

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
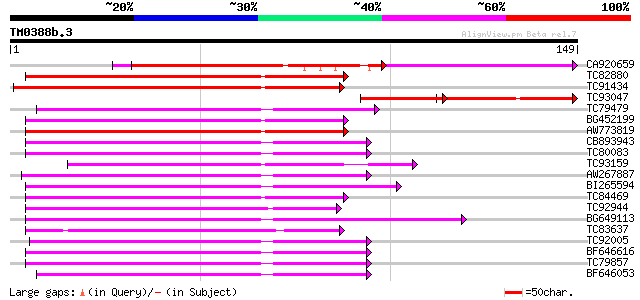
Score E
Sequences producing significant alignments: (bits) Value
CA920659 similar to GP|21740675|em OSJNBa0084K20.18 {Oryza sativ... 89 7e-19
TC82880 similar to GP|9858483|gb|AAG01054.1| resistance protein ... 70 4e-13
TC91434 weakly similar to PIR|T06144|T06144 disease resistance p... 69 5e-13
TC93047 46 1e-11
TC79479 similar to GP|12056928|gb|AAG48132.1 putative resistance... 62 9e-11
BG452199 weakly similar to GP|10178211|db TMV resistance protein... 60 3e-10
AW773819 similar to GP|9858483|gb|A resistance protein MG55 {Gly... 60 4e-10
CB893943 weakly similar to PIR|T52347|T52 disease resistance pro... 59 6e-10
TC80083 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 58 2e-09
TC93159 weakly similar to GP|10177477|dbj|BAB10868. disease resi... 57 2e-09
AW267887 weakly similar to GP|22037367|gb| disease resistance-li... 56 5e-09
BI265594 weakly similar to GP|9965103|gb| resistance protein LM6... 56 6e-09
TC84469 similar to GP|9858483|gb|AAG01054.1| resistance protein ... 55 1e-08
TC92944 similar to GP|9858483|gb|AAG01054.1| resistance protein ... 54 3e-08
BG649113 similar to GP|18033111|gb functional candidate resistan... 53 4e-08
TC83637 similar to GP|9279731|dbj|BAB01321.1 disease resistance ... 53 5e-08
TC92005 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 52 9e-08
BF646616 similar to GP|9965103|gb|A resistance protein LM6 {Glyc... 52 1e-07
TC79857 51 2e-07
BF646053 weakly similar to GP|18033111|g functional candidate re... 51 2e-07
>CA920659 similar to GP|21740675|em OSJNBa0084K20.18 {Oryza sativa}, partial
(2%)
Length = 790
Score = 89.0 bits (219), Expect = 7e-19
Identities = 60/148 (40%), Positives = 76/148 (50%), Gaps = 26/148 (17%)
Frame = +3
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNS-----LRIL-- 80
LERL+ G + + + PS+ L+ LA+L+L CS L SL +C +S ++L
Sbjct: 339 LERLNLEG-NSFVSLPPSMERLSSLAYLNLAHCSWLQSLPKLQLCATSSYGGRYFKMLSG 515
Query: 81 ---HLSG-----CTRLESTPN-----------FIGNPSHFRCGFDIVVPGNKIPYWFDSQ 121
H SG C LE + NP HFRCGFDIVVPG KIP WF+ Q
Sbjct: 516 SHNHRSGLYIFNCPLLEIAEGGQSLALSWLERLVENPHHFRCGFDIVVPGEKIPEWFNHQ 695
Query: 122 FKGGSRIREVDHVYEQDHWLGFAFCAVF 149
F G I+ D E D+WLGFAFC F
Sbjct: 696 FTGKFIIKISDFYNEFDNWLGFAFCVAF 779
Score = 50.8 bits (120), Expect = 2e-07
Identities = 30/67 (44%), Positives = 42/67 (61%)
Frame = +3
Query: 33 FTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
F+ C++L + SI +LT+L FLSL C++LVS+ C + SL L L GC RLE+ P
Sbjct: 6 FSPCSSLSTIGQSIAVLTQLKFLSLRDCTNLVSIPKSINC-MTSLVTLDLCGCLRLENLP 182
Query: 93 NFIGNPS 99
+GN S
Sbjct: 183--LGNVS 197
Score = 37.0 bits (84), Expect = 0.003
Identities = 31/94 (32%), Positives = 40/94 (41%), Gaps = 20/94 (21%)
Frame = +3
Query: 27 RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVC-------------- 72
+L+ L CTNL+ + SI +T L L L GC L +L LG V
Sbjct: 60 QLKFLSLRDCTNLVSIPKSINCMTSLVTLDLCGCLRLENLPLGNVSVSAENMDYFSVSGE 239
Query: 73 ------ELNSLRILHLSGCTRLESTPNFIGNPSH 100
L SL L LS C L + P+ IG+ H
Sbjct: 240 IVDHSYYLESLIFLDLSFC-NLSTVPDAIGDLWH 338
>TC82880 similar to GP|9858483|gb|AAG01054.1| resistance protein MG55
{Glycine max}, partial (46%)
Length = 1238
Score = 69.7 bits (169), Expect = 4e-13
Identities = 37/85 (43%), Positives = 57/85 (66%)
Frame = +2
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS+SKYL TP+F LE+L C +LL+VHPSIG L L ++L+ C+SL
Sbjct: 308 LKILNLSHSKYLKRTPDFSKLPNLEKLIMKDCQSLLEVHPSIGDLNNLLLINLKDCTSLS 487
Query: 65 SLYLGTVCELNSLRILHLSGCTRLE 89
+L + +L +++ L LSGC++++
Sbjct: 488 NL-PREIYQLRTVKTLILSGCSKID 559
>TC91434 weakly similar to PIR|T06144|T06144 disease resistance protein
homolog F24J7.70 - Arabidopsis thaliana, partial (4%)
Length = 940
Score = 69.3 bits (168), Expect = 5e-13
Identities = 38/87 (43%), Positives = 59/87 (67%)
Frame = +1
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+P L+RLDLS+SK L + +F LE L+ C L+++ PSIGLL +L +L+LE C
Sbjct: 286 LPNLRRLDLSDSKKLEKIEDFGQFPNLEWLNLERCIKLVELDPSIGLLRKLVYLNLERCY 465
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRL 88
+LVS+ + L+SL+ L++SGC++L
Sbjct: 466 NLVSI-PNNIFGLSSLKYLNMSGCSKL 543
>TC93047
Length = 812
Score = 45.8 bits (107), Expect(2) = 1e-11
Identities = 19/37 (51%), Positives = 25/37 (67%)
Frame = +3
Query: 113 KIPYWFDSQFKGGSRIREVDHVYEQDHWLGFAFCAVF 149
+ P WFD QF G SR++ D+ + D+WLGFAFC F
Sbjct: 111 QFPLWFDHQFAGNSRVKITDY-NKFDNWLGFAFCVAF 218
Score = 38.9 bits (89), Expect(2) = 1e-11
Identities = 15/23 (65%), Positives = 17/23 (73%)
Frame = +2
Query: 93 NFIGNPSHFRCGFDIVVPGNKIP 115
N + NP HFRCG DIVVP + IP
Sbjct: 50 NLVKNPCHFRCGLDIVVPSDTIP 118
>TC79479 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (21%)
Length = 1087
Score = 62.0 bits (149), Expect = 9e-11
Identities = 33/90 (36%), Positives = 49/90 (53%)
Frame = +1
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L+ K + P+ G+ LERL C NL+++H S+G L +L L+L C+ L +L
Sbjct: 319 LNFDECKIITHIPDVSGAPNLERLSLDSCENLVEIHDSVGFLDKLEILNLGSCAKLRNL- 495
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L SL+ L+LS C+ L S P +GN
Sbjct: 496 --PPIHLTSLQHLNLSHCSSLVSFPEILGN 579
Score = 28.1 bits (61), Expect = 1.4
Identities = 25/86 (29%), Positives = 39/86 (45%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ L+LS+ L+ P G+ + T + + SIG L L L L GC +L
Sbjct: 517 LQHLNLSHCSSLVSFPEILGNMKNITSLSLEYTAIREFPYSIGNLPRLKSLELHGCGNL- 693
Query: 65 SLYLGTVCELNSLRILHLSGCTRLES 90
L ++ L+ L L + C L+S
Sbjct: 694 -LLPSSIILLSELEELSIWQCEGLKS 768
>BG452199 weakly similar to GP|10178211|db TMV resistance protein N
{Arabidopsis thaliana}, partial (4%)
Length = 666
Score = 60.5 bits (145), Expect = 3e-10
Identities = 36/85 (42%), Positives = 51/85 (59%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS+S +L +TP+F LE+L T C L +V SIG L ++ ++LE C SL
Sbjct: 405 LKILNLSHSHFLSKTPDFSNLPNLEQLVLTDCPKLSEVSQSIGDLNKILLINLEDCISLQ 584
Query: 65 SLYLGTVCELNSLRILHLSGCTRLE 89
SL ++ L SL L LSGC ++
Sbjct: 585 SL-PRSIYNLKSLNTLILSGCLMID 656
>AW773819 similar to GP|9858483|gb|A resistance protein MG55 {Glycine max},
partial (20%)
Length = 633
Score = 59.7 bits (143), Expect = 4e-10
Identities = 35/85 (41%), Positives = 52/85 (61%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS+S YL +TP+F LE+L T C L +V SI L ++ ++LE C L
Sbjct: 361 LKILNLSHSHYLTQTPDFSNLPNLEQLVLTDCPRLSEVSHSIEHLNKILLINLEDCIGLQ 540
Query: 65 SLYLGTVCELNSLRILHLSGCTRLE 89
SL ++ +L SL+ L LSGC+ ++
Sbjct: 541 SL-PRSIYKLKSLKTLILSGCSMID 612
>CB893943 weakly similar to PIR|T52347|T52 disease resistance protein
RPP1-WsB [imported] - Arabidopsis thaliana (fragment),
partial (2%)
Length = 545
Score = 59.3 bits (142), Expect = 6e-10
Identities = 35/91 (38%), Positives = 47/91 (51%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K L L N +YL PN G LE+ F C NL+ +H SIG L +L L+ C+ L
Sbjct: 105 MKVLTLDNCRYLTHIPNVSGLPNLEKFSFRFCDNLIAIHDSIGNLNKLEILNAWACTKLE 284
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
+ L SL+ L LS C RL+S P +
Sbjct: 285 NF---PPLWLPSLKELDLSFCKRLKSFPELL 368
>TC80083 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (4%)
Length = 1269
Score = 57.8 bits (138), Expect = 2e-09
Identities = 35/91 (38%), Positives = 47/91 (51%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K L L KYL P+ G LE+L F C NL+ +H SIG L +L LS GC+ +
Sbjct: 249 MKVLILDRCKYLTHIPDVSGLSNLEKLSFERCYNLITIHNSIGHLNKLERLSAYGCNKVE 428
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
L SL+ L+LS C L+S P +
Sbjct: 429 HF---PPLGLASLKELNLSSCRSLKSFPELL 512
>TC93159 weakly similar to GP|10177477|dbj|BAB10868. disease resistance
protein-like {Arabidopsis thaliana}, partial (4%)
Length = 729
Score = 57.4 bits (137), Expect = 2e-09
Identities = 33/92 (35%), Positives = 54/92 (57%)
Frame = +1
Query: 16 LMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELN 75
L++ +F LE+L+ GC NL+++ PSIGLL +L +L+L C +LVS+ + L+
Sbjct: 61 LVKILDFGEFPNLEKLNLEGCINLVELDPSIGLLRKLVYLNLYECKNLVSI-PNNIFSLS 237
Query: 76 SLRILHLSGCTRLESTPNFIGNPSHFRCGFDI 107
SL L++ GC+++ NP H + DI
Sbjct: 238 SLEDLNMYGCSKV------FKNPMHLKKKHDI 315
>AW267887 weakly similar to GP|22037367|gb| disease resistance-like protein
GS4B-1 {Glycine max}, partial (34%)
Length = 609
Score = 56.2 bits (134), Expect = 5e-09
Identities = 31/92 (33%), Positives = 49/92 (52%)
Frame = +2
Query: 4 FLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
++K L L++ +YL + P+ G LE+L F C NL+ +H S+G L L L + C L
Sbjct: 155 YMKVLKLNSCQYLTQIPDVSGLPNLEKLSFQFCENLITIHNSVGFLNRLEILDAKYCIKL 334
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
S+ +L L+ L L+ C L+S P +
Sbjct: 335 QSV---PPLQLPCLKRLELAMCKSLKSFPELL 421
>BI265594 weakly similar to GP|9965103|gb| resistance protein LM6 {Glycine
max}, partial (15%)
Length = 664
Score = 55.8 bits (133), Expect = 6e-09
Identities = 35/99 (35%), Positives = 50/99 (50%)
Frame = +2
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K L L +++YL P+ G Q LE+ F C NL+ + SIG L +L LS GCS L
Sbjct: 194 MKVLTLDDNEYLTHIPDLSGLQNLEKFSFKYCENLITIDNSIGHLNKLERLSAFGCSKLE 373
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRC 103
L SL+ L+L C L+S P + ++ C
Sbjct: 374 RF---PPLGLASLKELNLCCCDSLKSFPKLLCEMTNIDC 481
>TC84469 similar to GP|9858483|gb|AAG01054.1| resistance protein MG55
{Glycine max}, partial (39%)
Length = 735
Score = 55.1 bits (131), Expect = 1e-08
Identities = 34/85 (40%), Positives = 49/85 (57%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS+S +L +TP+F LE+L C L QV SIG L ++ ++L+ C SL
Sbjct: 189 LKILNLSHSHHLTQTPDFSYLPNLEKLVLEDCPRLSQVSHSIGHLKKVVLINLKDCISLC 368
Query: 65 SLYLGTVCELNSLRILHLSGCTRLE 89
SL + L +L L LSGC ++
Sbjct: 369 SL-PRNIYTLKTLNTLILSGCLMID 440
>TC92944 similar to GP|9858483|gb|AAG01054.1| resistance protein MG55
{Glycine max}, partial (12%)
Length = 673
Score = 53.5 bits (127), Expect = 3e-08
Identities = 32/83 (38%), Positives = 50/83 (59%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS+S L TP+F LE+L C +L V +IG L ++ ++L+ C+ L
Sbjct: 324 LKILNLSHSHNLRHTPDFSKLPNLEKLILKDCPSLSSVSSNIGHLKKILLINLKDCTGLR 503
Query: 65 SLYLGTVCELNSLRILHLSGCTR 87
L ++ +L+SL+ L LSGCT+
Sbjct: 504 ELPR-SIYKLDSLKTLILSGCTK 569
>BG649113 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (2%)
Length = 782
Score = 53.1 bits (126), Expect = 4e-08
Identities = 36/116 (31%), Positives = 57/116 (49%)
Frame = +1
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K L L + +YL P+ LE+L F C NL+ +H SIG L++L LS+ G L
Sbjct: 70 MKILTLDDCEYLTHIPDVSSLSNLEKLSFEHCKNLITIHNSIGHLSKLERLSVTGYRKLK 249
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDS 120
L SL+ L+L G + LE+ P + +H + + K+P+ F +
Sbjct: 250 HF---PPLGLASLKELNLMGGSCLENFPELLCKMAHIKEIDIFYISIGKLPFSFQN 408
>TC83637 similar to GP|9279731|dbj|BAB01321.1 disease resistance protein
RPP1-WsB {Arabidopsis thaliana}, partial (2%)
Length = 1045
Score = 52.8 bits (125), Expect = 5e-08
Identities = 35/84 (41%), Positives = 46/84 (54%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK + +S + L E P+F + L+ L T C NL VHPSI L +L L L GC SL
Sbjct: 6 LKEVTISLAS-LKELPDFSKATNLKVLTVTVCPNLESVHPSIFTLEKLVRLDLGGCRSLT 182
Query: 65 SLYLGTVCELNSLRILHLSGCTRL 88
+ + L+SL L LSGC +L
Sbjct: 183TFTSNS--NLSSLHYLSLSGCEKL 248
Score = 29.6 bits (65), Expect = 0.48
Identities = 29/106 (27%), Positives = 45/106 (42%), Gaps = 18/106 (16%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTEL--------AFLS 56
L RLDL + L + L L +GC L + ++ + EL A S
Sbjct: 144 LVRLDLGGCRSLTTFTSNSNLSSLHYLSLSGCEKLSEFSVTLENIVELDLSWCPINALPS 323
Query: 57 LEGC-SSLVSLYL---------GTVCELNSLRILHLSGCTRLESTP 92
GC S+L +L L ++ +L LR L++ GC +L + P
Sbjct: 324 SFGCQSNLETLVLKATQIESIPSSIKDLTRLRKLNICGCKKLLALP 461
>TC92005 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (7%)
Length = 858
Score = 52.0 bits (123), Expect = 9e-08
Identities = 32/90 (35%), Positives = 43/90 (47%)
Frame = +1
Query: 6 KRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVS 65
K L L + +YL P+ G +E+ F C NL+ + SIG +L F+S GCS L
Sbjct: 1 KVLTLDDCEYLTHIPDVSGLSNIEKFSFKFCRNLITIDDSIGHQNKLEFISAIGCSKLKR 180
Query: 66 LYLGTVCELNSLRILHLSGCTRLESTPNFI 95
L SL+ L LS C L S P +
Sbjct: 181F---PPLGLASLKELELSFCVSLNSFPELL 261
>BF646616 similar to GP|9965103|gb|A resistance protein LM6 {Glycine max},
partial (5%)
Length = 671
Score = 51.6 bits (122), Expect = 1e-07
Identities = 33/91 (36%), Positives = 42/91 (45%)
Frame = +2
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L L +YL + N LE F C NL+ +H SIG L +L + GC L
Sbjct: 170 LKVLILDRCQYLTDISNVSNLPNLEEFSFQDCKNLITIHNSIGHLNKLEIIRATGCCKLE 349
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
S L SL+ L LS C L+S P +
Sbjct: 350 SF---PPLLLPSLKELELSYCESLKSFPELL 433
>TC79857
Length = 1217
Score = 50.8 bits (120), Expect = 2e-07
Identities = 31/91 (34%), Positives = 44/91 (48%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
+K L N + L+ TP+ LE+ F C NL+ +H S+ L L L+ EGC L
Sbjct: 207 MKVLIFDNCQDLIYTPDVSWLPNLEKFSFARCHNLVTIHNSLRYLNRLEILNAEGCEKLE 386
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
S + SL+ L LS C L+S P +
Sbjct: 387 SF---PPLQSPSLQNLELSNCKSLKSFPELL 470
>BF646053 weakly similar to GP|18033111|g functional candidate resistance
protein KR1 {Glycine max}, partial (9%)
Length = 644
Score = 50.8 bits (120), Expect = 2e-07
Identities = 32/88 (36%), Positives = 44/88 (49%)
Frame = +3
Query: 8 LDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLY 67
L L + +YL P+ G LE+L F C NL+ +H SIG L +L LS GC L
Sbjct: 90 LILDHCEYLTHIPDVSGLSNLEKLSFECCYNLITIHNSIGHLNKLERLSAFGCRKLKRF- 266
Query: 68 LGTVCELNSLRILHLSGCTRLESTPNFI 95
L SL+ L + C+ L+S P +
Sbjct: 267 --PPLGLASLKELDICCCSSLKSFPELL 344
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,881,091
Number of Sequences: 36976
Number of extensions: 93227
Number of successful extensions: 636
Number of sequences better than 10.0: 101
Number of HSP's better than 10.0 without gapping: 595
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 619
length of query: 149
length of database: 9,014,727
effective HSP length: 87
effective length of query: 62
effective length of database: 5,797,815
effective search space: 359464530
effective search space used: 359464530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0388b.3


