

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.11
(490 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
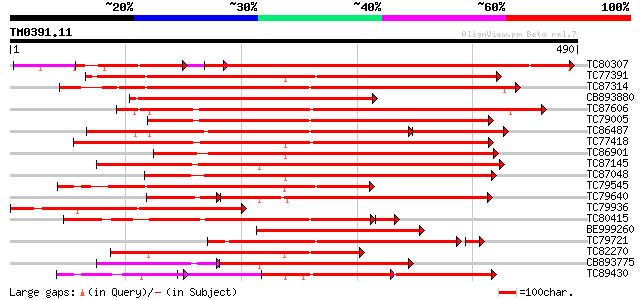
Score E
Sequences producing significant alignments: (bits) Value
TC80307 similar to GP|20160990|dbj|BAB89924. putative protein ki... 472 e-154
TC77391 similar to GP|14719339|gb|AAK73157.1 putative protein ki... 443 e-125
TC87314 similar to PIR|T00848|T00848 probable serine/threonine-s... 425 e-119
CB893880 similar to PIR|T51453|T51 serine/threonine specific pro... 400 e-112
TC87606 similar to PIR|C84473|C84473 probable protein kinase [im... 396 e-111
TC79005 similar to GP|19423982|gb|AAL87287.1 putative protein ki... 392 e-109
TC86487 similar to PIR|C84473|C84473 probable protein kinase [im... 309 e-105
TC77418 similar to GP|15146244|gb|AAK83605.1 At2g17220/T23A1.8 {... 350 9e-97
TC86901 similar to GP|13937147|gb|AAK50067.1 AT5g13160/T19L5_120... 344 5e-95
TC87145 similar to PIR|E71446|E71446 probable protein kinase - A... 330 6e-91
TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsi... 327 5e-90
TC79545 similar to GP|20160935|dbj|BAB89871. putative serine/thr... 310 1e-84
TC79640 similar to PIR|T02726|T02726 probable protein kinase [im... 269 6e-82
TC79936 similar to PIR|T51453|T51453 serine/threonine specific p... 298 4e-81
TC80415 homologue to GP|9542860|emb|CAB99493.1 protein kinase-li... 273 2e-79
BE999260 similar to GP|20160990|db putative protein kinase {Oryz... 281 5e-76
TC79721 similar to GP|22324431|dbj|BAC10348. putative serine/thr... 265 9e-72
TC82270 similar to GP|8778537|gb|AAF79545.1| F22G5.5 {Arabidopsi... 259 2e-69
CB893775 similar to GP|8978074|dbj protein serine/threonine kina... 224 7e-69
TC89430 similar to SP|P46573|APKB_ARATH Protein kinase APK1B (EC... 181 6e-46
>TC80307 similar to GP|20160990|dbj|BAB89924. putative protein kinase {Oryza
sativa (japonica cultivar-group)}, partial (69%)
Length = 1929
Score = 472 bits (1215), Expect(2) = e-154
Identities = 239/322 (74%), Positives = 270/322 (83%), Gaps = 2/322 (0%)
Frame = +1
Query: 169 GLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSL 228
GL + K L G + K AE+N+LGDL+H NLVKLI +CIEDDQRLLVYEFMPRGSL
Sbjct: 766 GLQLL*KPLTTMGFRVIKNGWAEVNFLGDLVHQNLVKLISYCIEDDQRLLVYEFMPRGSL 945
Query: 229 ENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSD 288
ENHLFRR +PLPWSIRMKIALGAAKGLAFLHE+++RP+IYRDFKTSNILLDA+YNAKLSD
Sbjct: 946 ENHLFRRSMPLPWSIRMKIALGAAKGLAFLHEEAERPVIYRDFKTSNILLDADYNAKLSD 1125
Query: 289 FGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSID 348
FGLAKDGPEG+KTHVSTRVMGTYGYAAPEYVMTGHL+SKSDVYSFGVVLLEM++GRRS+D
Sbjct: 1126 FGLAKDGPEGDKTHVSTRVMGTYGYAAPEYVMTGHLTSKSDVYSFGVVLLEMISGRRSMD 1305
Query: 349 KKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARP 408
K RPNGEHNLVEWARP LG RR F+++IDPRLEGHFSVKGAQKAAQLA CLSRDPKARP
Sbjct: 1306 KHRPNGEHNLVEWARPHLGERRRFYRLIDPRLEGHFSVKGAQKAAQLAHHCLSRDPKARP 1485
Query: 409 LMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKGQPLRIL 468
LMSEVV L PLPNLKDMA SSY+FQ+ + +R + PN ++ + Q SL + GQ R L
Sbjct: 1486 LMSEVVEALMPLPNLKDMASSSYYFQSMQAERFGASPNSRS-VHAQGASLARNGQQRRSL 1662
Query: 469 SSPNCPNGSPYSRY--SKSPKP 488
S PN + SPY +SPKP
Sbjct: 1663 SIPNGAHVSPYHHKYPQQSPKP 1728
Score = 95.1 bits (235), Expect = 5e-20
Identities = 70/199 (35%), Positives = 92/199 (46%), Gaps = 13/199 (6%)
Frame = +3
Query: 4 GDNNAIQAGSLDVDKSKGSKKK--------VDDGAKKERGWWFRFSFFGSCIPSRPKVD- 54
G +N S D+ KSKGSKK V D K + G W F G + +
Sbjct: 273 GSDNVKMVESWDMSKSKGSKKMKKDKENEDVIDAGKTKTGCWVGLRFIGKLHFFKIQS** 452
Query: 55 -SSISGTSTHNGNSIKSVITQSEENKS---GFEKNTEETEAPPESSTTTSNEESIASTPK 110
S ++ H + ++ Q E N+ + +A P+ + + +
Sbjct: 453 LSQMAVAQVHIMLKVNQLMIQVEANELLQ*SLLQPLVTQKAIPQLLNLKRSLKLLPGCEN 632
Query: 111 LFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGL 170
S L + +GL+V L + G GCVFKGWIEENGTAPVKPGTGL
Sbjct: 633 SLSMNLSWQQEI-----SGLRV---------FLAKVGLGCVFKGWIEENGTAPVKPGTGL 770
Query: 171 TVAVKILNHNGHQGHKEWL 189
TVAVK LNH+G QGHKEWL
Sbjct: 771 TVAVKTLNHDGLQGHKEWL 827
Score = 92.4 bits (228), Expect(2) = e-154
Identities = 55/96 (57%), Positives = 64/96 (66%)
Frame = +2
Query: 58 SGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELK 117
SGTSTH E+KS + + + AP SSTTTSN ES +ST KL EELK
Sbjct: 467 SGTSTHYA-----------ESKSTNDTSRGQRTAPVISSTTTSNAESNSSTTKL-EEELK 610
Query: 118 VASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFK 153
+AS LRKF+FN LK+ATRNFRPES LGEGGFG F+
Sbjct: 611 IASRLRKFSFNELKLATRNFRPESFLGEGGFGLCFQ 718
>TC77391 similar to GP|14719339|gb|AAK73157.1 putative protein kinase {Oryza
sativa}, partial (82%)
Length = 1753
Score = 443 bits (1140), Expect = e-125
Identities = 223/363 (61%), Positives = 278/363 (76%), Gaps = 3/363 (0%)
Frame = +3
Query: 66 NSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKF 125
N +KSV + +GF + SST+ ++ SI+ T + E L+ +S+L+ F
Sbjct: 312 NKVKSV----SPSNTGFTSRSVSRSGHDISSTSRNSSASISVTSRSEGEILQ-SSNLKSF 476
Query: 126 TFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGH 185
++N ++ ATRNFRP+S+LGEGGFG VFKGWI+E+ A KPG G+ VAVK LN GHQGH
Sbjct: 477 SYNEVRAATRNFRPDSVLGEGGFGSVFKGWIDEHSHAATKPGMGIIVAVKRLNQEGHQGH 656
Query: 186 KEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPWS 242
+EWLAE+NYLG L HPNLVKLIG+C ED+ RLLVYEFMP+GS+ENHLFRR P WS
Sbjct: 657 REWLAEINYLGQLQHPNLVKLIGYCFEDEHRLLVYEFMPKGSMENHLFRRGSYFQPFSWS 836
Query: 243 IRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTH 302
+RMKIALGAAKGLAFLH + +IYRDFKTSNILLD+ Y+AKLSDFGLA+DGP G+K+H
Sbjct: 837 LRMKIALGAAKGLAFLHSTEPK-VIYRDFKTSNILLDSNYDAKLSDFGLARDGPTGDKSH 1013
Query: 303 VSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWA 362
VSTRVMGT GYAAPEY+ TGHL++KSDVYSFGVVLLE+++GRR+IDK P+GEHNLVEWA
Sbjct: 1014VSTRVMGTRGYAAPEYLATGHLTAKSDVYSFGVVLLEIISGRRAIDKNLPSGEHNLVEWA 1193
Query: 363 RPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPN 422
+P L ++R F+++DPRLE +S A AA LA+QCLS +P+ RP M EVV TL+ L
Sbjct: 1194KPYLSNKRRVFRVMDPRLERQYSHSRAHAAAALASQCLSVEPRIRPNMDEVVKTLEQLQE 1373
Query: 423 LKD 425
KD
Sbjct: 1374PKD 1382
>TC87314 similar to PIR|T00848|T00848 probable serine/threonine-specific
protein kinase T20F6.6 (EC 2.7.1.-) - Arabidopsis
thaliana, partial (80%)
Length = 1575
Score = 425 bits (1093), Expect = e-119
Identities = 226/402 (56%), Positives = 289/402 (71%), Gaps = 4/402 (0%)
Frame = +1
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ S KVD++ S ST SG K T + P S + SN
Sbjct: 211 GNCLDSSAKVDAAQSSRST-----------------SGTSKTTPSSLTIP-SYSDKSNSS 336
Query: 104 SIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
S+ TP+ E L + +L+ F+FN LK ATRNFRP+SLLGEGGFG V+KGWI+E+
Sbjct: 337 SLLPTPRSEGEILS-SPNLKAFSFNELKNATRNFRPDSLLGEGGFGHVYKGWIDEHTFTA 513
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
KPG+G+ VAVK L G+QGHKEWL E+NYLG L HPNLVKLIG+C+E + RLLVYEFM
Sbjct: 514 AKPGSGMVVAVKRLKPEGYQGHKEWLTEVNYLGQLHHPNLVKLIGYCLEGENRLLVYEFM 693
Query: 224 PRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEY 282
P+GSLENHLFRR P PL WSIRMK+A+GAA+GL+FLH +++ +IYRDFK SNILLDAE+
Sbjct: 694 PKGSLENHLFRRGPQPLSWSIRMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDAEF 870
Query: 283 NAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT 342
N+KLSDFGLAK GP G++THVST+V+GT GYAAPEYV TG L++KSDVYSFGVV+LE+L+
Sbjct: 871 NSKLSDFGLAKAGPTGDRTHVSTQVVGTQGYAAPEYVATGRLTAKSDVYSFGVVMLELLS 1050
Query: 343 GRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSR 402
GRR++DK + NLV+WA+P LG +R F+I+D +LEG + KGA AA LA QCL+R
Sbjct: 1051GRRAVDKTIAGVDQNLVDWAKPYLGDKRRLFRIMDSKLEGQYPQKGAFMAATLALQCLNR 1230
Query: 403 DPKARPLMSEVVHTLKPLPNLKDM---AISSYHFQTARVDRT 441
+ KARP M+EV+ TL+ + K ++S +H A V R+
Sbjct: 1231EAKARPSMTEVLATLEQIEAPKHASRNSLSEHHRVHAPVRRS 1356
>CB893880 similar to PIR|T51453|T51 serine/threonine specific protein
kinase-like - Arabidopsis thaliana, partial (42%)
Length = 645
Score = 400 bits (1027), Expect = e-112
Identities = 194/215 (90%), Positives = 205/215 (95%)
Frame = +3
Query: 104 SIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
SI STP +FSEELKV+S LRKFTFN LK+ATRNFRPESLLGEGGFGCVFKGWIEENGTAP
Sbjct: 3 SIPSTP-MFSEELKVSSDLRKFTFNELKMATRNFRPESLLGEGGFGCVFKGWIEENGTAP 179
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
VKPGTGLTVAVK LNH+G QGHKEWLAELN LGD++HPNLVKLIGFCIEDDQRLLVY+FM
Sbjct: 180 VKPGTGLTVAVKTLNHDGLQGHKEWLAELNILGDIVHPNLVKLIGFCIEDDQRLLVYQFM 359
Query: 224 PRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYN 283
PRGSLENHLFRR LPLPWSIRMKIALGAAKGL FLHE++QRPIIYRDFKTSNILLDAEYN
Sbjct: 360 PRGSLENHLFRRSLPLPWSIRMKIALGAAKGLNFLHEEAQRPIIYRDFKTSNILLDAEYN 539
Query: 284 AKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEY 318
AKLSDFGLAKDGP+GE TH+STRVMGTYGYAAPEY
Sbjct: 540 AKLSDFGLAKDGPQGENTHISTRVMGTYGYAAPEY 644
>TC87606 similar to PIR|C84473|C84473 probable protein kinase [imported] -
Arabidopsis thaliana, partial (69%)
Length = 1803
Score = 396 bits (1018), Expect = e-111
Identities = 220/398 (55%), Positives = 276/398 (69%), Gaps = 26/398 (6%)
Frame = +3
Query: 93 PESSTTTSNEESIA--STPKLFSEELKVA---SSLRKFTFNGLKVATRNFRPESLLGEGG 147
P S++T+++ SI S P +E+L ++ S+L FT LKV T+ F + LG GG
Sbjct: 345 PRSASTSTHRISIEELSNPNA-TEDLSISLAGSNLYAFTVAELKVITQQFSSSNHLGAGG 521
Query: 148 FGCVFKGWIEENGTAPVKPGT-GLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKL 206
FG V KG+I++ V+PG +VAVK+L+ QGHKEWL E+ LG L P+LVKL
Sbjct: 522 FGPVHKGFIDDK----VRPGLKAQSVAVKLLDLESKQGHKEWLTEVVVLGQLRDPHLVKL 689
Query: 207 IGFCIEDDQRLLVYEFMPRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRP 265
IG+CIED+ RLLVYE++PRGSLEN LFRR LPWS RMKIA+GAAKGLAFLHE +++P
Sbjct: 690 IGYCIEDEHRLLVYEYLPRGSLENQLFRRFSASLPWSTRMKIAVGAAKGLAFLHE-AEQP 866
Query: 266 IIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLS 325
+I+RDFK SNILLD++YNAKLSDFGLAKDGPEG+ THVSTRVMGT GYAAPEYVMTGHL+
Sbjct: 867 VIFRDFKASNILLDSDYNAKLSDFGLAKDGPEGDDTHVSTRVMGTQGYAAPEYVMTGHLT 1046
Query: 326 SKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFS 385
+KSDVYSFGVVLLE+LTGR+S+DK RP E NLV+WARP+L R +I+DP+LEG +S
Sbjct: 1047AKSDVYSFGVVLLELLTGRKSVDKNRPQREQNLVDWARPMLIDSRKISKIMDPKLEGQYS 1226
Query: 386 VKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSY-------------- 431
GA+KAA LA QCLS PK+RP MS VV L+PL + D+ I +
Sbjct: 1227EMGAKKAASLAYQCLSHRPKSRPTMSNVVKILEPLQDFDDIPIGPFVYTVPIDNNEVAKE 1406
Query: 432 -----HFQTARVDRTMSMPNHKNGIRTQLVSLPKKGQP 464
++T R R + H+NG R + PK P
Sbjct: 1407KDQAKEYETNRERRRENDKYHRNGHRHHPLKSPKSPSP 1520
>TC79005 similar to GP|19423982|gb|AAL87287.1 putative protein kinase
{Arabidopsis thaliana}, partial (70%)
Length = 1723
Score = 392 bits (1008), Expect = e-109
Identities = 197/301 (65%), Positives = 242/301 (79%), Gaps = 2/301 (0%)
Frame = +3
Query: 120 SSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLT-VAVKILN 178
S++ FT+N L++AT+ FRP+ +LGEGGFG V+KG I+++ V+ G T VA+K LN
Sbjct: 324 SNVHIFTYNELRLATKQFRPDFILGEGGFGVVYKGVIDDS----VRAGYNSTEVAIKELN 491
Query: 179 HNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP-L 237
G QG +EWLAE+NYLG HPNLVKL G+C ED+ RLLVYE+M SLE HLFRR
Sbjct: 492 REGFQGDREWLAEVNYLGQFSHPNLVKLFGYCCEDEHRLLVYEYMASDSLEKHLFRRAGS 671
Query: 238 PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPE 297
L WS RMKIAL AA+GLAFLH ++RPIIYRDFKTSNILLDA++NAKLSDFGLAKDGP
Sbjct: 672 TLTWSKRMKIALHAARGLAFLH-GAERPIIYRDFKTSNILLDADFNAKLSDFGLAKDGPM 848
Query: 298 GEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
G++THVSTRVMGTYGYAAPEYVMTGHL+++SDVY FGVVLLE+L GRR++DK RP+ EHN
Sbjct: 849 GDQTHVSTRVMGTYGYAAPEYVMTGHLTARSDVYGFGVVLLELLIGRRALDKSRPSREHN 1028
Query: 358 LVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTL 417
LVEWARP+L H + +I+DP++EG +S K A K A LA QCLS++PK+RPLMS+VV L
Sbjct: 1029LVEWARPLLNHNKKLLKILDPKVEGQYSSKTATKVALLAYQCLSQNPKSRPLMSQVVEIL 1208
Query: 418 K 418
+
Sbjct: 1209E 1211
>TC86487 similar to PIR|C84473|C84473 probable protein kinase [imported] -
Arabidopsis thaliana, partial (75%)
Length = 2229
Score = 309 bits (792), Expect(2) = e-105
Identities = 168/292 (57%), Positives = 215/292 (73%), Gaps = 9/292 (3%)
Frame = +3
Query: 67 SIKSVITQSEENKSGFEKNTEETEAPPESSTTTS-NEESIAS----TPKLFSEELKVA-- 119
SI S Q E + + T+ET ++ T+S N S+ + SE+L ++
Sbjct: 417 SIFSSCYQGEGEYTSPKATTKETSTKVVATKTSSFNRISLTDLSFPSATTLSEDLSISLA 596
Query: 120 -SSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILN 178
S+L FT LK+ T+ F + LGEGGFG V KG+I++ A ++P VAVK+L+
Sbjct: 597 GSNLYVFTLAELKIITQGFSSSNFLGEGGFGPVHKGFIDDKVRAGLEPQP---VAVKLLD 767
Query: 179 HNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR-PL 237
+G QGH+EWL E+ +LG L H +LVKLIG+C E++ RLLVYE++PRGSLEN LF+R
Sbjct: 768 LDGTQGHREWLTEVVFLGQLRHQHLVKLIGYCCEEEHRLLVYEYLPRGSLENQLFKRYST 947
Query: 238 PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPE 297
LPWS RMKIA+GAAKGLAFLH D+++P+IYRDFK SNILLD++YNAKLSDFGLAKDGPE
Sbjct: 948 SLPWSTRMKIAVGAAKGLAFLH-DAKKPVIYRDFKGSNILLDSDYNAKLSDFGLAKDGPE 1124
Query: 298 GEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDK 349
G+ THVSTRVMGT GYAAPEY+MTGHL++ SDVYSFGVVLLE+LTGRRS+D+
Sbjct: 1125GDDTHVSTRVMGTQGYAAPEYIMTGHLTAMSDVYSFGVVLLELLTGRRSLDQ 1280
Score = 92.8 bits (229), Expect(2) = e-105
Identities = 47/83 (56%), Positives = 57/83 (68%)
Frame = +1
Query: 349 KKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARP 408
K RP E NLVEWARPVL R +I+DPRLEG +S GA+KAA LA QCLS P+ RP
Sbjct: 1279 KVRPPREQNLVEWARPVLNDARKLSRIMDPRLEGQYSEMGARKAAALAYQCLSHRPRNRP 1458
Query: 409 LMSEVVHTLKPLPNLKDMAISSY 431
MS VV+TL+PL + D+ I +
Sbjct: 1459 TMSTVVNTLEPLKDFDDIPIGPF 1527
>TC77418 similar to GP|15146244|gb|AAK83605.1 At2g17220/T23A1.8 {Arabidopsis
thaliana}, partial (74%)
Length = 1786
Score = 350 bits (897), Expect = 9e-97
Identities = 182/366 (49%), Positives = 244/366 (65%), Gaps = 3/366 (0%)
Frame = +1
Query: 56 SISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEE 115
S S ++ S IT S + F ++ + SS++ + + + +
Sbjct: 340 SYSQAASSGSYGTNSTITSSTATTTTFTTSSGSSSNMSSSSSSINRSNFSSVSDYSIGGQ 519
Query: 116 LKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVK 175
+ S+LR FTF LK AT+NFR ++LLGEGGFG V+KGW+E + + +G TVAVK
Sbjct: 520 ILPTSNLRVFTFAELKTATKNFRLDNLLGEGGFGKVYKGWLESS-----RNSSGTTVAVK 684
Query: 176 ILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR 235
LN G+QG +EW +E+++LG L HPNLVKL+G+C E+ + LLVYE+M RGSLENHLF R
Sbjct: 685 KLNTEGYQGFEEWQSEIHFLGRLYHPNLVKLLGYCYEETELLLVYEYMQRGSLENHLFGR 864
Query: 236 PL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLA 292
PLPW +R+KIA+GAA GL+FLH S R IIYRDFK SNILLD YNAK+SDFGLA
Sbjct: 865 GAAVQPLPWDLRLKIAIGAACGLSFLHT-SDREIIYRDFKASNILLDGSYNAKISDFGLA 1041
Query: 293 KDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRP 352
K GP ++H+ST VMGT GYAAPEY+ TGHL KSDVY FGVVL+E+LTG R++D RP
Sbjct: 1042KLGPSASQSHLSTTVMGTPGYAAPEYMQTGHLYVKSDVYGFGVVLVEILTGLRAVDLNRP 1221
Query: 353 NGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSE 412
+G H L +W +P L R+ +++DP+L + +K A A+LA CL+ +PK RP M +
Sbjct: 1222SGRHILTDWIKPELQDRKKLKKVMDPQLGDKYPIKAALPIAKLAISCLAPEPKLRPSMRD 1401
Query: 413 VVHTLK 418
V+ L+
Sbjct: 1402VLERLQ 1419
>TC86901 similar to GP|13937147|gb|AAK50067.1 AT5g13160/T19L5_120
{Arabidopsis thaliana}, partial (83%)
Length = 2028
Score = 344 bits (882), Expect = 5e-95
Identities = 176/301 (58%), Positives = 216/301 (71%), Gaps = 3/301 (0%)
Frame = +1
Query: 125 FTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQG 184
FTF L AT+NFRP+S LGEGGFG V+KG +E G A VAVK L+ NG QG
Sbjct: 493 FTFRELAAATKNFRPQSFLGEGGFGRVYKGRLETTGQA---------VAVKQLDRNGLQG 645
Query: 185 HKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPW 241
++E+L E+ L L PNLV LIG+C + DQRLLVYEFMP GSLE+HL P PL W
Sbjct: 646 NREFLVEVLMLSLLHSPNLVSLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPADKEPLDW 825
Query: 242 SIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKT 301
+ RMKIA GAAKGL +LH+ + P+IYRDFK+SNILLD Y+ KLSDFGLAK GP G+K+
Sbjct: 826 NTRMKIAAGAAKGLEYLHDKANPPVIYRDFKSSNILLDEGYHPKLSDFGLAKLGPVGDKS 1005
Query: 302 HVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEW 361
HVSTRVMGTYGY APEY MTG L+ KSDVYSFGVV LE++TGR++ID RP+GE NLV W
Sbjct: 1006HVSTRVMGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSTRPHGEQNLVTW 1185
Query: 362 ARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLP 421
ARP+ RR F ++ DPRL+G + ++G +A +A+ C+ ARPL+ +VV L L
Sbjct: 1186ARPLFNDRRKFSKLADPRLQGRYPMRGLYQALAVASMCIQEQAAARPLIGDVVTALSYLA 1365
Query: 422 N 422
N
Sbjct: 1366N 1368
>TC87145 similar to PIR|E71446|E71446 probable protein kinase - Arabidopsis
thaliana, partial (74%)
Length = 1443
Score = 330 bits (847), Expect = 6e-91
Identities = 174/357 (48%), Positives = 233/357 (64%), Gaps = 5/357 (1%)
Frame = +2
Query: 76 EENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATR 135
E++KS E + P S T++ SI+S + + S R FTF L+ AT
Sbjct: 239 EKSKSAPELQNKNKNKSPASKRATNSTGSISSPRSVKDLYREKEQSFRVFTFQELRDATN 418
Query: 136 NFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYL 195
F +GEGGFG V+KG I+ P G + VA+K LN G QGHKEWLAE+ +L
Sbjct: 419 GFNRMLKIGEGGFGSVYKGSIKP----PDGQGDPVIVAIKRLNTRGFQGHKEWLAEVQFL 586
Query: 196 GDLLHPNLVKLIGFCIEDD----QRLLVYEFMPRGSLENHLFRRPLP-LPWSIRMKIALG 250
+ HPNLVKL+G+C D QRLLVYEFMP SLENHLF R LP LPW R++I LG
Sbjct: 587 SIVNHPNLVKLLGYCSVDGERGIQRLLVYEFMPNRSLENHLFSRTLPVLPWKTRLEIMLG 766
Query: 251 AAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGT 310
AA+GLA+LHE + +IYRDFK+SN+LLD +++ KLSDFGLA++GP+G++THVST V+GT
Sbjct: 767 AAEGLAYLHEGLEIQVIYRDFKSSNVLLDEDFHPKLSDFGLAREGPQGDQTHVSTAVVGT 946
Query: 311 YGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRR 370
GYAAPEY+ TGHL +SD++SFGVVL E+LTGRRS+++ RP E L++W +
Sbjct: 947 QGYAAPEYIETGHLKVQSDMWSFGVVLYEILTGRRSLERNRPTTEQKLLDWVKQYPADTS 1126
Query: 371 MFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMA 427
F I+DPRL + + A+K A+LA CL ++ + RP MS++V LK +M+
Sbjct: 1127RFSMIMDPRLRNQYPIVAARKIAKLADSCLKKNAEDRPSMSQIVEGLKQALQFSEMS 1297
>TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsis
thaliana}, partial (77%)
Length = 1571
Score = 327 bits (839), Expect = 5e-90
Identities = 168/307 (54%), Positives = 214/307 (68%), Gaps = 3/307 (0%)
Frame = +1
Query: 117 KVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKI 176
K +++ F F L ATR F+ +L+GEGGFG VFKG + TG VAVK
Sbjct: 406 KSSTAAASFGFRELATATRGFKEANLIGEGGFGKVFKGRLS----------TGELVAVKQ 555
Query: 177 LNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRP 236
L+H+G QG +E++ E+ L L H NLVKLIG+C + DQRLLVYE+MP GSLE+HLF P
Sbjct: 556 LSHDGRQGFQEFVTEVLMLSLLHHSNLVKLIGYCTDGDQRLLVYEYMPMGSLEDHLFDLP 735
Query: 237 L---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAK 293
PL WS RMKIA+GAA+GL +LH + P+IYRD K++NILLD++++ KLSDFGLAK
Sbjct: 736 QDKEPLSWSSRMKIAVGAARGLEYLHCKADPPVIYRDLKSANILLDSDFSPKLSDFGLAK 915
Query: 294 DGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPN 353
GP G+ THVSTRVMGTYGY APEY M+G L+ KSD+YSFGVVLLE++TGRR+ID +
Sbjct: 916 LGPVGDNTHVSTRVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELITGRRAIDASKKP 1095
Query: 354 GEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEV 413
GE NLV W+RP RR F + DP L+GHF V+ +A + A CL PK RPL+ ++
Sbjct: 1096GEQNLVSWSRPYFSDRRKFVHMADPLLQGHFPVRCLHQAIAITAMCLQEQPKFRPLIGDI 1275
Query: 414 VHTLKPL 420
V L+ L
Sbjct: 1276VVALEYL 1296
>TC79545 similar to GP|20160935|dbj|BAB89871. putative
serine/threonine-specific protein kinase {Oryza sativa
(japonica cultivar-group)}, partial (44%)
Length = 848
Score = 310 bits (793), Expect = 1e-84
Identities = 162/277 (58%), Positives = 193/277 (69%), Gaps = 3/277 (1%)
Frame = +3
Query: 42 FFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSN 101
F G C+ +R K +S NG+S K + ET SS S
Sbjct: 81 FMGCCLSARIKAESP-----PRNGSSSKD--------------RSRETGLSGRSSGKVST 203
Query: 102 EESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGT 161
+ TP+ E LK +S+++ FTF+ LK ATRNFRP+S++GEGGFG VFKGWI+EN
Sbjct: 204 APTAPPTPRTEGEILK-SSNMKSFTFSELKTATRNFRPDSVVGEGGFGAVFKGWIDENTL 380
Query: 162 APVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYE 221
PV+PGTG+ +AVK LN G QGH EWL E+NYLG L HPNLVKLIG+C ED+ RLLVYE
Sbjct: 381 VPVRPGTGVVIAVKRLNQEGLQGHSEWLTEINYLGQLHHPNLVKLIGYCFEDEHRLLVYE 560
Query: 222 FMPRGSLENHLFRRP---LPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILL 278
F+ +GSL+NHLFRR PL WSIRMK+AL AAKGLA+LH D + +IYRDFKTSNILL
Sbjct: 561 FLTKGSLDNHLFRRASYFQPLSWSIRMKVALDAAKGLAYLHSDEAK-VIYRDFKTSNILL 737
Query: 279 DAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAA 315
D YNAKLSDFGLAKDGP G+ +HVSTRVMGTYGYAA
Sbjct: 738 DTNYNAKLSDFGLAKDGPAGDNSHVSTRVMGTYGYAA 848
>TC79640 similar to PIR|T02726|T02726 probable protein kinase [imported] -
Arabidopsis thaliana, partial (60%)
Length = 1210
Score = 269 bits (688), Expect(2) = 6e-82
Identities = 137/242 (56%), Positives = 175/242 (71%), Gaps = 7/242 (2%)
Frame = +1
Query: 183 QGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD----QRLLVYEFMPRGSLENHLFRRPLP 238
QGHKEW+ E+N LG + HPNLV+L+G+C EDD QRLLVYEFM SLE+HL R +P
Sbjct: 337 QGHKEWINEVNLLGVIKHPNLVRLVGYCAEDDERGIQRLLVYEFMSNKSLEDHLLVR-VP 513
Query: 239 ---LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDG 295
LPW R++IA AA+GLA+LHE+ +I+RDFKTSNILLD ++NAKLSDFGLA+ G
Sbjct: 514 SSVLPWITRLRIAQDAARGLAYLHEEMDFQLIFRDFKTSNILLDEDFNAKLSDFGLARQG 693
Query: 296 PEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGE 355
P +VST V+GT GYAAPEYV T L++KSDV+SFGVVL E++TGRR++++ P E
Sbjct: 694 PAEGFGYVSTAVVGTIGYAAPEYVHTEKLTAKSDVWSFGVVLYELITGRRAVERNLPKNE 873
Query: 356 HNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVH 415
L++W RP + + F I+DPRLE +K A K A LA +CL + PK+RP MSEVV
Sbjct: 874 QKLLDWVRPYVSDPKKFHHIVDPRLEAQDCIKSAHKLAVLANKCLMKQPKSRPKMSEVVE 1053
Query: 416 TL 417
L
Sbjct: 1054IL 1059
Score = 53.5 bits (127), Expect(2) = 6e-82
Identities = 28/65 (43%), Positives = 40/65 (61%)
Frame = +3
Query: 119 ASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILN 178
A+ L F+F+ LK+ATR F L+GEGGFG V++G + + + + VA+K LN
Sbjct: 150 ANHLHLFSFSDLKIATRGFNRALLIGEGGFGSVYRGTLIHHN----HHRSQIDVAIKQLN 317
Query: 179 HNGHQ 183
NGHQ
Sbjct: 318 RNGHQ 332
>TC79936 similar to PIR|T51453|T51453 serine/threonine specific protein
kinase-like - Arabidopsis thaliana, partial (22%)
Length = 869
Score = 298 bits (762), Expect = 4e-81
Identities = 158/207 (76%), Positives = 169/207 (81%), Gaps = 3/207 (1%)
Frame = +1
Query: 1 MGYGDNNAIQAGSLDVDKSKGSKKKVDDGAKKERGWWFRFSFFGSCIPSRPKVDSSI--- 57
MG+G+ N I+AGSL+VDKSK D KKER WFRFS G CIPSR K+D SI
Sbjct: 268 MGFGNKNVIKAGSLEVDKSKV------DVVKKERRCWFRFSCVGCCIPSRSKLDRSICCN 429
Query: 58 SGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELK 117
TST NG++ KSVI QSEENKS E NT+E+ +SSTTTSN ESI+ST K FSEELK
Sbjct: 430 GNTSTQNGDNTKSVIIQSEENKSTSETNTKESIGLVDSSTTTSNGESISSTSK-FSEELK 606
Query: 118 VASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL 177
VAS LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL
Sbjct: 607 VASCLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL 786
Query: 178 NHNGHQGHKEWLAELNYLGDLLHPNLV 204
NHNGHQGHKEWLAELNYLGDL HPNLV
Sbjct: 787 NHNGHQGHKEWLAELNYLGDLGHPNLV 867
>TC80415 homologue to GP|9542860|emb|CAB99493.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (49%)
Length = 1069
Score = 273 bits (697), Expect(2) = 2e-79
Identities = 146/271 (53%), Positives = 188/271 (68%), Gaps = 1/271 (0%)
Frame = +2
Query: 47 IPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIA 106
+P+ + +S+SGT +S++ +T S + S ET + P + +N +
Sbjct: 233 LPAGARNQTSLSGTGERKNSSLRHHLTDSASDLS-------ETCSTPRWNNANNNNTLLY 391
Query: 107 STPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKP 166
+ + FT L+ T++FR + +LGEGGFG V+KG+I+EN +K
Sbjct: 392 T-------------HVIAFTLYELETITKSFRADYILGEGGFGTVYKGYIDENVRVGLK- 529
Query: 167 GTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRG 226
L VAVK+LN G QGH+EWL E+N+LG L HPNLVKLIG+C EDD RLLVYEFM RG
Sbjct: 530 --SLPVAVKVLNKEGLQGHREWLTEVNFLGQLRHPNLVKLIGYCCEDDHRLLVYEFMFRG 703
Query: 227 SLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAK 285
SLENHLFR+ +PL W+ RM IALGAAKGLAFLH +++RP+IYRDFKTSNILLD++Y AK
Sbjct: 704 SLENHLFRKATVPLTWATRMMIALGAAKGLAFLH-NAERPVIYRDFKTSNILLDSDYTAK 880
Query: 286 LSDFGLAKDGPEGEKTHVSTRVMGTYGYAAP 316
LSDFGLAK GP+G++THVSTRVMGTYGYAAP
Sbjct: 881 LSDFGLAKAGPQGDETHVSTRVMGTYGYAAP 973
Score = 42.0 bits (97), Expect(2) = 2e-79
Identities = 18/21 (85%), Positives = 21/21 (99%)
Frame = +3
Query: 317 EYVMTGHLSSKSDVYSFGVVL 337
EYVMTGHL+++SDVYSFGVVL
Sbjct: 975 EYVMTGHLTARSDVYSFGVVL 1037
>BE999260 similar to GP|20160990|db putative protein kinase {Oryza sativa
(japonica cultivar-group)}, partial (29%)
Length = 442
Score = 281 bits (718), Expect = 5e-76
Identities = 138/145 (95%), Positives = 141/145 (97%)
Frame = +3
Query: 214 DQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKT 273
DQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHE +QRP+IYRDFKT
Sbjct: 6 DQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEGAQRPVIYRDFKT 185
Query: 274 SNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSF 333
SNILLDAEYN+KLSDFGLAKD PEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSF
Sbjct: 186 SNILLDAEYNSKLSDFGLAKDAPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSF 365
Query: 334 GVVLLEMLTGRRSIDKKRPNGEHNL 358
VVLLEMLTGRRSIDK RPNGEHNL
Sbjct: 366 *VVLLEMLTGRRSIDKIRPNGEHNL 440
>TC79721 similar to GP|22324431|dbj|BAC10348. putative
serine/threonine-specific protein kinase NAK {Oryza
sativa (japonica cultivar-group)}, partial (54%)
Length = 719
Score = 265 bits (677), Expect(2) = 9e-72
Identities = 133/221 (60%), Positives = 168/221 (75%), Gaps = 2/221 (0%)
Frame = +3
Query: 172 VAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENH 231
VA+K L QGHK AE+NYLG L H NLVKLIG+C++ RLLVYEFM +GSLENH
Sbjct: 6 VAIKNLKQESFQGHK---AEVNYLGQLHHENLVKLIGYCLQGKNRLLVYEFMQKGSLENH 176
Query: 232 LFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFG 290
LFR+ + P+ W+ R+ IA+G AKGLAFLH + +I+RD K SNILLD+++NA+LSDFG
Sbjct: 177 LFRKNVQPIAWATRVNIAVGVAKGLAFLHSLTAN-VIFRDLKASNILLDSDFNARLSDFG 353
Query: 291 LAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKK 350
LA+DGP G+ THVSTR++GT GYAAPEYV TGHL+ +SDVYSFGVVLLE+LTG+R+++
Sbjct: 354 LARDGPTGDNTHVSTRIIGTQGYAAPEYVATGHLTPRSDVYSFGVVLLELLTGKRAVEDA 533
Query: 351 RPN-GEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQ 390
RP E LV+WA P L R +I+D R+ G +S KGAQ
Sbjct: 534 RPGFSEETLVDWAMPFLSDSRRVLRIMDTRIGGQYSKKGAQ 656
Score = 23.9 bits (50), Expect(2) = 9e-72
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +2
Query: 395 LAAQCLSRDPKARPLM 410
LA QCL+ +PK RP M
Sbjct: 671 LALQCLNSNPKFRPPM 718
>TC82270 similar to GP|8778537|gb|AAF79545.1| F22G5.5 {Arabidopsis
thaliana}, partial (46%)
Length = 741
Score = 259 bits (662), Expect = 2e-69
Identities = 124/224 (55%), Positives = 167/224 (74%), Gaps = 5/224 (2%)
Frame = +2
Query: 88 ETEAPPESSTTTSNEESIASTPKLFSEELKV--ASSLRKFTFNGLKVATRNFRPESLLGE 145
+TE P + ++N+ S S P E ++ +S+L+ +T+ L AT NF P+++LGE
Sbjct: 65 KTELPCNTGLNSNNKVSAVSVPPTAQSEGEILPSSTLKSYTYAELSTATGNFFPDNVLGE 244
Query: 146 GGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVK 205
G FG VFKGWI+EN + KPG G+ +A+K LN + H+EWLAE+NY G HP+LV+
Sbjct: 245 GSFGAVFKGWIDENSFSAAKPGIGIVIAIKSLNQESFRSHREWLAEVNYHGRFSHPHLVR 424
Query: 206 LIGFCIEDDQRLLVYEFMPRGSLENHLFRRP---LPLPWSIRMKIALGAAKGLAFLHEDS 262
LIG+C+E++ RLLVYEFMPRGSLENHLFRR PL WS+R+K+AL AAKGLAFLH +
Sbjct: 425 LIGYCLENEHRLLVYEFMPRGSLENHLFRRGSYFQPLSWSLRLKVALDAAKGLAFLH-SA 601
Query: 263 QRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTR 306
+ +IYRDFKTSN+LLD++YNAKLSDFGLA+DGP G+K+H+ T+
Sbjct: 602 ENKVIYRDFKTSNVLLDSDYNAKLSDFGLARDGPTGDKSHIYTQ 733
>CB893775 similar to GP|8978074|dbj protein serine/threonine kinase-like
{Arabidopsis thaliana}, partial (50%)
Length = 858
Score = 224 bits (570), Expect(2) = 7e-69
Identities = 109/173 (63%), Positives = 138/173 (79%), Gaps = 5/173 (2%)
Frame = +1
Query: 182 HQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD----QRLLVYEFMPRGSLENHLFRRPL 237
H EWLAE+ +L + HPNLVKL+G+C D QRLLVYEFMP SLENHLF R L
Sbjct: 304 HSWISEWLAEVQFLSIVNHPNLVKLLGYCSVDGERGIQRLLVYEFMPNRSLENHLFSRTL 483
Query: 238 P-LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGP 296
P LPW R++I LGAA+GLA+LHE + +IYRDFK+SN+LLD +++ KLSDFGLA++GP
Sbjct: 484 PVLPWKTRLEIMLGAAEGLAYLHEGLEIQVIYRDFKSSNVLLDEDFHPKLSDFGLAREGP 663
Query: 297 EGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDK 349
+G++THVST V+GT GYAAPEY+ TGHL +SD++SFGVVL E+LTGRRS++K
Sbjct: 664 QGDQTHVSTAVVGTQGYAAPEYIETGHLKVQSDMWSFGVVLYEILTGRRSLEK 822
Score = 55.5 bits (132), Expect(2) = 7e-69
Identities = 38/108 (35%), Positives = 52/108 (47%)
Frame = +2
Query: 76 EENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATR 135
E++KS E + P S T++ SI+S + + S R FTF L+ AT
Sbjct: 8 EKSKSAPELQNKNKNKSPASKRATNSTGSISSPRSVKDLYREKEQSFRVFTFQELRDATN 187
Query: 136 NFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQ 183
F +GEGGFG V+KG I+ P G + VA+K LN G Q
Sbjct: 188 GFNRMLKIGEGGFGSVYKGSIK----PPDGQGDPVIVAIKRLNTRGFQ 319
>TC89430 similar to SP|P46573|APKB_ARATH Protein kinase APK1B (EC 2.7.1.-).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (71%)
Length = 1663
Score = 181 bits (459), Expect = 6e-46
Identities = 105/206 (50%), Positives = 132/206 (63%), Gaps = 3/206 (1%)
Frame = +2
Query: 218 LVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTS 274
LVYEFMPRGSLENHLF R PL WS+R+K+ALGAAK LAFLH ++ IY++ KTS
Sbjct: 680 LVYEFMPRGSLENHLFMRGSYFQPLSWSLRLKVALGAAKCLAFLHS-AETKSIYQNLKTS 856
Query: 275 NILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFG 334
NILLD+ YNAKL +FGLAKDG G VST T + L + S
Sbjct: 857 NILLDSNYNAKLFNFGLAKDGSTG*SKPVSTPGKSTDMQLQNTWPPVITLLRVMSIVS-E 1033
Query: 335 VVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQ 394
V +LE+L+GRR +DK RP +HNLVEWA+P L ++R + D RLEG + ++ A K A
Sbjct: 1034VGMLEILSGRRIVDKNRPLRQHNLVEWAKPCLSNKRKILHVFDNRLEGQYEIEDAYKVAI 1213
Query: 395 LAAQCLSRDPKARPLMSEVVHTLKPL 420
L+ +CLS + K RP M EVV L+ L
Sbjct: 1214LSLRCLSIEAKLRPNMDEVVTNLEQL 1291
Score = 107 bits (266), Expect(2) = 8e-31
Identities = 72/190 (37%), Positives = 94/190 (48%), Gaps = 2/190 (1%)
Frame = +1
Query: 146 GGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVK 205
G FG VFKG I+E+ + KPGTG+ VAVK LN +G + H E +AE+NYLG L HP+LVK
Sbjct: 463 GDFGSVFKGRIDEHSLSAAKPGTGIAVAVKRLNQDGVKSHIELMAEVNYLGQLTHPHLVK 642
Query: 206 LIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAA--KGLAFLHEDSQ 263
LIG+C+ED+ L L L L S G + ++ + +
Sbjct: 643 LIGYCLEDENSLFGL*IYATWQLGESLVHERLIFSTSFLEPTFEGCSWCCQMSCIPSQCR 822
Query: 264 RPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGH 323
I F+ L E + F K G YGYAAPEY+ TG+
Sbjct: 823 NKIDLPKFEDFKYLAGFEL*CETL*FWAGK-GWVHRVIEARVYTGQIYGYAAPEYLATGN 999
Query: 324 LSSKSDVYSF 333
+SKSDVYSF
Sbjct: 1000HTSKSDVYSF 1029
Score = 45.1 bits (105), Expect(2) = 8e-31
Identities = 35/116 (30%), Positives = 57/116 (48%), Gaps = 2/116 (1%)
Frame = +3
Query: 41 SFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTS 100
S G C+ + K + ++G S KSVI +E++ K +++ ++S
Sbjct: 177 SAMGVCLGGQVKAEGP-----KNSGLSSKSVIVNTEDHNILCYKGSDDL-------ISSS 320
Query: 101 NEESIASTPKLF--SEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKG 154
+E S S P+ E+ +S+L+ FT L+ ATRNFR + +LGE F F G
Sbjct: 321 SEVSATSVPRTLPCEGEILQSSNLKSFTLTELQAATRNFRVDIVLGEWRFWISF*G 488
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,124,792
Number of Sequences: 36976
Number of extensions: 219973
Number of successful extensions: 2584
Number of sequences better than 10.0: 646
Number of HSP's better than 10.0 without gapping: 1962
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1998
length of query: 490
length of database: 9,014,727
effective HSP length: 100
effective length of query: 390
effective length of database: 5,317,127
effective search space: 2073679530
effective search space used: 2073679530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0391.11


