

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.14
(76 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
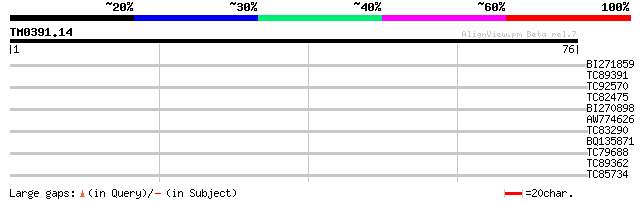
Score E
Sequences producing significant alignments: (bits) Value
BI271859 similar to GP|15810549|gb putative caffeoyl-CoA O-methy... 27 1.1
TC89391 weakly similar to GP|5679843|emb|CAB51836.1 Putitive Ser... 27 1.1
TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane... 27 1.5
TC82475 similar to GP|20146289|dbj|BAB89071. peptide deformylase... 27 1.9
BI270898 25 4.2
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 25 4.2
TC83290 similar to GP|19920013|gb|AAM08453.1 hypothetical protei... 25 4.2
BQ135871 25 5.5
TC79688 similar to GP|21450872|gb|AAK44106.2 unknown protein {Ar... 25 7.2
TC89362 similar to PIR|T05723|T05723 peroxidase (EC 1.11.1.7) pr... 25 7.2
TC85734 homologue to GP|9187460|emb|CAB96990.1 putative 14-kDa p... 24 9.5
>BI271859 similar to GP|15810549|gb putative caffeoyl-CoA O-methyltransferase
{Arabidopsis thaliana}, partial (69%)
Length = 662
Score = 27.3 bits (59), Expect = 1.1
Identities = 9/24 (37%), Positives = 15/24 (62%)
Frame = -2
Query: 31 QWHAFIKNGNTVYVGLLWIPAVLI 54
QWH+ I + + V VG W P++ +
Sbjct: 133 QWHSLILSDSLVLVGKRWFPSIYL 62
>TC89391 weakly similar to GP|5679843|emb|CAB51836.1 Putitive Ser/Thr
protein kinase {Oryza sativa}, partial (54%)
Length = 803
Score = 27.3 bits (59), Expect = 1.1
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = -3
Query: 19 SRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLM 57
S+ VLEL Y + KN TVY+ L W+ L++++
Sbjct: 801 SQFVLELLFTTYYFK--YKNSKTVYISLSWLTVHLMHIL 691
>TC92570 similar to GP|15290605|gb|AAK94907.1 seven transmembrane protein
MLO2 {Oryza sativa} [Oryza sativa (indica
cultivar-group)], partial (47%)
Length = 944
Score = 26.9 bits (58), Expect = 1.5
Identities = 14/46 (30%), Positives = 24/46 (51%), Gaps = 7/46 (15%)
Frame = +3
Query: 27 DVEYQWHAFIKNGNT----VYVGL---LWIPAVLIYLMDIQIWYAI 65
D ++ +H +IK V VG+ LW A++ LM++ WY +
Sbjct: 708 DSKFDFHKYIKRSMEDDFKVVVGISVPLWAFAIIFLLMNVYNWYTL 845
>TC82475 similar to GP|20146289|dbj|BAB89071. peptide deformylase-like
protein {Oryza sativa (japonica cultivar-group)},
partial (52%)
Length = 654
Score = 26.6 bits (57), Expect = 1.9
Identities = 8/20 (40%), Positives = 16/20 (80%)
Frame = +1
Query: 56 LMDIQIWYAIYSSLKMKLWI 75
LMD+ +W+ + +LK+++WI
Sbjct: 595 LMDLXLWWNVTLTLKLQVWI 654
>BI270898
Length = 653
Score = 25.4 bits (54), Expect = 4.2
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +2
Query: 1 IIFFSFSYFLQIQPIIASSR 20
+I SFSYFLQ+ I A+S+
Sbjct: 29 VILLSFSYFLQVFAIPATSK 88
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 25.4 bits (54), Expect = 4.2
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = +1
Query: 39 GNTVYVGLLWIPAVLIYLMDIQ 60
G +Y GLLW+P +L +M I+
Sbjct: 76 GKIMYCGLLWLPVMLKTVMVIR 141
>TC83290 similar to GP|19920013|gb|AAM08453.1 hypothetical protein
{Dictyostelium discoideum}, partial (3%)
Length = 837
Score = 25.4 bits (54), Expect = 4.2
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = +1
Query: 1 IIFFSFSYFLQIQPIIASS 19
I+F F++FL+I+P I+SS
Sbjct: 538 ILFVFFTFFLRIEPSISSS 594
>BQ135871
Length = 789
Score = 25.0 bits (53), Expect = 5.5
Identities = 7/27 (25%), Positives = 16/27 (58%)
Frame = +3
Query: 48 WIPAVLIYLMDIQIWYAIYSSLKMKLW 74
W P YL+ +++ Y++Y + + +W
Sbjct: 222 WNPRSYNYLLHVELCYSLYKTTYVTIW 302
>TC79688 similar to GP|21450872|gb|AAK44106.2 unknown protein {Arabidopsis
thaliana}, partial (57%)
Length = 1334
Score = 24.6 bits (52), Expect = 7.2
Identities = 8/19 (42%), Positives = 14/19 (73%)
Frame = -1
Query: 31 QWHAFIKNGNTVYVGLLWI 49
+W AF +GN V +G++W+
Sbjct: 191 EW*AFSIHGNYVEIGVIWL 135
>TC89362 similar to PIR|T05723|T05723 peroxidase (EC 1.11.1.7) precursor
seed coat - soybean, partial (82%)
Length = 1030
Score = 24.6 bits (52), Expect = 7.2
Identities = 10/30 (33%), Positives = 16/30 (53%)
Frame = +1
Query: 4 FSFSYFLQIQPIIASSRVVLELKDVEYQWH 33
+ FSY L+ I SS + +++QWH
Sbjct: 427 YRFSYTLRCSYIWKSSLQCIHQPSIQFQWH 516
>TC85734 homologue to GP|9187460|emb|CAB96990.1 putative 14-kDa proline-rich
protein {Cicer arietinum}, partial (62%)
Length = 848
Score = 24.3 bits (51), Expect = 9.5
Identities = 17/44 (38%), Positives = 23/44 (51%)
Frame = -3
Query: 33 HAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYSSLKMKLWIF 76
H I N N +V + + IY+ I I+ IY + MKLWIF
Sbjct: 540 HTRITNTNQ-HVNMSTNINIYIYIY-IYIYIYIYIYIYMKLWIF 415
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,082,273
Number of Sequences: 36976
Number of extensions: 38997
Number of successful extensions: 428
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 427
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 428
length of query: 76
length of database: 9,014,727
effective HSP length: 52
effective length of query: 24
effective length of database: 7,091,975
effective search space: 170207400
effective search space used: 170207400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0391.14


