

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
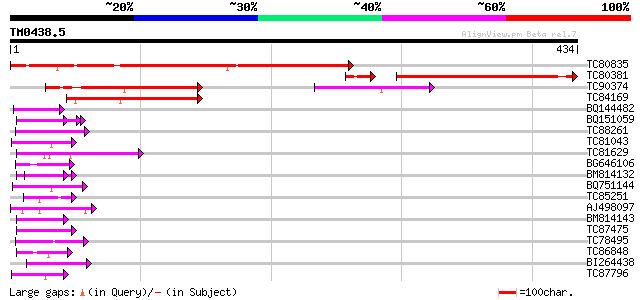
Query= TM0438.5
(434 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF com... 395 e-110
TC80381 similar to PIR|F84747|F84747 probable SWI/SNF complex su... 187 2e-51
TC90374 similar to PIR|T00422|T00422 probable SWI/SNF family tra... 106 2e-23
TC84169 similar to GP|18377706|gb|AAL67003.1 unknown protein {Ar... 93 2e-19
BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 51 1e-06
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 47 2e-05
TC88261 similar to PIR|S22697|S22697 extensin - Volvox carteri (... 45 4e-05
TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 45 5e-05
TC81629 similar to GP|15081654|gb|AAK82482.1 AT4g26750/F10M23_90... 45 7e-05
BG646106 similar to GP|3549654|emb| metal-transporting P-type AT... 44 9e-05
BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 44 9e-05
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 44 2e-04
TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean... 43 3e-04
AJ498097 weakly similar to PIR|T14441|T144 glycine-rich protein ... 42 4e-04
BM814143 GP|21322711|e pherophorin-dz1 protein {Volvox carteri f... 42 4e-04
TC87475 similar to PIR|T11622|T11622 extensin class 1 precursor ... 42 5e-04
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 42 5e-04
TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {A... 42 5e-04
BI264438 similar to GP|21427467|gb actin-related protein 6 {Arab... 42 5e-04
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 42 5e-04
>TC80835 weakly similar to PIR|F84747|F84747 probable SWI/SNF complex
subunit SW13 [imported] - Arabidopsis thaliana, partial
(34%)
Length = 830
Score = 395 bits (1015), Expect = e-110
Identities = 191/272 (70%), Positives = 227/272 (83%), Gaps = 9/272 (3%)
Frame = +2
Query: 1 MATKSPTTATEPPPSSSAGTP-PPPLPPPPLHQPQP---AKPDAPPSEPKLSATPDANVI 56
MAT +P EPP SS+ GTP PP PPP + PQP AKP+ P E K S+ DAN+I
Sbjct: 38 MATSTPNRTVEPP-SSATGTPLPPTTTPPPSNPPQPPPVAKPEVPLPEAKTSS--DANLI 208
Query: 57 LVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKSPRVYKYYRNSIIKYFRYNPNRKITFTDV 116
LVPSHSRWFSWDSIH+CE+R++PE SSK+PRVYKYYRNSI+K+FR+NPNRKITFTDV
Sbjct: 209 LVPSHSRWFSWDSIHECEIRNIPE----SSKNPRVYKYYRNSIVKFFRFNPNRKITFTDV 376
Query: 117 RKMLVGDVGSIRRVFEFLEAWGLINYHPSSSFSKPFKWDDKDTKVDSAE-----PPPPPV 171
RK LVGDVGSIRRVF+FLEAWGLINYHPSSS SKP KW+DKDTK D+A P P PV
Sbjct: 377 RKTLVGDVGSIRRVFDFLEAWGLINYHPSSSLSKPLKWEDKDTKPDAASNSAESPSPAPV 556
Query: 172 RETAKRVCSGCKALCTIACFVCDKYDLTLCARCYVRGNYRVGVSSSDFKRVEISEETKTE 231
+E AKR+CSGCK LC +ACF C+K ++TLCARC++RGNYR+G+S+++FKRVEISEETK E
Sbjct: 557 KE-AKRICSGCKNLCVMACFACEKNNMTLCARCFIRGNYRIGMSNTEFKRVEISEETKNE 733
Query: 232 WTEKETLNLVEAISHYGDDWKRVSHQVVGRTE 263
WTEKETLNL+EAI+++GDDWKRV+HQVVGRT+
Sbjct: 734 WTEKETLNLLEAITNFGDDWKRVAHQVVGRTD 829
>TC80381 similar to PIR|F84747|F84747 probable SWI/SNF complex subunit SW13
[imported] - Arabidopsis thaliana, partial (10%)
Length = 892
Score = 187 bits (475), Expect(2) = 2e-51
Identities = 103/138 (74%), Positives = 118/138 (84%)
Frame = +1
Query: 297 ESETVASAESNKRMRLTPLADASNPIMAQAAFLSALAGPEVAQAAAQAALTTLSDVYKST 356
ESETV S +S+KRM LTPLADASNPIMAQAAFLSALAG EVAQAAAQAALT+LS+VYKST
Sbjct: 160 ESETVPSDKSSKRMCLTPLADASNPIMAQAAFLSALAGTEVAQAAAQAALTSLSNVYKST 339
Query: 357 RINYRSLPKNTLQQDAGVASNGGNNSDSLQGARLNASLQLEKDESDVEKAISEIIEVQVC 416
RINYRS P+NTLQQDA VASNGGN SDS+QG+ L A+LQLEK+ESDVEKAISE+ EVQ+
Sbjct: 340 RINYRSFPRNTLQQDAAVASNGGNASDSIQGSLLRANLQLEKEESDVEKAISEVTEVQMK 519
Query: 417 PILNCLMVFTLVKFKNLD 434
I + L+ F++LD
Sbjct: 520 NIQD-----KLINFEDLD 558
Score = 33.5 bits (75), Expect(2) = 2e-51
Identities = 16/23 (69%), Positives = 19/23 (82%)
Frame = +3
Query: 258 VVGRTEKECVAHFLKLPFGEQFL 280
+VG T K CVA FL+LPFG+QFL
Sbjct: 15 LVGPTRK-CVARFLELPFGDQFL 80
>TC90374 similar to PIR|T00422|T00422 probable SWI/SNF family transcription
activator [imported] - Arabidopsis thaliana, partial
(40%)
Length = 1543
Score = 106 bits (264), Expect = 2e-23
Identities = 54/122 (44%), Positives = 76/122 (62%), Gaps = 2/122 (1%)
Frame = +1
Query: 28 PPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDSSS- 86
PP H D P S+ +L + +PS S+WF+WD IH+ E E+FD +S
Sbjct: 79 PPAHFSMEGSKD-PISDSELE------LYTIPSSSKWFAWDEIHETEKTAFKEYFDGTSI 237
Query: 87 -KSPRVYKYYRNSIIKYFRYNPNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHPS 145
++P++YK YR+ II +R P+R++TFT+VRK LVGDV + +VF FLE WGLINY
Sbjct: 238 TRTPKIYKEYRDFIINKYREEPSRRLTFTEVRKSLVGDVTFLNKVFLFLECWGLINYGAP 417
Query: 146 SS 147
S+
Sbjct: 418 SA 423
Score = 55.8 bits (133), Expect = 3e-08
Identities = 33/123 (26%), Positives = 56/123 (44%), Gaps = 31/123 (25%)
Frame = +3
Query: 234 EKETLNLVEAISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLGSA---------- 283
E ETL L+E++ +GDDW+ V+ V +T+ EC++ ++LPFGE L S
Sbjct: 846 EGETLLLLESVLKHGDDWELVAQSVRTKTKLECISKLIELPFGELMLASVRRNDNSNSVT 1025
Query: 284 ---------------------VSDDGCELKQHADESETVASAESNKRMRLTPLADASNPI 322
D E K +++ + +KR R++ L+D+S+ +
Sbjct: 1026GIVNNRNQVQVSSSDHQETSMTQDQSSEPKNEVEQNGDAVNENPSKRRRVSTLSDSSSSL 1205
Query: 323 MAQ 325
M Q
Sbjct: 1206MKQ 1214
>TC84169 similar to GP|18377706|gb|AAL67003.1 unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 884
Score = 92.8 bits (229), Expect = 2e-19
Identities = 46/108 (42%), Positives = 66/108 (60%), Gaps = 4/108 (3%)
Frame = +1
Query: 44 EPKLSA--TPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFD--SSSKSPRVYKYYRNSI 99
EP+ A + DAN +VP+H WFSW IH E R +P FF+ S +++P Y RN I
Sbjct: 361 EPEFKAIRSRDANAHVVPTHCGWFSWSDIHSIEKRMMPSFFNGISENRTPDKYMEIRNWI 540
Query: 100 IKYFRYNPNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHPSSS 147
+K F NPN +I D+ ++ +GD + + + EFL+ WGLIN+HP S
Sbjct: 541 MKKFHSNPNIQIELKDLSELDIGDSDARQEIMEFLDYWGLINFHPFPS 684
>BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (16%)
Length = 993
Score = 50.8 bits (120), Expect = 1e-06
Identities = 22/39 (56%), Positives = 23/39 (58%)
Frame = +1
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
KSP PPPS S PPPPLPP PL +P P P PP
Sbjct: 169 KSPRAHLFPPPSPSISAPPPPLPPLPLFRPPPPPPPPPP 285
Score = 37.7 bits (86), Expect = 0.009
Identities = 19/53 (35%), Positives = 22/53 (40%), Gaps = 5/53 (9%)
Frame = +3
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPP-----LHQPQPAKPDAPPSEPKLSATP 51
+ P+ + PPP S PPP PP P L P P P PP P P
Sbjct: 216 RPPSPSPSPPPLSPPSPSPPPPPPQPLLGPGLRPPGPRPPAPPPPAPAPRPAP 374
Score = 35.0 bits (79), Expect = 0.056
Identities = 19/44 (43%), Positives = 20/44 (45%), Gaps = 13/44 (29%)
Frame = +3
Query: 12 PPPSSSAGTP--PPPLP-----------PPPLHQPQPAKPDAPP 42
PPP A P PPPLP PPPL P P+ P PP
Sbjct: 156 PPPPKVAPRPPFPPPLPFYLRPPSPSPSPPPLSPPSPSPPPPPP 287
Score = 34.7 bits (78), Expect = 0.074
Identities = 20/53 (37%), Positives = 25/53 (46%), Gaps = 3/53 (5%)
Frame = +2
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAP---PSEPKLSATP 51
A P A + P+ + PPP L PPPL P+ AP P P SA+P
Sbjct: 140 AAPRPAPAPQSRPAPTFSPPPPLLSPPPLPLSLPSPSFAPLPLPPPPPPSASP 298
Score = 33.1 bits (74), Expect = 0.21
Identities = 18/46 (39%), Positives = 21/46 (45%), Gaps = 3/46 (6%)
Frame = +2
Query: 13 PPSSSAGTPPPPLPPP---PLHQPQPAKPDAPPSEPKLSATPDANV 55
P S A P PP PPP P +P P +P AP + P P V
Sbjct: 239 PSPSFAPLPLPPPPPPSASPWPRPSPPRPPAPRAPPPRPGPPPGPV 376
Score = 32.3 bits (72), Expect = 0.37
Identities = 18/60 (30%), Positives = 25/60 (41%), Gaps = 5/60 (8%)
Frame = +3
Query: 1 MATKSPTTATEPP-----PSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
++ SP+ PP P P PP PPPP P+PA P+ +A A +
Sbjct: 249 LSPPSPSPPPPPPQPLLGPGLRPPGPRPPAPPPPAPAPRPAPSRPSVGRPRRAAPSLARI 428
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 46.6 bits (109), Expect = 2e-05
Identities = 19/53 (35%), Positives = 25/53 (46%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILV 58
P PPP PPPP PPPP P P P PP P +P+ ++++
Sbjct: 159 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPGFSPERGIVVL 1
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/50 (40%), Positives = 20/50 (40%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
P PPP PPPP PPPP P P P PP P P V
Sbjct: 178 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPGV 29
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 300 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 181
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 296 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 177
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 291 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 172
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 289 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 170
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 284 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 165
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 260 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 141
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 281 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 162
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 276 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 157
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 274 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 155
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 271 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 152
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 268 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 149
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 265 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 146
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 212 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 93
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 215 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 96
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 257 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 138
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 253 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 134
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 250 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 131
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 246 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 127
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 243 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 124
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 242 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 123
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 237 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 118
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 235 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 116
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 231 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 112
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 229 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 110
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 225 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 106
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 222 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 103
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 220 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 101
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 216 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 97
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 263 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 144
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 287 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 168
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 208 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 89
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 204 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 85
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 201 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 82
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 199 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 80
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 197 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 78
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 194 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 75
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 191 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 72
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 188 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 69
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 183 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 64
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 181 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 62
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 298 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 179
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 305 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 186
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 307 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 188
>TC88261 similar to PIR|S22697|S22697 extensin - Volvox carteri (fragment),
partial (7%)
Length = 1516
Score = 45.4 bits (106), Expect = 4e-05
Identities = 21/57 (36%), Positives = 28/57 (48%)
Frame = +2
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
+P + PPP +P PP PPPP QP P S P +S P ++ + PSH
Sbjct: 350 TPPLPSSPPPPPPPSSPSPPPPPPPPPILQPLPLSLPSSAPPVSLVPQSSGLARPSH 520
Score = 28.9 bits (63), Expect = 4.0
Identities = 17/43 (39%), Positives = 19/43 (43%), Gaps = 2/43 (4%)
Frame = +2
Query: 19 GTPPPPLPPPPLHQPQPAK--PDAPPSEPKLSATPDANVILVP 59
G+ P PL PP P P+ P PPS P P IL P
Sbjct: 314 GSIPLPLDSPPPTPPLPSSPPPPPPPSSPSPPPPPPPPPILQP 442
>TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (19%)
Length = 555
Score = 45.1 bits (105), Expect = 5e-05
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Frame = +1
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLP-PPPL--HQPQPAKPDAPPSEPKLSATP 51
A SP + +PP S G+PP P P PPP+ P P P +PPS P S P
Sbjct: 217 APPSPGSPPKPPAPPSPGSPPAPAPAPPPIPPSPPAPPSPGSPPSPPPASPAP 375
Score = 35.0 bits (79), Expect = 0.056
Identities = 14/31 (45%), Positives = 17/31 (54%)
Frame = +1
Query: 21 PPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPPP PP P P+P P +P S P + P
Sbjct: 205 PPPPAPPSPGSPPKPPAPPSPGSPPAPAPAP 297
Score = 34.3 bits (77), Expect = 0.096
Identities = 18/44 (40%), Positives = 19/44 (42%), Gaps = 1/44 (2%)
Frame = +1
Query: 5 SPTTATEPPPSSSAGTPPPPLPP-PPLHQPQPAKPDAPPSEPKL 47
+P A P P S P P PP PP P P KP P P L
Sbjct: 280 APAPAPPPIPPSPPAPPSPGSPPSPPPASPAPPKPTPPKGSPLL 411
Score = 34.3 bits (77), Expect = 0.096
Identities = 17/44 (38%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Frame = +1
Query: 11 EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAP-PSEPKLSATPDA 53
+PPP + PP PP P P P P AP P+ P + +P A
Sbjct: 202 KPPPPAPPSPGSPPKPPAP---PSPGSPPAPAPAPPPIPPSPPA 324
Score = 33.9 bits (76), Expect = 0.13
Identities = 19/46 (41%), Positives = 23/46 (49%), Gaps = 6/46 (13%)
Frame = +1
Query: 12 PPPSSSAGTPP-PPLPP----PPLHQPQPAK-PDAPPSEPKLSATP 51
PP S G+PP PP PP PP P P P +PP+ P + P
Sbjct: 211 PPAPPSPGSPPKPPAPPSPGSPPAPAPAPPPIPPSPPAPPSPGSPP 348
Score = 33.5 bits (75), Expect = 0.16
Identities = 18/50 (36%), Positives = 20/50 (40%)
Frame = +1
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A P PP S G+PP P P P P+P P P P TP
Sbjct: 286 APAPPPIPPSPPAPPSPGSPPSPPPASPA-PPKPTPPKGSPLLPMPEKTP 432
Score = 31.6 bits (70), Expect = 0.62
Identities = 16/42 (38%), Positives = 20/42 (47%)
Frame = +1
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS 43
A SP + PPP+S P PP P PP P P+ P+
Sbjct: 322 APPSPGSPPSPPPAS----PAPPKPTPPKGSPLLPMPEKTPA 435
Score = 29.3 bits (64), Expect = 3.1
Identities = 18/42 (42%), Positives = 19/42 (44%)
Frame = +1
Query: 19 GTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
G P PP PP P P KP APPS A A + PS
Sbjct: 193 G*PKPP-PPAPPSPGSPPKPPAPPSPGSPPAPAPAPPPIPPS 315
>TC81629 similar to GP|15081654|gb|AAK82482.1 AT4g26750/F10M23_90
{Arabidopsis thaliana}, partial (19%)
Length = 824
Score = 44.7 bits (104), Expect = 7e-05
Identities = 35/107 (32%), Positives = 47/107 (43%), Gaps = 10/107 (9%)
Frame = +2
Query: 6 PTTATEPPPSSSAGTPPPPL----PPPP---LHQPQPAKPDAPPSEP--KLSATPDANVI 56
P + PPP S PPPP PPPP HQP PA+ D+ SEP +PD +
Sbjct: 74 PPSQDYPPPPSQDYHPPPPSQDYQPPPPSQDYHQPPPARSDSSYSEPYNHQQYSPDQSQN 253
Query: 57 LVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKS-PRVYKYYRNSIIKY 102
L P++ + + P F +SS S P + YY+ Y
Sbjct: 254 LGPNYPSHETPPPYSLPHFQSYPSFTESSLPSVPVNHTYYQGPDASY 394
>BG646106 similar to GP|3549654|emb| metal-transporting P-type ATPase
(fragment) {Arabidopsis thaliana}, partial (10%)
Length = 687
Score = 44.3 bits (103), Expect = 9e-05
Identities = 25/46 (54%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Frame = -3
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS-EPKLSA 49
SPT T P P S PP PLPPPP P PA P PP+ EPK +A
Sbjct: 397 SPTNFTLPSPDS----PPEPLPPPPPPFPTPAPPFPPPAEEPKEAA 272
>BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (7%)
Length = 164
Score = 44.3 bits (103), Expect = 9e-05
Identities = 20/46 (43%), Positives = 20/46 (43%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPP PPPP PPPP P P P PP P A P
Sbjct: 141 PPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPP 4
Score = 44.3 bits (103), Expect = 9e-05
Identities = 19/40 (47%), Positives = 19/40 (47%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P PA P PP P
Sbjct: 154 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPPPPPPPP 35
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/46 (41%), Positives = 20/46 (43%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPP PPPP PPPP P P P PP P + P
Sbjct: 138 PPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPP 1
Score = 42.7 bits (99), Expect = 3e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 156 PPPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPP 37
Score = 42.4 bits (98), Expect = 4e-04
Identities = 17/34 (50%), Positives = 18/34 (52%)
Frame = -3
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
PPP PPPP PPPP +P P P PP P
Sbjct: 159 PPPPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPP 58
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 148 PPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPP 29
Score = 42.0 bits (97), Expect = 5e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 163 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPPPPP 44
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/40 (47%), Positives = 19/40 (47%)
Frame = -1
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP PPPP PPPP P PA P PP P P
Sbjct: 164 PPPPPPPPPPPPPPPPPP--PPPPAPPPPPPPPPPRPPPP 51
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/48 (39%), Positives = 19/48 (39%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDA 53
P PPP PPP PPPP P P P PP P P A
Sbjct: 153 PPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPPPA 10
Score = 39.3 bits (90), Expect = 0.003
Identities = 18/46 (39%), Positives = 18/46 (39%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPP PPP PPPP P P P PP P P
Sbjct: 144 PPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPPPAP 7
Score = 38.9 bits (89), Expect = 0.004
Identities = 17/40 (42%), Positives = 17/40 (42%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPP P P P PP P
Sbjct: 159 PPPPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPPPPP 40
Score = 38.9 bits (89), Expect = 0.004
Identities = 18/41 (43%), Positives = 19/41 (45%), Gaps = 1/41 (2%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLP-PPPLHQPQPAKPDAPPSEP 45
P PPP PPPP P PPP P P +P PP P
Sbjct: 161 PPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPPPPPP 39
Score = 37.4 bits (85), Expect = 0.011
Identities = 16/33 (48%), Positives = 16/33 (48%)
Frame = -2
Query: 21 PPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDA 53
PPPP PPPP P P P PP P P A
Sbjct: 160 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPA 62
Score = 35.0 bits (79), Expect = 0.056
Identities = 16/35 (45%), Positives = 17/35 (47%)
Frame = -3
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP 38
+ P PPP PPPP PPPP P PA P
Sbjct: 99 RPPPPPPPPPPPPPPPPPPPPPPPPP---PPPAPP 4
Score = 28.5 bits (62), Expect = 5.3
Identities = 13/32 (40%), Positives = 13/32 (40%)
Frame = -1
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQP 33
A P P P PPPP PPPP P
Sbjct: 98 APPPPPPPPPPRPPPPPPPPPPPPPPPPPPPP 3
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 43.5 bits (101), Expect = 2e-04
Identities = 23/66 (34%), Positives = 30/66 (44%), Gaps = 9/66 (13%)
Frame = +1
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPL---------HQPQPAKPDAPPSEPKLSATPDA 53
T+ + + PPPS S PPPP P PL +P P P +PP P S +
Sbjct: 295 TRPSSLFSPPPPSPSRSRPPPPSTPSPLLPLPLLPLATEPSPFPPSSPPPRPASSPSLSP 474
Query: 54 NVILVP 59
V+L P
Sbjct: 475 PVLLFP 492
Score = 38.5 bits (88), Expect = 0.005
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Frame = +1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD------APPSEPKLSATPDAN 54
P ++ P P+SS PP L PP P P P APPS P L A+P ++
Sbjct: 424 PPSSPPPRPASSPSLSPPVLLFPPSASPPPWSPSRPPSALAPPSRPSLPASPPSS 588
Score = 33.5 bits (75), Expect = 0.16
Identities = 19/46 (41%), Positives = 25/46 (54%)
Frame = +1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P +A+ PP S S PP L PP +P+ P +PPS P+ S P
Sbjct: 490 PPSASPPPWSPSR--PPSALAPPS----RPSLPASPPSSPRSSTRP 609
Score = 31.6 bits (70), Expect = 0.62
Identities = 17/56 (30%), Positives = 23/56 (40%)
Frame = +1
Query: 4 KSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
+SPT P S A +P P P L P P P P + +P + L+P
Sbjct: 232 RSPTAPPAAPLSPPASSPSSPTRPSSLFSPPPPSPSRSRPPPPSTPSPLLPLPLLP 399
>TC85251 homologue to PIR|JQ2128|JQ2128 metallothionein - soybean, partial
(93%)
Length = 1554
Score = 42.7 bits (99), Expect = 3e-04
Identities = 21/43 (48%), Positives = 25/43 (57%), Gaps = 2/43 (4%)
Frame = -2
Query: 11 EPPPSSSAGTP--PPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+PP SS TP PPPLPPPP P P + PP+ + ATP
Sbjct: 653 QPPKSSPVSTPTLPPPLPPPPKISPTPVQ--TPPAPAPVKATP 531
Score = 34.3 bits (77), Expect = 0.096
Identities = 19/53 (35%), Positives = 24/53 (44%), Gaps = 2/53 (3%)
Frame = -2
Query: 1 MATKSPTTATEPPPSSSAGTP--PPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+A KS + P + A +P PPP PP P P P PK+S TP
Sbjct: 731 VAPKSAPVTSPAPKIAPASSPKVPPPQPPKSSPVSTPTLPPPLPPPPKISPTP 573
Score = 32.0 bits (71), Expect = 0.48
Identities = 14/38 (36%), Positives = 16/38 (41%)
Frame = -2
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
SPT PP + P P P P P PA +PP
Sbjct: 584 SPTPVQTPPAPAPVKATPVPAPAPAKQAPTPAPATSPP 471
Score = 31.6 bits (70), Expect = 0.62
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = -2
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
+P A + P + A +PP P P P + P PA + P
Sbjct: 524 APAPAKQAPTPAPATSPPIPAPTPAIEAPVPAPESSKP 411
Score = 28.9 bits (63), Expect = 4.0
Identities = 21/58 (36%), Positives = 26/58 (44%), Gaps = 9/58 (15%)
Frame = -2
Query: 12 PPPSSSAGTP--PPPLPPP----PLHQPQPAK---PDAPPSEPKLSATPDANVILVPS 60
PPP + TP PP P P P+ P PAK AP + P + A A VP+
Sbjct: 602 PPPPKISPTPVQTPPAPAPVKATPVPAPAPAKQAPTPAPATSPPIPAPTPAIEAPVPA 429
>AJ498097 weakly similar to PIR|T14441|T144 glycine-rich protein - wild
cabbage (fragment), partial (25%)
Length = 592
Score = 42.4 bits (98), Expect = 4e-04
Identities = 27/78 (34%), Positives = 37/78 (46%), Gaps = 12/78 (15%)
Frame = -2
Query: 1 MATKSPTT------ATEPPPSSSAGTP--PPPLPPPPLHQPQPAKPDAPPSEPKLSATPD 52
+AT PTT AT PP +A P P P+P PP+ P P +P PP P D
Sbjct: 423 LATAPPTTPPATPPATAPPIIPAAAPPDLPLPIPLPPIPLPWPPEPPCPPPPPPFLIVTD 244
Query: 53 ANVI----LVPSHSRWFS 66
++ + L + S WF+
Sbjct: 243 SSCLCAF*LSTNPSLWFN 190
>BM814143 GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (4%)
Length = 132
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 129 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 10
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 125 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 6
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 121 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 2
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 120 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 1
Score = 42.4 bits (98), Expect = 4e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PPPP PPPP P P P PP P
Sbjct: 131 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 12
>TC87475 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (16%)
Length = 396
Score = 42.0 bits (97), Expect = 5e-04
Identities = 19/47 (40%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPP-PLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + + PPP PPP P PPPP P P PP P L ++P
Sbjct: 93 PPSPSPPPPYGYKSPPPPSPSPPPPYIYKSPPSPSPPPYHPYLYSSP 233
Score = 38.5 bits (88), Expect = 0.005
Identities = 18/47 (38%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPP-PLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P +A+ PPP PPP P PPPP P P P P + +P
Sbjct: 45 PPSASPPPPYYYKSPPPPSPSPPPPYGYKSPPPPSPSPPPPYIYKSP 185
Score = 33.5 bits (75), Expect = 0.16
Identities = 15/36 (41%), Positives = 17/36 (46%), Gaps = 2/36 (5%)
Frame = +3
Query: 12 PPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEP 45
PPP +PPPP PPPP + P P P P
Sbjct: 12 PPPPYVYKSPPPPSASPPPPYYYKSPPPPSPSPPPP 119
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 42.0 bits (97), Expect = 5e-04
Identities = 24/56 (42%), Positives = 26/56 (45%)
Frame = +3
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
SP AT PP + TPPP L P PL P PA AP S P +L PS
Sbjct: 531 SPPPATPPPATPPPATPPPALTPTPLSSP-PATTPAPAPAKLKSKAPALAPVLSPS 695
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/46 (43%), Positives = 24/46 (51%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP+S PPP PPP P PA P PP+ P + TP
Sbjct: 471 PPVQSTPPPASPPPASPPPFSPPPA-TPPPATP--PPATPPPALTP 599
Score = 39.7 bits (91), Expect = 0.002
Identities = 20/49 (40%), Positives = 23/49 (46%)
Frame = +3
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T P +T PP S +PPP PPP P P PP P L+ TP
Sbjct: 465 TPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPATPP--PALTPTP 605
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/51 (45%), Positives = 25/51 (48%), Gaps = 3/51 (5%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPPPLPPP---PLHQPQPAKPDAPPSEPKLSATPDA 53
P T PPP S TPPP PPP P P PA P PP+ P + P A
Sbjct: 450 PPTPLTPPPVQS--TPPPASPPPASPPPFSPPPATP--PPATPPPATPPPA 590
Score = 37.7 bits (86), Expect = 0.009
Identities = 20/53 (37%), Positives = 24/53 (44%), Gaps = 1/53 (1%)
Frame = +3
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQ-PQPAKPDAPPSEPKLSATPDA 53
A +P PP PP PL PPP+ P PA P PP+ P + P A
Sbjct: 393 AQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASP--PPASPPPFSPPPA 545
Score = 35.8 bits (81), Expect = 0.033
Identities = 21/62 (33%), Positives = 26/62 (41%), Gaps = 4/62 (6%)
Frame = +3
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLH-QPQPAKPDAPP---SEPKLSATPDANVIL 57
A PTT PP + PP PPPL P PA+ PP S P + +P +
Sbjct: 288 ANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSSPPPVQQSPPPTPLT 467
Query: 58 VP 59
P
Sbjct: 468 PP 473
Score = 35.0 bits (79), Expect = 0.056
Identities = 21/50 (42%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPP-PLPPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
P + PPP+ S TPPP PPP+ Q P P PP P S P A+
Sbjct: 366 PPLQSSPPPAQS--TPPPVQSSPPPVQQSPPPTPLTPP--PVQSTPPPAS 503
Score = 33.5 bits (75), Expect = 0.16
Identities = 18/47 (38%), Positives = 20/47 (42%)
Frame = +3
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP A+ PPP S PPP PPP P P S P + P
Sbjct: 501 SPPPAS-PPPFSPPPATPPPATPPPATPPPALTPTPLSSPPATTPAP 638
Score = 33.5 bits (75), Expect = 0.16
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 5/51 (9%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPP--PLPPPPLHQPQPAKPDAPPSE---PKLSATP 51
PT PP++ +PPP PPP P P + PP++ P + ++P
Sbjct: 276 PTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSSP 428
Score = 33.5 bits (75), Expect = 0.16
Identities = 20/59 (33%), Positives = 29/59 (48%), Gaps = 9/59 (15%)
Frame = +3
Query: 2 ATKSPTTA---TEPPPSSSAGTPP--PPLPPPPLHQPQPAKPDAPP----SEPKLSATP 51
A ++P+T+ + PP+ A TPP P PPP+ P +PP S P +TP
Sbjct: 231 AQQAPSTSPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTP 407
Score = 32.0 bits (71), Expect = 0.48
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 5/51 (9%)
Frame = +3
Query: 6 PTTATEPPPSSSAGTPPPP--LPPPPLHQPQPAKPDAPP---SEPKLSATP 51
P + PPP S+ PPP PPP P P + PP + P + +TP
Sbjct: 345 PPVQSSPPPLQSS--PPPAQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTP 491
Score = 30.8 bits (68), Expect = 1.1
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 2/49 (4%)
Frame = +3
Query: 5 SPTTATEPPPSSSAGTPPPPLPP--PPLHQPQPAKPDAPPSEPKLSATP 51
S A + P +S +P PP PP P PQ + P S P + ++P
Sbjct: 219 SSVGAQQAPSTSPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSP 365
>TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {Arabidopsis
thaliana}, partial (92%)
Length = 1250
Score = 42.0 bits (97), Expect = 5e-04
Identities = 23/49 (46%), Positives = 23/49 (46%), Gaps = 6/49 (12%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPPP------PLHQPQPAKPDAPPSEPKLS 48
P PPP PPPPLPPP PL P P KP PP P LS
Sbjct: 253 PLPPPRPPPRPPL--PPPPLPPPLALNGVPLSPPLPPKPPLPPPLPPLS 113
Score = 33.1 bits (74), Expect = 0.21
Identities = 19/36 (52%), Positives = 21/36 (57%), Gaps = 2/36 (5%)
Frame = -2
Query: 12 PPPSSSAGTP-PPPLPP-PPLHQPQPAKPDAPPSEP 45
PPP + G P PPLPP PPL P P P +PP P
Sbjct: 199 PPPLALNGVPLSPPLPPKPPL--PPPLPPLSPPLPP 98
Score = 30.0 bits (66), Expect = 1.8
Identities = 17/39 (43%), Positives = 17/39 (43%), Gaps = 2/39 (5%)
Frame = -2
Query: 6 PTTATEPPPSSSAGTPPPPLPP--PPLHQPQPAKPDAPP 42
P A P S P PPLPP PPL P P PP
Sbjct: 196 PPLALNGVPLSPPLPPKPPLPPPLPPLSPPLPPLNPPPP 80
>BI264438 similar to GP|21427467|gb actin-related protein 6 {Arabidopsis
thaliana}, partial (43%)
Length = 695
Score = 42.0 bits (97), Expect = 5e-04
Identities = 21/50 (42%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Frame = +1
Query: 14 PSSSAGTP-PPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P+ + TP PPP PPPP ++ P P A PS +TP +N I P+ S
Sbjct: 4 PAQTLSTPLPPPPPPPPPNKTSPPPPSAAPSIAAT*STPTSNAISGPTFS 153
Score = 28.1 bits (61), Expect = 6.9
Identities = 16/58 (27%), Positives = 22/58 (37%)
Frame = +1
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
+P PPP + +PPPP P + S P S A+ +L P S
Sbjct: 22 TPLPPPPPPPPPNKTSPPPPSAAPSIAAT*STPTSNAISGPTFSPLSSASTLLNPLSS 195
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 42.0 bits (97), Expect = 5e-04
Identities = 21/47 (44%), Positives = 21/47 (44%), Gaps = 3/47 (6%)
Frame = -1
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLP---PPPLHQPQPAKPDAPPSEP 45
A P EPPP PPP P PPPL P P P APP P
Sbjct: 188 APPPPPPPPEPPPPPPPPGDPPPYPPPVPPPLPYPTPCAPPAPPPPP 48
Score = 38.5 bits (88), Expect = 0.005
Identities = 18/40 (45%), Positives = 20/40 (50%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P+ PPP + PPPP PPPP P P P PP P
Sbjct: 230 PSPPPIPPPYAPPCAPPPP-PPPPEPPPPPPPPGDPPPYP 114
Score = 36.6 bits (83), Expect = 0.019
Identities = 18/42 (42%), Positives = 20/42 (46%), Gaps = 2/42 (4%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPP--PPLPPPPLHQPQPAKPDAPPSEP 45
P PPP PP PP PPPP P+P P PP +P
Sbjct: 242 PWPPPSPPPIPPPYAPPCAPPPPPPP---PEPPPPPPPPGDP 126
Score = 36.2 bits (82), Expect = 0.025
Identities = 16/35 (45%), Positives = 17/35 (47%)
Frame = -1
Query: 13 PPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKL 47
PP + PPPP PPPP P P PP P L
Sbjct: 197 PPCAPPPPPPPPEPPPPPPPPGDPPPYPPPVPPPL 93
Score = 34.3 bits (77), Expect = 0.096
Identities = 18/43 (41%), Positives = 19/43 (43%), Gaps = 3/43 (6%)
Frame = -1
Query: 12 PPPSSSAGTPPPPLPPP---PLHQPQPAKPDAPPSEPKLSATP 51
PPPS PPP+PPP P P P P PP P P
Sbjct: 236 PPPS------PPPIPPPYAPPCAPPPPPPPPEPPPPPPPPGDP 126
Score = 32.3 bits (72), Expect = 0.37
Identities = 18/43 (41%), Positives = 19/43 (43%), Gaps = 2/43 (4%)
Frame = -1
Query: 5 SPTTATEPPPSSSAGTPPPP--LPPPPLHQPQPAKPDAPPSEP 45
+P A PPP PPP PPPP P P P PP P
Sbjct: 200 APPCAPPPPP------PPPEPPPPPPPPGDPPPYPPPVPPPLP 90
Score = 32.3 bits (72), Expect = 0.37
Identities = 15/41 (36%), Positives = 18/41 (43%)
Frame = -1
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
+P + P P S PPP PP P P P+ PP P
Sbjct: 263 NPQGSYPPWPPPSPPPIPPPYAPPCAPPPPPPPPEPPPPPP 141
Score = 31.2 bits (69), Expect = 0.82
Identities = 19/53 (35%), Positives = 19/53 (35%), Gaps = 13/53 (24%)
Frame = -1
Query: 6 PTTATEPPPSSSAGTPPP--PLPPPPLHQPQPAK-----------PDAPPSEP 45
P PPP G PPP P PPPL P P P PP P
Sbjct: 173 PPPPEPPPPPPPPGDPPPYPPPVPPPLPYPTPCAPPAPPPPPPPLPPPPPYPP 15
Score = 29.3 bits (64), Expect = 3.1
Identities = 16/38 (42%), Positives = 16/38 (42%), Gaps = 1/38 (2%)
Frame = -1
Query: 9 ATEPPPSSSAGTPPPPLP-PPPLHQPQPAKPDAPPSEP 45
A P S PP P P PPP P P PP EP
Sbjct: 269 ALNPQGSYPPWPPPSPPPIPPPYAPPCAPPPPPPPPEP 156
Score = 29.3 bits (64), Expect = 3.1
Identities = 11/24 (45%), Positives = 12/24 (49%)
Frame = -1
Query: 5 SPTTATEPPPSSSAGTPPPPLPPP 28
+P PPP PPPP PPP
Sbjct: 83 TPCAPPAPPPPPPPLPPPPPYPPP 12
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.131 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,672,346
Number of Sequences: 36976
Number of extensions: 330411
Number of successful extensions: 15871
Number of sequences better than 10.0: 758
Number of HSP's better than 10.0 without gapping: 4780
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10022
length of query: 434
length of database: 9,014,727
effective HSP length: 99
effective length of query: 335
effective length of database: 5,354,103
effective search space: 1793624505
effective search space used: 1793624505
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0438.5


