

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0476a.1
(57 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
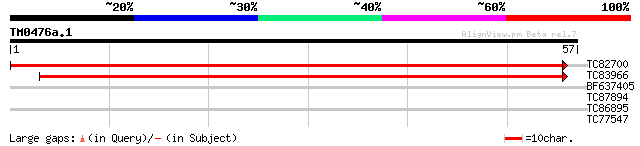
TC82700 similar to GP|23617236|dbj|BAC20903. hypothetical protei... 82 4e-17
TC83966 weakly similar to GP|23617236|dbj|BAC20903. hypothetical... 50 1e-07
BF637405 26 3.6
TC87894 similar to GP|12005328|gb|AAG44394.1 unknown {Hevea bras... 25 8.0
TC86895 similar to PIR|E96688|E96688 unknown protein 33791-3152... 25 8.0
TC77547 similar to PIR|T48354|T48354 hypothetical protein F12E4.... 25 8.0
>TC82700 similar to GP|23617236|dbj|BAC20903. hypothetical protein~similar
to Arabidopsis thaliana chromosome 4 At4g13970, partial
(39%)
Length = 1066
Score = 82.0 bits (201), Expect = 4e-17
Identities = 41/56 (73%), Positives = 48/56 (85%)
Frame = +3
Query: 1 RRRRVIYRSTHELDANDVVSISMWVESHLNCVFFYEDFSNSDPFILGIQTE*QLQR 56
R+ R I RST+ELDA+D VSISMWVES + VFFY+DFS+SDPFILGIQTE QLQ+
Sbjct: 729 RQEREIRRSTYELDADDSVSISMWVESRQSNVFFYQDFSDSDPFILGIQTEWQLQQ 896
>TC83966 weakly similar to GP|23617236|dbj|BAC20903. hypothetical
protein~similar to Arabidopsis thaliana chromosome 4
At4g13970, partial (13%)
Length = 720
Score = 50.4 bits (119), Expect = 1e-07
Identities = 21/53 (39%), Positives = 36/53 (67%)
Frame = +3
Query: 4 RVIYRSTHELDANDVVSISMWVESHLNCVFFYEDFSNSDPFILGIQTE*QLQR 56
R I+ S+ EL N+ S+ +W++ H +F+++D S S+ FI+ IQT+ QLQ+
Sbjct: 201 RTIHNSSRELLGNEECSVKIWIQRHQKDIFYFQDNSGSESFIVAIQTDWQLQQ 359
>BF637405
Length = 666
Score = 25.8 bits (55), Expect = 3.6
Identities = 8/19 (42%), Positives = 15/19 (78%)
Frame = +2
Query: 17 DVVSISMWVESHLNCVFFY 35
++ S+ +W+ESH+ V+FY
Sbjct: 500 EMSSLPLWLESHVALVYFY 556
>TC87894 similar to GP|12005328|gb|AAG44394.1 unknown {Hevea brasiliensis},
partial (26%)
Length = 997
Score = 24.6 bits (52), Expect = 8.0
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = -2
Query: 1 RRRRVIYRSTHELDANDVVSISMWVESHLNCVF 33
+ R + Y+S ++S S+WVES + C+F
Sbjct: 618 KEREISYKSGSSYGQRSILSNSIWVES-ICCLF 523
>TC86895 similar to PIR|E96688|E96688 unknown protein 33791-31527
[imported] - Arabidopsis thaliana, partial (63%)
Length = 1857
Score = 24.6 bits (52), Expect = 8.0
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = -2
Query: 10 THELDANDVVSISMWVESHLNCVFFYEDFSNSDPFILGIQTE*QLQ 55
T+ + D V +S+ ES C F F N FILG + LQ
Sbjct: 302 TNRNQSFDKVDLSLRTESLFRCPFSSLPFENCILFILGDEDPCDLQ 165
>TC77547 similar to PIR|T48354|T48354 hypothetical protein F12E4.60 -
Arabidopsis thaliana, partial (53%)
Length = 2073
Score = 24.6 bits (52), Expect = 8.0
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = +3
Query: 18 VVSISMWVESHLNCVFFYEDFSNSD 42
V+ IS W E H N ++ D ++S+
Sbjct: 1545 VIFISFWAEVHYNSIYPQGDITSSE 1619
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.144 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,085,729
Number of Sequences: 36976
Number of extensions: 23192
Number of successful extensions: 235
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 235
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 235
length of query: 57
length of database: 9,014,727
effective HSP length: 33
effective length of query: 24
effective length of database: 7,794,519
effective search space: 187068456
effective search space used: 187068456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0476a.1


