

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
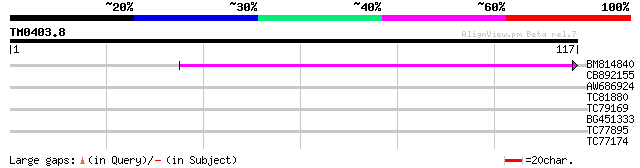
Query= TM0403.8
(117 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 60 1e-10
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 35 0.003
AW686924 similar to GP|22136670|gb| putative serine/threonine ki... 26 2.0
TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [... 25 3.3
TC79169 type IIB calcium ATPase [Medicago truncatula] 25 4.4
BG451333 similar to GP|16604434|gb| AT5g10910/T30N20_180 {Arabid... 25 5.7
TC77895 similar to GP|18087577|gb|AAL58919.1 AT4g00150/F6N15_20 ... 24 7.4
TC77174 similar to PIR|H84453|H84453 probable heat shock protein... 24 9.7
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 60.1 bits (144), Expect = 1e-10
Identities = 33/82 (40%), Positives = 48/82 (58%)
Frame = +3
Query: 36 SSWFV**NPMRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVT 95
S W + + + L + +I A V G G+ +YIPR++L S + + F R QF +
Sbjct: 12 SIWSLHGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLV 191
Query: 96 LCFAMTINNNQGQSLSHVSLYL 117
L FAMTIN +QGQ+L+ V LYL
Sbjct: 192 LSFAMTINKSQGQTLTSVGLYL 257
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 35.4 bits (80), Expect = 0.003
Identities = 15/24 (62%), Positives = 19/24 (78%)
Frame = -2
Query: 94 VTLCFAMTINNNQGQSLSHVSLYL 117
V + FAMTIN +QGQSL H+ +YL
Sbjct: 370 VEVYFAMTINKSQGQSLKHIGVYL 299
>AW686924 similar to GP|22136670|gb| putative serine/threonine kinase
{Arabidopsis thaliana}, partial (4%)
Length = 649
Score = 26.2 bits (56), Expect = 2.0
Identities = 14/41 (34%), Positives = 22/41 (53%), Gaps = 7/41 (17%)
Frame = -2
Query: 83 LPFKFSRRQFLVTLCFAMT-------INNNQGQSLSHVSLY 116
L FKF RQF++ CF M+ ++N G ++ H+ Y
Sbjct: 231 LLFKFLARQFVLLFCFHMSNFLIFSILDNLCG*AIEHIKNY 109
>TC81880 similar to PIR|B85431|B85431 trichohyalin like protein [imported] -
Arabidopsis thaliana, partial (4%)
Length = 1043
Score = 25.4 bits (54), Expect = 3.3
Identities = 13/42 (30%), Positives = 23/42 (53%)
Frame = -3
Query: 74 ISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHVSL 115
+SL+ S+S + + S L CF+ +++ G L+H SL
Sbjct: 522 LSLSLSNSSIFLRRSFSNSLSIFCFSSSLSTEAGVLLTHFSL 397
>TC79169 type IIB calcium ATPase [Medicago truncatula]
Length = 3565
Score = 25.0 bits (53), Expect = 4.4
Identities = 14/35 (40%), Positives = 17/35 (48%)
Frame = +2
Query: 68 TAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTI 102
T Y IS+ S LPF+F L LCF M +
Sbjct: 14 TRYKYNISILSIQQTLPFRFCALITLN*LCFVMVL 118
>BG451333 similar to GP|16604434|gb| AT5g10910/T30N20_180 {Arabidopsis
thaliana}, partial (14%)
Length = 536
Score = 24.6 bits (52), Expect = 5.7
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = -1
Query: 70 YIPRISLTSSDSGLPFKFSRRQFLVTL 96
++ RI++T S LPF+FSR + V +
Sbjct: 110 WMKRITVTIS*KXLPFRFSRPKLAVVV 30
>TC77895 similar to GP|18087577|gb|AAL58919.1 AT4g00150/F6N15_20
{Arabidopsis thaliana}, partial (41%)
Length = 2591
Score = 24.3 bits (51), Expect = 7.4
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 5/38 (13%)
Frame = +1
Query: 74 ISLTSSDSGLPFK-----FSRRQFLVTLCFAMTINNNQ 106
ISL ++ + FK + QFL+ +CF ++INNNQ
Sbjct: 826 ISLLVAEG*IKFKTLISVLHKFQFLMVVCF-ISINNNQ 936
>TC77174 similar to PIR|H84453|H84453 probable heat shock protein [imported]
- Arabidopsis thaliana, partial (90%)
Length = 2727
Score = 23.9 bits (50), Expect = 9.7
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = -1
Query: 79 SDSGLPFKFSRRQFLVTLCFAMTINNNQGQS 109
S + + KFS + FLVT+ T+NN QS
Sbjct: 474 SITSITHKFSEKNFLVTI---QTVNNQIHQS 391
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.353 0.153 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,385,703
Number of Sequences: 36976
Number of extensions: 37867
Number of successful extensions: 418
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 418
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 418
length of query: 117
length of database: 9,014,727
effective HSP length: 93
effective length of query: 24
effective length of database: 5,575,959
effective search space: 133823016
effective search space used: 133823016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0403.8


