDLS-FDHGYYWISLIKFICACPSLQTLTIRKI 1095
>TC82492 similar to GP|14596145|gb|AAK68800.1 Unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 1286
Score = 80.1 bits (196), Expect(2) = 2e-26
Identities = 68/201 (33%), Positives = 98/201 (47%), Gaps = 24/201 (11%)
Frame = +3
Query: 112 HVDICIYSLSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQL-SQRR 170
H+ + + L+P+ IF+ +TLV+LKL+ L + DLP LK L L+ +Q+
Sbjct: 318 HLGMYGHILTPI----IFTSRTLVILKLQNLKIVADNLCFDLPSLKTLVLEREYFKNQKN 485
Query: 171 CLAQLLNGCPVLEDLKAK----AIYFKKYKETGFEYKTL--SKLVRADISGTSTDLFRME 224
+LLN CP+LEDL I +K E+K+L SKLVRA I +
Sbjct: 486 DFVKLLNACPILEDLHTYYPRLRIMKRKDNNEVQEFKSLFLSKLVRAYIHSMDVPF---D 656
Query: 225 VINNVDFLRI---------------HEIYIRIPHDDQFTDM--FHNLTHIELGYYDYNTN 267
INNV++L I H I+I + F D+ F NL IEL ++
Sbjct: 657 AINNVEYLCIIQDEEERSYGMNIEEHSFEIKI-EEASFKDIPVFQNLICIELWFFSVFRG 833
Query: 268 WIEVVEFLNYCPKLQVLVINQ 288
W +VE L +CPKLQ+ I +
Sbjct: 834 WDGIVELLRHCPKLQIFFIRK 896
Score = 56.6 bits (135), Expect(2) = 2e-26
Identities = 40/113 (35%), Positives = 60/113 (52%), Gaps = 3/113 (2%)
Frame = +1
Query: 8 LPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQSVH 67
LP+E+L +ILSFLPTK A T +LSKRW PL + + + FD + +HS +
Sbjct: 4 LPNELLRHILSFLPTKLAFTTMLLSKRWTPLSYTPSVVRFDVVP-ELDEAFHSFCLFVDT 180
Query: 68 RVIHTPDFLQPIQKLRMSFVSRHSEVPSN---VNVWVNAAVQRGGFEHVDICI 117
++ QPI+ ++F R+ V S+ VN WV AA+ R + C+
Sbjct: 181 LMLSLRSLNQPIKTFSLNF--RNVVVDSDQPIVNAWVEAALLRRVRNFISACM 333
>BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.13~similar
to unknown protein {Arabidopsis thaliana}, partial (11%)
Length = 677
Score = 103 bits (258), Expect = 6e-23
Identities = 91/261 (34%), Positives = 125/261 (47%), Gaps = 2/261 (0%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS LPD+IL +ILS LPTK+A TS+LSKRWK LW V L+F S++
Sbjct: 25 DRISILPDDILVHILSPLPTKQAFVTSVLSKRWKNLWCLVPALEFVGTKTVTV----SKY 192
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSP 122
V VSR + ++ W IY P
Sbjct: 193 V-----------------------VSRQAAGNHYMHNWER--------------IYPEFP 261
Query: 123 LNLSSIFSCKTLVVLKLEGLDLNTTVSS--VDLPFLKVLHLQVLQLSQRRCLAQLLNGCP 180
++IF+ +TLV+LKL L + + + + LP LK LHL+ ++ Q L LL GCP
Sbjct: 262 ---NTIFTRETLVILKLSELFMGSGFCNYLIRLPSLKTLHLKDIEFQQYGDLDCLLEGCP 432
Query: 181 VLEDLKAKAIYFKKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEIYIR 240
VLEDL+ I + + KTL+KL RADI + RM+ ++NV+FL I +
Sbjct: 433 VLEDLQLYDISYVSLLFSNACRKTLTKLNRADIIQCDCGV-RMKALSNVEFLHIK---LS 600
Query: 241 IPHDDQFTDMFHNLTHIELGY 261
+ F+NLTH L Y
Sbjct: 601 KGYLHYVFPAFNNLTHFVLNY 663
>TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein F17J16.10
- Arabidopsis thaliana, partial (8%)
Length = 1019
Score = 95.5 bits (236), Expect(2) = 4e-22
Identities = 88/295 (29%), Positives = 134/295 (44%), Gaps = 28/295 (9%)
Frame = +2
Query: 22 TKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKG--WHSRFVQSVHRVIHT------P 73
TK AV TS+LSKRW LW V LDF + + W ++F +V T
Sbjct: 95 TKDAVTTSVLSKRWTNLWCFVPFLDFSDIKLADHESFLWFNQFFCTVMLFRETYASNSID 274
Query: 74 DFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIY-------------SL 120
F IQ +++ S +N W+ +R +++++ + +L
Sbjct: 275 SFSLDIQYAHLNYFSI-----LRLNNWLYLLGKRN-VKYLNLHLNFLNALLVDVQHRKTL 436
Query: 121 SPLNLSSIFSCKTLVVLKL-----EGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQL 175
+P S+IF+CKTLVVL + +G ++ P LK LH + + L
Sbjct: 437 TPKLPSTIFTCKTLVVLNISWFAFKGFSFSSVGFGFGFPSLKTLHFNHIYFNHLSDFLLL 616
Query: 176 LNGCPVLEDLKAKAIYFKKYKETGFEYKT--LSKLVRADISGTSTDLFRMEVINNVDFLR 233
L GCPVLED KA ++ + +E + T LS L+R DI T+ D+ + N+V FLR
Sbjct: 617 LAGCPVLEDFKACHVFTLEEEEEATQESTLNLSNLIRTDIIDTNFDIPMKALFNSV-FLR 793
Query: 234 IHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVINQ 288
I +D F+NLTH+ + YY ++ L +CP LQ L + Q
Sbjct: 794 IQLCQRYTSYD---FPTFNNLTHLVINYY-----LDMFIKVLYHCPNLQKLELYQ 934
Score = 26.2 bits (56), Expect(2) = 4e-22
Identities = 11/23 (47%), Positives = 18/23 (77%)
Frame = +1
Query: 2 EDRISSLPDEILCYILSFLPTKK 24
EDRIS L D+I+ +ILS++ ++
Sbjct: 34 EDRISKLDDKIIHHILSYVRNQR 102
>BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.300 -
Arabidopsis thaliana, partial (4%)
Length = 628
Score = 89.7 bits (221), Expect = 1e-18
Identities = 69/195 (35%), Positives = 104/195 (52%), Gaps = 6/195 (3%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVAT-SILSKRWKPLWRSVTTLDF---DEANFRRTKGW 58
DR+SSLPD +LC+I+SFLPT +VAT LS RW+ LW+ + F D +F+R
Sbjct: 5 DRLSSLPDSVLCHIMSFLPTITSVATIDRLSSRWRHLWKDLQVFRFSSDDSYSFKR---- 172
Query: 59 HSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIY 118
+ FV +V + + + I+K +F R EV + +WV+AA+ E +D+ IY
Sbjct: 173 FAFFVNAVLALRRS----RHIRKFDFTFEFREYEVEC-IEMWVHAAI-GPRLEELDLDIY 334
Query: 119 SLS-PLNLSSIFSCKTLVVLKLEG-LDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLL 176
L LS SC LV L+L G ++ S V P LK ++L + + L
Sbjct: 335 EADINLPLSFFTSCNNLVSLRLGGEYNMKFKHSLVQFPSLKKVYLSTMDSDS---IVAFL 505
Query: 177 NGCPVLEDLKAKAIY 191
+GCP L+DL+ ++Y
Sbjct: 506 SGCPKLQDLEIFSVY 550
>TC92783 similar to PIR|T49020|T49020 hypothetical protein F3C22.70 -
Arabidopsis thaliana (fragment), partial (7%)
Length = 679
Score = 77.0 bits (188), Expect = 8e-15
Identities = 67/206 (32%), Positives = 107/206 (51%), Gaps = 22/206 (10%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVAT-SILSKRWKPLWRSVTTLDF---------DEANF 52
D I++LPD +LC+ILSFLPTK V T ++S+RW+ LW ++ DF + +F
Sbjct: 94 DWINTLPDSLLCHILSFLPTKITVTTIPLVSRRWRHLWENLQVFDFYLKYNKHTINNNDF 273
Query: 53 RRTKGWHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEV----PSNVNVWVNAAVQRG 108
R+ + FV SV + + D I+K+R+S HSEV ++++ W+ +A
Sbjct: 274 RK----FAAFVNSVLSIRRSRD----IRKMRLS--CGHSEVDPFFSNSIDSWIRSAT-GS 420
Query: 109 GFEHVDICIYSLSPLNLSSIF-----SCKTLVVLKLEGLDLNTTV---SSVDLPFLKVLH 160
+ +DI ++ + SIF C LV L L G D+ + S+ LP LK L
Sbjct: 421 CLQELDITLFYTDREYVFSIFLTILAPCTNLVSLSLCG-DIYVVLERSSAFFLPSLKKLQ 597
Query: 161 LQVLQLSQRRCLAQLLNGCPVLEDLK 186
L + + + + LL+ CP+LE L+
Sbjct: 598 LDIGYV-EMNSMDNLLSSCPILETLE 672
>BG646112 weakly similar to GP|10177368|dbj contains similarity to heat shock
transcription factor HSF30~gene_id:K16L22.13, partial
(5%)
Length = 742
Score = 75.9 bits (185), Expect = 2e-14
Identities = 73/221 (33%), Positives = 106/221 (47%), Gaps = 28/221 (12%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAV-ATSILSKRWKPLWRSVTTLDFDE---ANFRRTKGW 58
DR+SSLPD +LC+ILSFLPTK +V TS++S+RW+ LW + FD+ N R K +
Sbjct: 14 DRMSSLPDSLLCHILSFLPTKTSVMTTSLVSRRWRHLWEHLNVFVFDDKSNCNCRNPKKF 193
Query: 59 H--SRFVQSVHRVIHTPDFLQPIQKLRMSFVSR--HSEVPSNVNVWVNAAVQRGGFE--- 111
+ FV SV + + + I+K ++ + +S V +WV AA+ +
Sbjct: 194 RKFAFFVSSVLSLRKS----RHIRKFHLTCCTSDVYSFPGECVYMWVRAAIGPHLEDLSL 361
Query: 112 -----HVDICIYSLSPLNLSSIFSCKTLVVLKLEGL-DLNTTVSSVDLPFLKVLHLQVLQ 165
H D +Y L P S+ +C LV L L GL L S++ P LK+L ++
Sbjct: 362 NITNCHGDDMVY-LPP----SLLNCTNLVSLSLFGLIHLKFQPSAIHFPSLKMLKVEFSI 526
Query: 166 LSQR-----------RCLAQLLNGCPVLEDLKAKAIYFKKY 195
L + L+GCPVLE L YF Y
Sbjct: 527 LEHNIDIEDFGKHLTDSILVFLSGCPVLETLDT---YFSPY 640
>BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis thaliana},
partial (11%)
Length = 549
Score = 75.9 bits (185), Expect = 2e-14
Identities = 57/159 (35%), Positives = 79/159 (48%), Gaps = 4/159 (2%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRI +LPDEIL +ILSF+PTK+A TSILSKRW LWR V LDF E N
Sbjct: 122 DRIGTLPDEILIHILSFVPTKQAFTTSILSKRWIHLWRYVPILDFTETN----------- 268
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYS--L 120
++ VI +F+ + + SRHS ++N ++ + H + S +
Sbjct: 269 LEDRDSVIRFEEFIFSVIR------SRHSAGNHSINTFILDIQRHSSRLHALVFGNSNTI 430
Query: 121 SPLNLSSIFSCKTLVVLKLEGLDLNTT--VSSVDLPFLK 157
+P I TLV LKL D+ ++ + LP LK
Sbjct: 431 APEFPIPILGSTTLVXLKLASFDMGADLFLNLITLPSLK 547
>TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (8%)
Length = 692
Score = 74.7 bits (182), Expect = 4e-14
Identities = 41/89 (46%), Positives = 56/89 (62%)
Frame = +3
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
EDRIS+LPD I+ +ILSF+PTK A TSILSKRW PLW SV L F++ F+ + + S
Sbjct: 264 EDRISALPDPIIWHILSFVPTKTAAITSILSKRWNPLWLSVLILHFEDETFQNMESF-SH 440
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRH 90
F+ SV + D PI+ ++ R+
Sbjct: 441 FMSSVFLL---RDITLPIRSFHLNRSKRY 518
>TC78781 weakly similar to PIR|T02291|T02291 hypothetical protein T13D8.28 -
Arabidopsis thaliana, partial (12%)
Length = 1300
Score = 73.6 bits (179), Expect = 9e-14
Identities = 59/193 (30%), Positives = 96/193 (49%), Gaps = 2/193 (1%)
Frame = +3
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
EDRISSL D++L ILS L T+++V+T +LSKRW +++++T L D+ N
Sbjct: 126 EDRISSLSDDLLVRILSNLHTRESVSTCVLSKRWVHVFKALTCLRLDDEN--------GS 281
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
+ ++ + H D + +S +R S V + R + IY +
Sbjct: 282 LLDALDEMRHRTD----LTSFDLSIFTRRSGDIVTVAEKAVRNIVRLNLSLNSLSIYGIL 449
Query: 122 PLNLS-SIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCP 180
+ L+ +F+ +TLV LKL + + T+S LP LKVL L +Q + LL+
Sbjct: 450 NMYLTRPVFNSRTLVELKLHRVCIAETLSVASLPSLKVLSLSRVQFDTKSVFLSLLSAVS 629
Query: 181 -VLEDLKAKAIYF 192
VLE+L+ + F
Sbjct: 630 RVLEELRISSPTF 668
>TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ22.3~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 712
Score = 70.1 bits (170), Expect = 9e-13
Identities = 49/141 (34%), Positives = 77/141 (53%), Gaps = 3/141 (2%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEA--NFRRTKGWHS 60
D IS+L + IL ILSF+P AV+TS+LS+RW +W+ +T LDFD++ R+ +
Sbjct: 164 DMISTLHESILSQILSFIPIVDAVSTSVLSRRWVDVWKCITNLDFDDSLLGSRKKRMQKE 343
Query: 61 RFVQSVHRV-IHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYS 119
+FV V +V IH + IQ +S S H S ++ W++ ++R + + I
Sbjct: 344 QFVNFVEKVLIHFTN--SSIQSFSLSLTS-HQYDASKLSEWISFILER-RVQKLHIQYAD 511
Query: 120 LSPLNLSSIFSCKTLVVLKLE 140
L S+F C +LV L L+
Sbjct: 512 KVFLPSDSLFRCNSLVDLTLQ 574
>BG581615 similar to GP|10177368|dbj contains similarity to heat shock
transcription factor HSF30~gene_id:K16L22.13, partial
(3%)
Length = 825
Score = 69.7 bits (169), Expect = 1e-12
Identities = 54/154 (35%), Positives = 83/154 (53%), Gaps = 15/154 (9%)
Frame = +3
Query: 3 DRISSLPDEILCYILSFLPTKKAVAT-SILSKRWKPLWRSVTTLDFD------EANFRRT 55
DR+S LPD ILC+ILSFLPT+ +VAT +++S+RW+ LW+ DFD A++ R
Sbjct: 315 DRLSGLPDSILCHILSFLPTRTSVATMNLVSRRWRNLWKHAQVFDFDFDCDGVSADYERF 494
Query: 56 KGWHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSR---HSEVPSN-VNVWVNAAVQRGGFE 111
+ FV SV + + D IQK ++ S S++ +N V +W+ AA +
Sbjct: 495 R----FFVNSVLALRKSRD----IQKFHLTITSDCQFMSDIQNNYVEMWICAAT-GPHLQ 647
Query: 112 HVDICIYS----LSPLNLSSIFSCKTLVVLKLEG 141
+ + I S + L+ S +C LV L L+G
Sbjct: 648 ELSLIIPSYADQIVKLSPSLFMNCSNLVSLSLDG 749
>TC82256 similar to GP|20042913|gb|AAM08741.1 Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (9%)
Length = 897
Score = 66.6 bits (161), Expect = 1e-11
Identities = 64/188 (34%), Positives = 86/188 (45%), Gaps = 10/188 (5%)
Frame = +3
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
+D IS L D +L +ILSFL +AV T +LSKRW LW+S++T+ + R K R
Sbjct: 216 DDFISDLTDSVLLHILSFLNAIQAVQTCVLSKRWIILWKSLSTITLRSSYSRPRK----R 383
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDI---CIY 118
F + V R+ D I L RHS S + + AV +H+ I C
Sbjct: 384 FDEFVSRIFSLRDGSTAIHTL--DLYRRHSMKHSLLRKIIEYAVSH-NVQHLRIDYTCHI 554
Query: 119 SLSPLNLSSIFSCKTLVVLKLEGLDLNTTV-------SSVDLPFLKVLHLQVLQLSQRRC 171
P S +FSC TL L L G NT V +S++LP L L L+
Sbjct: 555 ENFP---SCLFSCHTLKSLNLSGFLYNTFVHHKPVFRNSLNLPSLTNLSLKYF------A 707
Query: 172 LAQLLNGC 179
A+ NGC
Sbjct: 708 FARSDNGC 731
>BI311381 weakly similar to PIR|T46128|T461 hypothetical protein T2J13.140 -
Arabidopsis thaliana, partial (12%)
Length = 803
Score = 65.9 bits (159), Expect = 2e-11
Identities = 33/46 (71%), Positives = 36/46 (77%)
Frame = +1
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDF 47
ED IS+LPD ILC+ILSFL T A ATS+LSKRW LWRSV TL F
Sbjct: 100 EDIISTLPDAILCHILSFLETNYADATSVLSKRWNHLWRSVHTLRF 237
>TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (5%)
Length = 696
Score = 65.5 bits (158), Expect = 2e-11
Identities = 59/202 (29%), Positives = 99/202 (48%), Gaps = 18/202 (8%)
Frame = +3
Query: 3 DRISSLPDEILCYILSFLPTKKAVATS-ILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
DRI+ LP+ +L +ILSFLPTK V T+ ++S++W+ LW+++ LDF +++ + + +
Sbjct: 105 DRINQLPNSLLHHILSFLPTKTCVQTTPLVSRKWRNLWKNLEALDFCDSSSPQIYEFDND 284
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSR-------HSEVP----SNVNVWVNAAVQRGGF 110
+ V F+ + LR S V R H ++ +++ W+NA +
Sbjct: 285 EQFLIFSV-----FVNTVLTLRKSRVVRKFCLSCYHVQLDPFYNHSIDTWINATI-GPNL 446
Query: 111 EHVDICIYSLSPLNL--SSIFSCKTLVVLKLEG---LDLNTTVSSVDLPFLKVLH-LQVL 164
E + + + + N S+FSC LV L L L S + LP LK+L L +
Sbjct: 447 EEFHLTLLTAAGFNRVPLSLFSCPNLVSLSFNDYIILQLQDN-SKICLPSLKLLQLLDMY 623
Query: 165 QLSQRRCLAQLLNGCPVLEDLK 186
L + C PVLE+L+
Sbjct: 624 NLDLKFCEMPFSLAVPVLENLE 689
>TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity to heat
shock transcription factor HSF30~gene_id:K16L22.13,
partial (6%)
Length = 713
Score = 64.7 bits (156), Expect = 4e-11
Identities = 30/54 (55%), Positives = 39/54 (71%)
Frame = +1
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRT 55
EDRIS+LPD ++ +ILSFLP+K A AT++LSKRWKPLW + F+ F T
Sbjct: 541 EDRISALPDSLVHHILSFLPSKDATATTVLSKRWKPLWLLELIIHFNHQPFPXT 702
>TC82690 weakly similar to GP|9802755|gb|AAF99824.1| Unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 762
Score = 63.9 bits (154), Expect = 7e-11
Identities = 63/188 (33%), Positives = 91/188 (47%), Gaps = 5/188 (2%)
Frame = +1
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRISSLP I+ ILS LP K+AV TSILS +W+ W ++ L FD T
Sbjct: 115 DRISSLPGHIIDQILSILPIKEAVRTSILSTKWRYKWATLPNLVFDSQCISDTSEDLLVI 294
Query: 63 VQSVHRVIHTPDFLQ--PIQKLRMSFVSRHSEV--PSNVNVWVNAAVQRGGFEHVDICIY 118
+ R+I L PI+K ++S H E+ ++++ W +R E V + I+
Sbjct: 295 KSKLSRIIDHVLLLHSGPIKKFKLS----HRELIGVTDIDRWTLHLTRRPVKEFV-LEIW 459
Query: 119 SLSPLNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLN 177
+ S +FSC+ L L+L L + LK L LQ + LSQ L++
Sbjct: 460 KGQRYKIPSCLFSCQGLHHLELFNCWLIPPSTFQGFRNLKSLDLQHVTLSQ-DAFENLIS 636
Query: 178 GCPVLEDL 185
CP+LE L
Sbjct: 637 TCPLLERL 660
>TC82108 similar to GP|21537320|gb|AAM61661.1 unknown {Arabidopsis
thaliana}, partial (7%)
Length = 817
Score = 62.8 bits (151), Expect = 2e-10
Identities = 47/147 (31%), Positives = 76/147 (50%), Gaps = 8/147 (5%)
Frame = +2
Query: 151 VDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKAKAIYFK---KYKETGFEYKTLSK 207
V LP LK+LHL+ + + + +LL+GCP+L++L+A + K + + +LS
Sbjct: 14 VILPSLKILHLESVCFTYYGHIRKLLSGCPILQELEANDLTIKIPCRVFLGSSQPPSLSN 193
Query: 208 LVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTN 267
LVRA+IS ++ + N +R+ Y H +FHNLTH+EL D+ N
Sbjct: 194 LVRANISNICPMHNLLDRLRNAQHMRLLAKYSYELH-----SVFHNLTHLEL-RLDFMHN 355
Query: 268 -----WIEVVEFLNYCPKLQVLVINQV 289
W +++ L P LQ L+I +V
Sbjct: 356 RGMLKWSWLIKLLQNFPLLQTLIIEEV 436
>BF644690 weakly similar to GP|20042923|gb Putative copia-type polyprotein
{Oryza sativa (japonica cultivar-group)}, partial (2%)
Length = 690
Score = 61.6 bits (148), Expect = 3e-10
Identities = 47/176 (26%), Positives = 82/176 (45%), Gaps = 16/176 (9%)
Frame = +1
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
EDR+S LP+ ++ ILSFL TK V T +LS+RWK +W+ + TL D + F + +
Sbjct: 163 EDRLSDLPNSVILCILSFLNTKDGVRTCVLSRRWKDIWKHIPTLVLDSSRFDTVRQFEI- 339
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQ---------RGGFEH 112
F+ I LR + ++ HS ++ + +Q + +
Sbjct: 340 -------------FMSKILTLRDNTIALHSLDFDHIGKMEHQLLQKILDYVYSHKTKLQR 480
Query: 113 VDICIYSLSPLNLSSIFSCKTLVVLKLE-----GLDLNTTV--SSVDLPFLKVLHL 161
+ I +++ + L + + SCK L L+L + T + S++LP L+ L L
Sbjct: 481 LRIFVHNDNGLIMQCVSSCKNLTSLRLSIYPXMSFCVKTILFPKSLNLPALETLDL 648
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,874,453
Number of Sequences: 36976
Number of extensions: 141916
Number of successful extensions: 1268
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 1237
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1242
length of query: 289
length of database: 9,014,727
effective HSP length: 95
effective length of query: 194
effective length of database: 5,502,007
effective search space: 1067389358
effective search space used: 1067389358
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0487.10


