

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
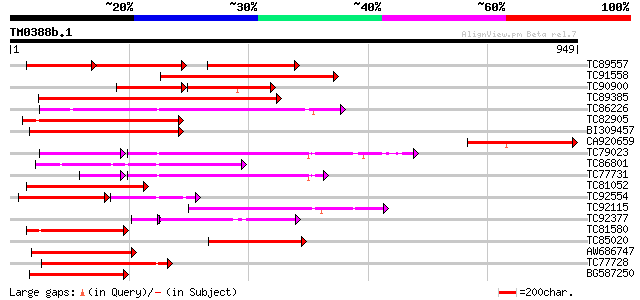
Query= TM0388b.1
(949 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 226 e-146
TC91558 weakly similar to GP|19774201|gb|AAL99077.1 NBS-LRR-Toll... 382 e-106
TC90900 weakly similar to PIR|D96753|D96753 Similar to disease r... 206 6e-99
TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 358 7e-99
TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 285 4e-77
TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanu... 254 1e-67
BI309457 weakly similar to GP|15787901|gb resistance gene analog... 249 3e-66
CA920659 similar to GP|21740675|em OSJNBa0084K20.18 {Oryza sativ... 237 2e-62
TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 230 2e-60
TC86801 similar to PIR|A54810|A54810 TMV resistance protein N - ... 218 1e-56
TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 205 6e-53
TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 204 1e-52
TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance ... 133 1e-49
TC92115 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 178 8e-45
TC92377 similar to GP|19774149|gb|AAL99051.1 NBS-LRR-Toll resist... 160 5e-44
TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 174 1e-43
TC85020 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 174 2e-43
AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein ... 173 3e-43
TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance... 169 5e-42
BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 168 8e-42
>TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 1567
Score = 226 bits (576), Expect(3) = e-146
Identities = 109/158 (68%), Positives = 126/158 (78%)
Frame = +1
Query: 138 KQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVR 197
KQ VFPVFYDVDPS V+ QNGVYENAFV H FKH++ +V RW+ M L G+AGWDVR
Sbjct: 523 KQVVFPVFYDVDPSHVKKQNGVYENAFVLHTEAFKHNSGKVARWRTTMTYLGGTAGWDVR 702
Query: 198 NKPEFREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWG 257
+PEF IE IVEAVI+ LGR FSG DDLIGIQP +E LENLLKL+S+ C+V+GIWG
Sbjct: 703 VEPEFEMIEKIVEAVIKKLGRTFSGSTDDLIGIQPHIEALENLLKLSSKDDGCRVLGIWG 882
Query: 258 MGGIGKTTLATVLYDRISHLFEARCFVENVSKVYRDGG 295
M GIGKTTLATVLYD+IS F+A CF+ENVSK+Y DGG
Sbjct: 883 MDGIGKTTLATVLYDKISFQFDACCFIENVSKIYEDGG 996
Score = 174 bits (440), Expect(3) = e-146
Identities = 84/153 (54%), Positives = 112/153 (72%)
Frame = +3
Query: 332 LKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNND 391
LK+L+VLDNV+QL QL++ + P S++II TRD+HIL+ YGA V+E LMN+ D
Sbjct: 1107 LKLLIVLDNVEQLEQLEKLDIEPKFLHPRSKIIIITRDKHILQAYGADEVFEAKLMNDED 1286
Query: 392 ARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLK 451
A +L RK FKSD SS AEL+P+VL YAQ LPLA++V GSFL +RNA +W LD+ +
Sbjct: 1287 AHKLLCRKAFKSDYPSSGFAELIPKVLVYAQRLPLAVKVLGSFLFSRNANEWSSTLDKFE 1466
Query: 452 NNPDNKVMDVLQISFEGLHSEDKEIFLHIACFF 484
NP NK+M LQ+S+EGL ++KE+FLH+A FF
Sbjct: 1467 KNPPNKIMKALQVSYEGLEKDEKEVFLHVAXFF 1565
Score = 160 bits (404), Expect(3) = e-146
Identities = 83/117 (70%), Positives = 95/117 (80%)
Frame = +3
Query: 29 LAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKK 88
L + G TASSS+T SN + Y+YDVFISFRGSDTRN+FVDHLYAHL RKGIF FKDDK+
Sbjct: 198 LPREGGTASSSSTD-SNSNQSYRYDVFISFRGSDTRNSFVDHLYAHLNRKGIFTFKDDKQ 374
Query: 89 LQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVF 145
LQKGE+IS QLLQAI+ SR+SIVVFSK+YA S WCLDEMAAI E + K + FP F
Sbjct: 375 LQKGEAISPQLLQAIQQSRISIVVFSKDYASSTWCLDEMAAINESRINLKTSCFPCF 545
Score = 36.6 bits (83), Expect = 0.047
Identities = 19/50 (38%), Positives = 32/50 (64%)
Frame = +2
Query: 284 VENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLK 333
++ ++K R AVQKQ++ QT++E N++T S +IS +R+RL K
Sbjct: 962 LKTLAKFMRMVVPVAVQKQIICQTIEEKNIDTCSARKISQTMRNRLCKTK 1111
>TC91558 weakly similar to GP|19774201|gb|AAL99077.1 NBS-LRR-Toll resistance
gene analog protein {Medicago edgeworthii}, partial
(60%)
Length = 942
Score = 382 bits (982), Expect = e-106
Identities = 196/300 (65%), Positives = 232/300 (77%), Gaps = 2/300 (0%)
Frame = +1
Query: 253 IGIWGMGGIGKTTLATVLYDRISHL--FEARCFVENVSKVYRDGGVTAVQKQVLRQTVDE 310
+GIW M GIGKTTLA VLYD ISH F+A CFVE+VSK+YRDGG AVQK++L QT+ E
Sbjct: 1 VGIWRMDGIGKTTLANVLYDTISHQYQFDACCFVEDVSKIYRDGGAIAVQKRILDQTIKE 180
Query: 311 MNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDE 370
NLE YSPSEISGI+ +RL LK+L+VLDNVDQ VQLQE +NP GSR+IITTRD+
Sbjct: 181 KNLEGYSPSEISGIISNRLYKLKLLLVLDNVDQSVQLQELHINPISLCAGSRIIITTRDK 360
Query: 371 HILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRV 430
HIL Y A IVYE L+N+NDA EL RK FKSD SS EL+PEVLKYAQGLPLAIRV
Sbjct: 361 HILIEYEADIVYEAELLNDNDAHELLCRKAFKSDYSSSDYEELIPEVLKYAQGLPLAIRV 540
Query: 431 TGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKEN 490
GSFL R QWR AL+ +NNPD+ +M VL+ SFEGL +KEIFLH+ACFF GE+E+
Sbjct: 541 MGSFLYKRKTAQWRAALEGWQNNPDSGIMKVLRSSFEGLELREKEIFLHVACFFDGERED 720
Query: 491 YVKRILDACGLHPHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSR 550
YV+RIL ACGL P+IGI +E+SLITIRNQE HMHEM+Q+LG+ VR+ P+EP WS+
Sbjct: 721 YVRRILHACGLQPNIGIPTSVEKSLITIRNQENHMHEMLQELGETNVRETHPDEPRLWSK 900
>TC90900 weakly similar to PIR|D96753|D96753 Similar to disease resistance
proteins [imported] - Arabidopsis thaliana, partial (7%)
Length = 826
Score = 206 bits (523), Expect(2) = 6e-99
Identities = 106/155 (68%), Positives = 121/155 (77%), Gaps = 8/155 (5%)
Frame = +3
Query: 298 AVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLF 357
++QKQ+LRQT+DE LETYSPSEISGIVR RL + K LVVLDNVD L Q++E A+NP L
Sbjct: 360 SLQKQILRQTIDEKYLETYSPSEISGIVRKRLCNRKFLVVLDNVDLLEQVEELAINPELV 539
Query: 358 QKGSRMIITTRDEHILKVYGAH--------IVYEVPLMNNNDARELFYRKGFKSDNLSSR 409
KGSRMIITTR+ HIL+VYG + YEVPL+NNNDARELFYRK FKS + +S
Sbjct: 540 GKGSRMIITTRNMHILRVYGEQLSLSHGTCVSYEVPLLNNNDARELFYRKAFKSQDPASE 719
Query: 410 CAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWR 444
C L PEVLKY +GLPLAIRV GSFLCTRNA QWR
Sbjct: 720 CLNLTPEVLKYVEGLPLAIRVVGSFLCTRNANQWR 824
Score = 174 bits (442), Expect(2) = 6e-99
Identities = 84/116 (72%), Positives = 97/116 (83%)
Frame = +2
Query: 180 RWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLEN 239
RW +AM LA GWDVRNKPEFREIENIV+ VI+ LG KFSGFADDLI QPRVE LE+
Sbjct: 5 RWTKAMGRLAELVGWDVRNKPEFREIENIVQEVIKTLGHKFSGFADDLIATQPRVEELES 184
Query: 240 LLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENVSKVYRDGG 295
LLKL+S+ + +V+GIWGM GIGKTTLA+VLYDRIS F+A CF+ENVSK+YRDGG
Sbjct: 185 LLKLSSDDDELRVVGIWGMAGIGKTTLASVLYDRISSQFDASCFIENVSKIYRDGG 352
>TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (98%)
Length = 1285
Score = 358 bits (918), Expect = 7e-99
Identities = 188/411 (45%), Positives = 262/411 (63%), Gaps = 3/411 (0%)
Frame = +3
Query: 48 RRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSR 107
RR YDVF++FRG DTRN F + L+A L RKGI+ F+DD L KGESI +LL+ I S+
Sbjct: 57 RRNCYDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGESIGPELLRTIEGSQ 236
Query: 108 VSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFH 167
V + V S+NYA S WCL E+ I EC + + V P+FY VDPS V+ Q+G+Y + F H
Sbjct: 237 VFVAVLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEVKKQSGIYWDDFAKH 416
Query: 168 MLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVIEALGRKFSGFADDL 227
RFK D +V RW+ A+ + AGWD+R+K + E+E IV+ ++ L K S + DL
Sbjct: 417 EQRFKQDPHKVSRWREALNQVGSIAGWDLRDKQQSVEVEKIVQTILNILKCKSSFVSKDL 596
Query: 228 IGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENV 287
+GI R E L++ L LNS +VIGIWGMGG GKTTLA LY +I H F+A CF+++V
Sbjct: 597 VGINSRTEALKHQLLLNS-VDGVRVIGIWGMGGKGKTTLAMNLYGQICHRFDASCFIDDV 773
Query: 288 SKVYR--DGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLV 345
SK++R DG + A QKQ+L QT+ + + + + ++R RL K L++LDNVDQ+
Sbjct: 774 SKIFRLHDGPIDA-QKQILHQTLGIEHHQICNHYSATDLIRHRLSREKTLLILDNVDQVE 950
Query: 346 QLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDN 405
QL+ V+ GSR++I +RDEHILK Y + Y+VPL++ ++ +LF +K FK +
Sbjct: 951 QLERIGVHREWLGAGSRIVIISRDEHILKEYKVDVDYKVPLLDWTESHKLFRQKAFKLEK 1130
Query: 406 -LSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPD 455
+ L E+L YA GLPL I V GSFL RN +W+ AL RL+ P+
Sbjct: 1131IIMKNYQNLAYEILNYANGLPLTITVLGSFLSGRNVTEWKSALARLRQRPN 1283
>TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (24%)
Length = 1769
Score = 285 bits (730), Expect = 4e-77
Identities = 175/526 (33%), Positives = 290/526 (54%), Gaps = 14/526 (2%)
Frame = +3
Query: 50 YKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDD--KKLQKGESISAQLLQAIRNSR 107
+ YDVF+ +G+DTR F +L L+ KGI F DD LQ+ + ++ +++ SR
Sbjct: 84 FSYDVFLICKGTDTRYGFTGNLLKALIDKGIRTFHDDDDSDLQRRDKVTPIIIE---ESR 254
Query: 108 VSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFH 167
+ I +FS NYA S CLD + I C + V PVF+ V+P+ VR+ G Y A H
Sbjct: 255 ILIPIFSANYASSSSCLDTLVHIIHCYKTKGCLVLPVFFGVEPTDVRHHTGRYGKALAEH 434
Query: 168 MLRFKHDA---DRVDRWKRAMRSLAGSAGW-DVRNKPEFREIENIVEAVIEALGRKFSGF 223
RF++D +R+ +WK A+ A + D + E+ I IV+ + + R+
Sbjct: 435 ENRFQNDTKNMERLQQWKVALSLAANLPSYHDDSHGYEYELIGKIVKYISNKISRQSLHV 614
Query: 224 ADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCF 283
A +G+Q RV+ +++LL + ++GI+G+GG GK+TLA +Y+ ++ FE CF
Sbjct: 615 ATYPVGLQSRVQQVKSLLDEGPDD-GVHMVGIYGIGGSGKSTLARAIYNFVADQFEGLCF 791
Query: 284 VENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQ 343
+E V + + Q+ +L +T+ ++ ++ SE I+++RL K+L++LD+VD
Sbjct: 792 LEQVRENSASNSLKRFQEMLLSKTL-QLKIKLADVSEGISIIKERLCRKKILLILDDVDN 968
Query: 344 LVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKS 403
+ QL A F GSR+IITTRD+H+L + Y V +N +A EL FK+
Sbjct: 969 MKQLNALAGGVDWFGPGSRVIITTRDKHLLACHEIEKTYAVKGLNVTEALELLRWMAFKN 1148
Query: 404 DNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQ 463
D + S +++ V+ YA GLP+ I + GS L +N + ++ LD + P+ ++ +L+
Sbjct: 1149DKVPSSYEKILNRVVAYASGLPVVIEIVGSNLFGKNIEECKNTLDWYEKIPNKEIQRILK 1328
Query: 464 ISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACGLHPHIG------IQNMIERSLIT 517
+S++ L E++ +FL IAC FKG K VK I LH H G ++ ++E+ LI
Sbjct: 1329VSYDSLEEEEQSVFLDIACCFKGCKWEKVKEI-----LHAHYGHCINHHVEVLVEKCLID 1493
Query: 518 IRNQEIH--MHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVL 561
+ H +H +++++GK++VR + P EPG SRLW + VL
Sbjct: 1494HFEYDSHVSLHNLIENMGKELVRLESPFEPGKRSRLWFEKDIFEVL 1631
>TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum
tuberosum}, partial (22%)
Length = 832
Score = 254 bits (648), Expect = 1e-67
Identities = 133/270 (49%), Positives = 178/270 (65%)
Frame = +1
Query: 22 SSHHLGPLAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIF 81
SS HL + S+ SSS V S+ ++ YDVF++FRG DTRN F D L+ L KGI
Sbjct: 7 SSKHLY-IQMASSSKSSSALVTSS--KKNHYDVFVTFRGEDTRNNFTDFLFDALETKGIM 177
Query: 82 VFKDDKKLQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTV 141
VF+D LQKGE I +L +AI S+V + +FSKNYA S WCL E+ I EC + + V
Sbjct: 178 VFRDVINLQKGECIGPELFRAIEISQVYVAIFSKNYASSTWCLQELEKICECIKGSGKHV 357
Query: 142 FPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPE 201
PVFYDVDPS VR Q+G+Y AFV H RF+ D+ +V RW+ A+ + +GWD+R++P
Sbjct: 358 LPVFYDVDPSEVRKQSGIYSEAFVKHEQRFQQDSMKVSRWREALEQVGSISGWDLRDEPL 537
Query: 202 FREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGI 261
REI+ IV+ +I L K+S + DL+GI ++ L+N L LNS + IGI GMGGI
Sbjct: 538 AREIKEIVQKIINILECKYSCVSKDLVGIDSPIQALQNHLLLNS-VDGVRAIGICGMGGI 714
Query: 262 GKTTLATVLYDRISHLFEARCFVENVSKVY 291
GKTTLAT LY +ISH F A CF+++V+K+Y
Sbjct: 715 GKTTLATTLYGQISHQFSASCFIDDVTKIY 804
>BI309457 weakly similar to GP|15787901|gb resistance gene analog NBS7
{Helianthus annuus}, partial (45%)
Length = 806
Score = 249 bits (637), Expect = 3e-66
Identities = 125/258 (48%), Positives = 172/258 (66%)
Frame = +3
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
++ S+S++V +RR YDVF++FRG DTRN F D L+ L KGI VF DD L KGE
Sbjct: 18 ASTSNSSSVLGTSSRRNYYDVFVTFRGEDTRNNFTDFLFDALQTKGIIVFSDDTNLPKGE 197
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SI +LL+AI S+V + VFS NYA S WCL E+ I EC + + V PVFYDVDPS V
Sbjct: 198 SIGPELLRAIEGSQVFVAVFSINYASSTWCLQELEKICECVKGSGKHVLPVFYDVDPSDV 377
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVI 213
R Q+G+Y AF+ H RF+ + +V +W+ A++ + +GWD+R+KP+ EI+ IV+ ++
Sbjct: 378 RKQSGIYGEAFIKHEQRFQQEFQKVSKWRDALKQVGSISGWDLRDKPQAGEIKKIVQTIL 557
Query: 214 EALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDR 273
L K S F+ DL+GI R++ L+N L L+S + IGI GMGGIGKTTLA LYD+
Sbjct: 558 NILKYKSSCFSKDLVGIDSRLDGLQNHLLLDS-VDSVRAIGICGMGGIGKTTLAMALYDQ 734
Query: 274 ISHLFEARCFVENVSKVY 291
ISH F A F+++V + Y
Sbjct: 735 ISHRFSASWFIDDVKQNY 788
>CA920659 similar to GP|21740675|em OSJNBa0084K20.18 {Oryza sativa}, partial
(2%)
Length = 790
Score = 237 bits (604), Expect = 2e-62
Identities = 127/201 (63%), Positives = 150/201 (74%), Gaps = 18/201 (8%)
Frame = +3
Query: 767 CVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLP 826
C SLST+ QSI VLT+L+FLSLRDC NL +IP S+N M SL+TLD CGCL+L++LPLG
Sbjct: 15 CSSLSTIGQSIAVLTQLKFLSLRDCTNLVSIPKSINCMTSLVTLDLCGCLRLENLPLGNV 194
Query: 827 SLSP----------------FTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFV 870
S+S + L+SLIFLDL FC+LS VP A+G++ LERLNLEGN+FV
Sbjct: 195 SVSAENMDYFSVSGEIVDHSYYLESLIFLDLSFCNLSTVPDAIGDLWHLERLNLEGNSFV 374
Query: 871 SLPSSLRWLSSLAYLNLAHCSKLEFLSELQLCDIASEGGRYFRTLSGSHNHRSGLYIFNC 930
SLP S+ LSSLAYLNLAHCS L+ L +LQLC +S GGRYF+ LSGSHNHRSGLYIFNC
Sbjct: 375 SLPPSMERLSSLAYLNLAHCSWLQSLPKLQLCATSSYGGRYFKMLSGSHNHRSGLYIFNC 554
Query: 931 PTLAIT--GLNLALLWLERLV 949
P L I G +LAL WLERLV
Sbjct: 555 PLLEIAEGGQSLALSWLERLV 617
>TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (37%)
Length = 2072
Score = 230 bits (586), Expect = 2e-60
Identities = 158/498 (31%), Positives = 266/498 (52%), Gaps = 12/498 (2%)
Frame = +3
Query: 198 NKPEFREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWG 257
N+ E+ I IVE + + R+ A +G+Q RV+ ++ L S+ + ++G++G
Sbjct: 684 NRYEYEFIGKIVEDISNRISREPLDVAKYPVGLQSRVQHVKGHLDEKSDD-EVHMVGLYG 860
Query: 258 MGGIGKTTLATVLYDRISHLFEARCFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYS 317
GGIGK+TLA +Y+ I+ FE CF+ENV + +Q+++L +TV ++++
Sbjct: 861 TGGIGKSTLAKAIYNFIADQFEVLCFLENVRVNSTSDNLKHLQEKLLLKTV-RLDIKLGG 1037
Query: 318 PSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYG 377
S+ I++ RL K+L++LD+VD+L QL+ A F GSR+IITTR++H+LK++G
Sbjct: 1038 VSQGIPIIKQRLCRKKILLILDDVDKLDQLEALAGGLDWFGPGSRVIITTRNKHLLKIHG 1217
Query: 378 AHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCT 437
+ V +N +A EL FK +N+ S +++ L YA GLPLAI + GS L
Sbjct: 1218 IESTHAVEGLNATEALELLRWMAFK-ENVPSSHEDILNRALTYASGLPLAIVIIGSNLVG 1394
Query: 438 RNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILD 497
R+ LD + P+ ++ +L++S++ L E++ +FL IAC FKG K VK IL
Sbjct: 1395 RSVQDSMSTLDGYEEIPNKEIQRILKVSYDSLEKEEQSVFLDIACCFKGCKWPEVKEILH 1574
Query: 498 A----CGLHPHIGIQNMIERSLITIRNQE--IHMHEMVQDLGKKIVRQQFPEEPGSWSRL 551
A C +H H+ + + E+SL+ + + +H++++D+GK++VRQ+ P+EPG SRL
Sbjct: 1575 AHYGHCIVH-HVAV--LAEKSLMDHLKYDSYVTLHDLIEDMGKEVVRQESPDEPGERSRL 1745
Query: 552 WLYQHFHHVLMSEMGTNKVKAIVLDQNEDISEYPQLRAE------GLSIMRGLIILILHH 605
W + HVL GT K+K I + ++P + ++ M L I +
Sbjct: 1746 WFERDIVHVLKKNTGTRKIKMINM-------KFPSMESDIDWNGNAFEKMTNLKTFITEN 1904
Query: 606 QNFSGSLHFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRKDLPFLK 665
+ S SL +L ++L+ + +P + P SS K +K
Sbjct: 1905 GHHSKSLEYLPSSLRVMK------GCIPKS-----------PSSS-----SSNKKFEDMK 2018
Query: 666 RMDLSNSKYLTETPNFEG 683
+ L+N +YLT P+ G
Sbjct: 2019 VLILNNCEYLTHIPDVSG 2072
Score = 127 bits (319), Expect = 2e-29
Identities = 65/147 (44%), Positives = 88/147 (59%), Gaps = 3/147 (2%)
Frame = +1
Query: 50 YKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVS 109
+ Y VF+SFRG+DTR+ F +LY L KGI+ F DD LQ+G+ I+ L AI SR+
Sbjct: 88 FTYQVFLSFRGADTRHGFTGNLYKALTDKGIYTFIDDNDLQRGDEITPSLKNAIEKSRIF 267
Query: 110 IVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHML 169
I VFS+NYA S +CLDE+ I C + V PVF VDP+ VR+ G Y A H
Sbjct: 268 IPVFSENYASSSFCLDELVHITHCYDTKGCLVLPVFIGVDPTDVRHHTGRYGEALAVHKK 447
Query: 170 RFKHDAD---RVDRWKRAMRSLAGSAG 193
+F++D D R+ +WK A+ A +G
Sbjct: 448 KFQNDKDNTERLQQWKEALSQAANLSG 528
>TC86801 similar to PIR|A54810|A54810 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (7%)
Length = 1268
Score = 218 bits (554), Expect = 1e-56
Identities = 134/361 (37%), Positives = 204/361 (56%), Gaps = 8/361 (2%)
Frame = +2
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ A + KYDVFISFRG DTR F HL+A L R + + D +++KG+ + +L +AI
Sbjct: 44 SHSAAQKKYDVFISFRGEDTRAGFTSHLHAALSRTYLHTYID-YRIEKGDEVWPELEKAI 220
Query: 104 RNSRVSIVVFSKNYAESRWCLDEMAAIAECC---EDFKQTVFPVFYDVDPSPVRNQNGVY 160
+ S + +VVFS+NYA S WCL+E+ + EC ED V PVFY VDPS VR Q G Y
Sbjct: 221 KQSTLFLVVFSENYASSTWCLNELVELMECRNKNEDDNIGVIPVFYHVDPSHVRKQTGSY 400
Query: 161 ENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAVIEAL-GRK 219
+A H D + WK A+ A +G+ + +R N++E + AL G+
Sbjct: 401 GSALAKHKQE-NQDDKMMQNWKNALFQAANLSGF---HSSTYRTESNMIEDITRALLGKL 568
Query: 220 FSGFADDL---IGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISH 276
+ D+L + + + +L+K +S Q+IG+WGMGG GKTTLA ++ R S
Sbjct: 569 NHQYRDELTCNLILDENYWAVRSLIKFDST--TVQIIGLWGMGGTGKTTLAAAMFQRFSF 742
Query: 277 LFEARCFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLV 336
+E CF+E V++V + G+ ++L + + E +L +P I +++ RLR +K +
Sbjct: 743 KYEGNCFLERVTEVSKKHGINYTCNKLLSKLLGE-DLRIDTPKVIPAMIKRRLRHMKSFI 919
Query: 337 VLDNVDQLVQLQE-FAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDAREL 395
VLD+V LQ+ V G GS +I+TTRD+H+L G +YEV MN+ ++ +L
Sbjct: 920 VLDDVHNSELLQDLIGVRGGWLGPGSIVIVTTRDKHVLISGGIDEIYEVKKMNSQNSLQL 1099
Query: 396 F 396
F
Sbjct: 1100F 1102
>TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (23%)
Length = 1651
Score = 205 bits (522), Expect = 6e-53
Identities = 124/342 (36%), Positives = 200/342 (58%), Gaps = 6/342 (1%)
Frame = +1
Query: 198 NKPEFREIENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWG 257
N+ E++ IE IVE + + F A +G+Q R+E ++ LL + SE + +++G++G
Sbjct: 493 NRYEYKFIEKIVEDISNNINHVFLNVAKYPVGLQSRIEQVKLLLDMGSED-EVRMVGLFG 669
Query: 258 MGGIGKTTLATVLYDRISHLFEARCFVENVSKVYRDGGVTAVQKQVLRQTVDEMNLETYS 317
GG+GK+TLA +Y+ ++ FE CF+ NV + + +QK++L + V + + +
Sbjct: 670 TGGMGKSTLAKAVYNFVADQFEGVCFLHNVRENSTHKNLKHLQKKLLSKIV-KFDGKLED 846
Query: 318 PSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYG 377
SE I+++RL K+L++LD+VD+L QL+ A F GSR+IITTRD+H+L +G
Sbjct: 847 VSEGIPIIKERLSRKKILLILDDVDKLEQLEALAGGLDWFGHGSRVIITTRDKHLLACHG 1026
Query: 378 AHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCT 437
+ V +N +A EL R FK+D + S E++ V+ YA GLPLAI G L
Sbjct: 1027ITSTHAVEELNETEALELLRRMAFKNDKVPSTYEEILNRVVTYASGLPLAIVTIGDNLFG 1206
Query: 438 RNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILD 497
R W+ LD +N P+ + +LQ+S++ L ++K +FL IAC FKG K VK+IL
Sbjct: 1207RKVEDWKRILDEYENIPNKDIQRILQVSYDALEPKEKSVFLDIACCFKGCKWTKVKKILH 1386
Query: 498 A----CGLHPHIGIQNMIERSLITIRNQEIHM--HEMVQDLG 533
A C H H+G+ + E+SLI + M H++++D+G
Sbjct: 1387AHYGHCIEH-HVGV--LAEKSLIGHWEYDTQMTLHDLIEDMG 1503
Score = 60.1 bits (144), Expect = 4e-09
Identities = 30/81 (37%), Positives = 47/81 (57%), Gaps = 3/81 (3%)
Frame = +2
Query: 117 YAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKH--- 173
YA S +CLDE+ I C + V PVFYDV+P+ +R+Q+G Y H RF++
Sbjct: 2 YASSSFCLDELVHIIHCYKTKSCLVLPVFYDVEPTHIRHQSGSYGEYLTKHEERFQNNEK 181
Query: 174 DADRVDRWKRAMRSLAGSAGW 194
+ +R+ +WK A+ A +G+
Sbjct: 182 NMERLRQWKIALTQAANLSGY 244
>TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 656
Score = 204 bits (519), Expect = 1e-52
Identities = 96/204 (47%), Positives = 134/204 (65%)
Frame = +2
Query: 29 LAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKK 88
+ S+ SSS+ + + R+ YDVF++FRG DTR F+DHL+A L RKGIF F+DD
Sbjct: 41 IQMASSSNSSSSALMALPRRKNYYDVFVTFRGEDTRFNFIDHLFAALQRKGIFAFRDDAN 220
Query: 89 LQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDV 148
LQKGESI +L++AI S+V I V SKNY+ S WCL E+ I +C + + V PVFYDV
Sbjct: 221 LQKGESIPPELIRAIEGSQVFIAVLSKNYSSSTWCLRELVHILDCSQVSGRRVLPVFYDV 400
Query: 149 DPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENI 208
DPS VR+Q G+Y AF H F+HD+ V W+ A+ + +GWD+R+KP++ EI+ I
Sbjct: 401 DPSEVRHQKGIYGEAFSKHEQTFQHDSHVVQSWREALTQVGNISGWDLRDKPQYAEIKKI 580
Query: 209 VEAVIEALGRKFSGFADDLIGIQP 232
VE ++ LG FS +L+G+ P
Sbjct: 581 VEEILNILGHNFSSLPKELVGMNP 652
>TC92554 similar to GP|9758876|dbj|BAB09430.1 disease resistance protein
{Arabidopsis thaliana}, partial (7%)
Length = 1043
Score = 133 bits (335), Expect(2) = 1e-49
Identities = 66/152 (43%), Positives = 96/152 (62%)
Frame = +1
Query: 16 LVSRRHSSHHLGPLAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHL 75
L S+H+ L G ASSS++ ++ + ++ VF++FRG DTR F HLYA L
Sbjct: 127 LAKESKSTHNWTCLLVGV*MASSSSSFGASHSNSKQHHVFLNFRGEDTRYNFTSHLYAAL 306
Query: 76 VRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCE 135
K I F DD+++++G++IS LL AI +S++ +++FS++YA S WCLDE+ I EC E
Sbjct: 307 CGKKIHTFMDDEEIERGDNISPTLLSAIESSKICVIIFSQDYASSSWCLDELVKIIECSE 486
Query: 136 DFKQTVFPVFYDVDPSPVRNQNGVYENAFVFH 167
K V PVFY +D S VR+Q G Y +AF H
Sbjct: 487 KKKLVVIPVFYHIDASHVRHQRGTYGDAFAKH 582
Score = 82.8 bits (203), Expect(2) = 1e-49
Identities = 49/154 (31%), Positives = 87/154 (55%), Gaps = 3/154 (1%)
Frame = +2
Query: 169 LRFKHDADRVDRWKRAMRSLAGSAGW-DVRNKPEFREIENIVEAVIEALGRKFSGFADD- 226
+RF+ + +V W+ A+ A AGW +N+ E I+ IVE +++ L + S +
Sbjct: 587 IRFRDNLTKVHMWRTALHKAANLAGWVSEKNRSEAVLIKEIVEVILDKL-KCMSPHVQNK 763
Query: 227 -LIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVE 285
L+GI V +E++L S D +IGIWGMGGIGKTT+A ++ ++S+ +E F
Sbjct: 764 GLVGISVHVAHVESMLCSGSA--DVHIIGIWGMGGIGKTTIADAVFTKVSYQYERYYFAA 937
Query: 286 NVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPS 319
NV + + G +Q +VL +++ +++ +P+
Sbjct: 938 NVRETW--GNRIKLQNEVLADVLEDQSIKISTPT 1033
>TC92115 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (24%)
Length = 1348
Score = 178 bits (452), Expect = 8e-45
Identities = 108/346 (31%), Positives = 193/346 (55%), Gaps = 10/346 (2%)
Frame = +2
Query: 299 VQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQ 358
+Q+++L + + ++ ++ SE +++RL + K L++LD+VD + QL A P F
Sbjct: 2 LQEELLLKAL-QLKIKLGGVSEGIPYIKERLHTKKTLLILDDVDDMEQLHALAGGPDWFG 178
Query: 359 KGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVL 418
+GSR+IITTRD+H+L+ +G +EV + +A EL FK++ + S +++ +
Sbjct: 179 RGSRVIITTRDKHLLRSHGIESTHEVEELYGTEALELLRWMAFKNNKVPSIYEDVLNRAV 358
Query: 419 KYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFL 478
YA GLPL + + GS L + +W+ LD + P+ K+ +L++S++ L E + +FL
Sbjct: 359 SYASGLPLVLEIVGSNLFGKTIEEWKGTLDGYEKIPNKKIHQILKVSYDALEEEQQSVFL 538
Query: 479 HIACFFKG---EKENYVKRILDACGLHPHIGIQNMIERSLITIRN------QEIHMHEMV 529
IAC FKG E+ Y+ R + H+ + + E+SL+ I + E+ +H+++
Sbjct: 539 DIACCFKGCGWEEFEYILRAHYGHRITHHLVV--LAEKSLVKITHPHYGSIYELTLHDLI 712
Query: 530 QDLGKKIVRQQFPEEPGSWSRLWLYQHFHHVLMSEMGTNKVKAIVLDQNEDISEYP-QLR 588
+++GK++VRQ+ P+EPG SRLW +VL GT+K++ I + N E+ +
Sbjct: 713 KEMGKEVVRQESPKEPGERSRLWCEDDIVNVLKENTGTSKIEMIYM--NFPSEEFVIDKK 886
Query: 589 AEGLSIMRGLIILILHHQNFSGSLHFLSNNLQYLLWHGYPFASLPS 634
+ M L LI+ + +FS L +L ++L+ L G SL S
Sbjct: 887 GKAFKKMTRLKTLIIENVHFSKGLKYLPSSLRVLKLRGCLSESLIS 1024
>TC92377 similar to GP|19774149|gb|AAL99051.1 NBS-LRR-Toll resistance gene
analog protein {Medicago sativa}, partial (57%)
Length = 945
Score = 160 bits (404), Expect(2) = 5e-44
Identities = 95/236 (40%), Positives = 135/236 (56%)
Frame = +1
Query: 251 QVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVENVSKVYRDGGVTAVQKQVLRQTVDE 310
QV+GI+GMGGIGKTTLAT LYDRISH F+A CF++++SKVYR+ G ++ + + +
Sbjct: 259 QVVGIYGMGGIGKTTLATALYDRISHQFDACCFIDDISKVYRNEGRLVLKSKHYIKLLAN 438
Query: 311 MNLETYSPSEISGIVRDRLRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDE 370
+ + ++R +L ++ L+VLDNVDQ+ Q ++ AV+ GSR++IT+RD
Sbjct: 439 STFRYAIFTTKTNLIRLKLGHIRALLVLDNVDQVEQQEKLAVSSECLGAGSRIVITSRDV 618
Query: 371 HILKVYGAHIVYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRV 430
HILK YEV + N F +K F+ + S EL + K
Sbjct: 619 HILK------EYEVDEIVN-----*FCQKAFQGGKIKSNYEELTCHI-KLC*WPTTGN*S 762
Query: 431 TGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKG 486
G LC RN +W AL RL+ NP+ +MDVL++SFEGL + IFL I C G
Sbjct: 763 IGLILCGRNISEWTSALARLRENPNKDIMDVLRLSFEGLEESENVIFLDIVCLLTG 930
Score = 37.4 bits (85), Expect(2) = 5e-44
Identities = 19/52 (36%), Positives = 25/52 (47%)
Frame = +3
Query: 205 IENIVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIW 256
IE IVE +I LG +FS DL+GI +E LE + Y +W
Sbjct: 123 IEKIVEEIINRLGYRFSSLPKDLVGIDSLIEELEKVFTFGIN*YYTGCRNLW 278
>TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (84%)
Length = 827
Score = 174 bits (442), Expect = 1e-43
Identities = 85/171 (49%), Positives = 114/171 (65%)
Frame = +2
Query: 29 LAQGGSTASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKK 88
L Q S+ +SS + ++ Y YDVF +FRG DTRN F D L+ L KGIF F+DD
Sbjct: 47 LNQMASSNNSSLALVTSSRNNY-YDVFGTFRGEDTRNNFTDFLFDALETKGIFAFRDDTN 223
Query: 89 LQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDV 148
LQKGESI +LL+AI SRV + VFS+NYA S WCL E+ I +C + ++ + PVFYDV
Sbjct: 224 LQKGESIEPELLRAIEGSRVFVAVFSRNYASSTWCLQELEKICKCVQRSRKHILPVFYDV 403
Query: 149 DPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNK 199
DPS VR Q+G+Y AFV H RF+ D + V RW+ A++ + +GWD+R+K
Sbjct: 404 DPSVVRKQSGIYCEAFVKHEQRFQQDFEMVSRWREALKHVGSISGWDLRDK 556
>TC85020 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (55%)
Length = 500
Score = 174 bits (440), Expect = 2e-43
Identities = 87/166 (52%), Positives = 117/166 (70%), Gaps = 1/166 (0%)
Frame = +1
Query: 333 KVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMNNNDA 392
K L++ DNVDQ+ QL++ AV+ GSR++I +RDEHILK YG +VY+VPLMN+ D+
Sbjct: 1 KALLIFDNVDQVEQLEKIAVHREWLGAGSRIVIISRDEHILKEYGVDVVYKVPLMNSTDS 180
Query: 393 RELFYRKGFKSDNL-SSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNAMQWRDALDRLK 451
ELF RK FK + + S L E+L YA+GLPLAI+V GSFL + +W+ AL RL+
Sbjct: 181 YELFCRKAFKVEKIIMSDYQNLANEILDYAKGLPLAIKVLGSFLFGHSVAEWKSALARLR 360
Query: 452 NNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILD 497
+P N VMDVL +SF+GL ++KEIFL IACFF + E YVK +L+
Sbjct: 361 ESPHNDVMDVLHLSFDGLDEKEKEIFLDIACFFNSQPEKYVKNVLN 498
>AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (10%)
Length = 644
Score = 173 bits (438), Expect = 3e-43
Identities = 84/176 (47%), Positives = 116/176 (65%)
Frame = +3
Query: 37 SSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESIS 96
+SS++ S R+ VF+SFRG DTR F DHL+A L R+GI FKDD L++GE IS
Sbjct: 75 ASSSSSPSIQTSRWTNHVFLSFRGEDTRQGFTDHLFASLERRGIKTFKDDHDLERGEVIS 254
Query: 97 AQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQ 156
+L +AI S +I++ S NYA S WCLDE+ I EC + F Q VFP+FY VDPS VR+Q
Sbjct: 255 YELNKAIEESMFAIIILSPNYASSTWCLDELKKIVECSKSFGQAVFPIFYGVDPSDVRHQ 434
Query: 157 NGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNKPEFREIENIVEAV 212
G ++ AF H +F+ D +V+RW+ A+R +AG +GWD + + E +E IVE +
Sbjct: 435 RGSFDEAFRKHEEKFRKDRTKVERWRDALREVAGYSGWDSKGRHEASLVETIVEHI 602
>TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (12%)
Length = 765
Score = 169 bits (428), Expect = 5e-42
Identities = 92/223 (41%), Positives = 140/223 (62%), Gaps = 4/223 (1%)
Frame = +1
Query: 54 VFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIVVF 113
VF+SFRGSDTRN F +LY LV KGI F DD L++G+ I+ L++AI SR+ I +F
Sbjct: 94 VFLSFRGSDTRNKFTGNLYKALVDKGIRTFIDDNDLERGDEITPSLVKAIEESRIFIPIF 273
Query: 114 SKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKH 173
S NYA S +CLDE+ I C + VFPVFYDV+P+ +RNQ+G+Y H RF++
Sbjct: 274 SANYASSSFCLDELVHIIHCYKTKSCLVFPVFYDVEPTHIRNQSGIYGEHLTKHEERFQN 453
Query: 174 ---DADRVDRWKRAMRSLAGSAGWDVR-NKPEFREIENIVEAVIEALGRKFSGFADDLIG 229
+ +R+ +WK A+ A +G+ + E++ IE IVE + + F A +G
Sbjct: 454 NEKNMERLRQWKIALIQAANLSGYHYSPHGYEYKFIEKIVEDISNNINHVFLNVAKYPVG 633
Query: 230 IQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYD 272
+Q R+E ++ LL + SE + +++G++G GG+GK+TLA +Y+
Sbjct: 634 LQSRIEEVKLLLDMGSE-DEVRMVGLFGTGGMGKSTLAKAVYN 759
>BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(9%)
Length = 655
Score = 168 bits (426), Expect = 8e-42
Identities = 79/166 (47%), Positives = 109/166 (65%)
Frame = +1
Query: 34 STASSSTTVFSNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGE 93
+++S+S+T RR YDVF++FRG DTRN F + L+A L RKGI+ F+DD L KGE
Sbjct: 16 ASSSNSSTALVPLPRRNCYDVFVTFRGEDTRNNFTNFLFAALERKGIYAFRDDTNLPKGE 195
Query: 94 SISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPV 153
SI +LL+ I S+V + V S+NYA S WCL E+ I EC + + V P+FY VDPS V
Sbjct: 196 SIGPELLRTIEGSQVFVAVLSRNYASSTWCLQELEKICECIKGSGKYVLPIFYGVDPSEV 375
Query: 154 RNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVRNK 199
+ Q+G+Y + F H RFK D +V RW+ A+ + AGWD+R+K
Sbjct: 376 KKQSGIYWDDFAKHEQRFKQDPHKVSRWREALNQVGSIAGWDLRDK 513
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,893,224
Number of Sequences: 36976
Number of extensions: 501382
Number of successful extensions: 3312
Number of sequences better than 10.0: 352
Number of HSP's better than 10.0 without gapping: 2933
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3093
length of query: 949
length of database: 9,014,727
effective HSP length: 105
effective length of query: 844
effective length of database: 5,132,247
effective search space: 4331616468
effective search space used: 4331616468
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0388b.1


