

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.4
(1489 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
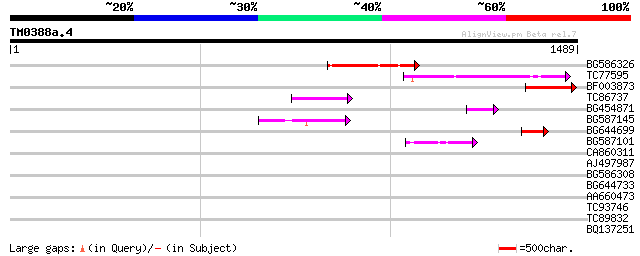
Score E
Sequences producing significant alignments: (bits) Value
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 217 2e-56
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 171 2e-42
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 168 1e-41
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 105 2e-22
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 67 7e-11
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 66 9e-11
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 64 6e-10
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 51 3e-06
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 42 0.002
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 38 0.026
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 37 0.044
BG644733 weakly similar to GP|15289942|db putative polyprotein {... 36 0.097
AA660473 34 0.37
TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical... 32 1.4
TC89832 homologue to GP|21618319|gb|AAM67369.1 unknown {Arabidop... 32 2.4
BQ137251 similar to GP|18026135|gb| cytochrome c oxidase subunit... 30 7.0
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 217 bits (553), Expect = 2e-56
Identities = 122/244 (50%), Positives = 161/244 (65%), Gaps = 2/244 (0%)
Frame = +2
Query: 834 TTALILALPKEGEPYDVYCDASHNGLGCVMMQEKKVIAYASRQLKIHEKNYPTHDLELAA 893
T+A IL LP E Y VY DAS GLGCV+ Q +KVIAYASRQL+ HE NYPTHDLE+AA
Sbjct: 8 TSAPILVLP-ELITYVVYTDASITGLGCVLTQHEKVIAYASRQLRKHEGNYPTHDLEMAA 184
Query: 894 IVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPGKT 953
+V+ALKIWR YLYG+ I++DHKSLKY+F Q +LN+RQRRWMEF+ DY+ D+ +PGK
Sbjct: 185 VVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYPGKA 364
Query: 954 NVVADALSRKAMHVSSLMVKELELLEAF-*DLSLDVKIAPGKLSFGMVTVS-SGLLDEIK 1011
N+VADALSR+ + VS+ +E + L+ L L+V + S G+ V+ + L I+
Sbjct: 365 NLVADALSRRRVDVSA--EREADDLDGMVRALRLNV-LTKATESLGLEAVNQADLFTRIR 535
Query: 1012 SKQETDEGLLEWRKLVTQGKAPEFSIGSDNILRCKGRVCVPNDATMRRLILDEGHKSRLS 1071
Q DE L + V Q E+ D + GR+ VPND +++ I+ E HKSR S
Sbjct: 536 LAQGQDENL----QKVAQNDRTEYQTAKDGTILVNGRISVPNDRSLKEEIMSEAHKSRFS 703
Query: 1072 IHPG 1075
+HPG
Sbjct: 704 VHPG 715
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 171 bits (433), Expect = 2e-42
Identities = 134/457 (29%), Positives = 215/457 (46%), Gaps = 17/457 (3%)
Frame = +2
Query: 1034 EFSIGSDNILRCKGRVCVPNDAT-------MRRLILDEGHKSRLSIHPGMNKMYQDLKLH 1086
E + S L +GR+ VP +R ++ E H S + HPG N + +
Sbjct: 77 ECQLDSLKRLTFRGRIWVPGSDDEESPLNELRTKLVQESHDSTAAGHPGRNGTLEIVSRK 256
Query: 1087 FWWPGMKKQVAEYVSTCLTC*NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRR 1146
F+WPG + V +V C C + Q G L+ L VP +SMDF+T+LP TR
Sbjct: 257 FFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGFLKPLPVPNRLHSDLSMDFITSLPPTRG 436
Query: 1147 RFDA-IWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSR 1205
R +WV+VDRL+K+ + + E A+ ++ R HG+P SIVSDR S + R
Sbjct: 437 RGSQYLWVIVDRLSKSVTLEEMD-TMEAEACAQRFLSCHYRFHGMPQSIVSDRGSNWVGR 613
Query: 1206 FWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTY 1265
FW + G LS++Y+PQTDG TER Q ++ +LRA V + +W +LLP ++
Sbjct: 614 FWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEIQAVLRAYVCWSQDNWGDLLPTVQLAL 793
Query: 1266 NNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPE-----LVQQTTEEVKRIQEKM 1320
N ++SIG P+ +G P+ +D +V E LV++ + IQ ++
Sbjct: 794 RNRHNSSIGATPFFVEHGYHVD-PIPTVEDTGGVVSEGEAAAQLLVKRMKDVTGFIQAEI 970
Query: 1321 RIS*SRQKSYADNRRKELE-FQAGDHVFLRVTPMTGVGRAIKSKKLTPKFIGPYQITERV 1379
+ R ++ A+ RR + +Q GD V+L V+ + K L K Y++T V
Sbjct: 971 VAAQQRSEASANKRRCPADRYQVGDKVWLNVSNYKSPRPSKKLDWLHHK----YEVTRFV 1138
Query: 1380 GPVAYRIALPPFLSHIHDVLHVSQLRKYMAD---DSHVLEPDDIQLKDDLTVVMPPIKIV 1436
P + +P ++ HV LR+ +D V++P + DD V ++ +
Sbjct: 1139 TPHVVELNVP---GTVYPKFHVDLLRRAASDPLPGQEVVDPQPPPIVDDDGEVEWEVEEI 1309
Query: 1437 DRSTKRLRNK*VSLVKVVWNQATGDATWELEDKMRES 1473
+ + +V + DATWE D +RE+
Sbjct: 1310 LAARWHQVGRGRRRQALVKWKGFVDATWEAADAIRET 1420
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 168 bits (426), Expect = 1e-41
Identities = 83/135 (61%), Positives = 106/135 (78%), Gaps = 1/135 (0%)
Frame = +2
Query: 1355 GVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHV 1414
GVGRA+KSKKLT +FIGPYQI+ERVG VAYR+ LPP L ++HDV HVSQLRKY+ D SHV
Sbjct: 2 GVGRALKSKKLTVRFIGPYQISERVGTVAYRVGLPPHLLNLHDVFHVSQLRKYVPDPSHV 181
Query: 1415 LEPDDIQLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWNQATGDA-TWELEDKMRES 1473
++ DD+Q++D+LTV P++I DR K LR K + LV+VVW++A G++ TWELE KM ES
Sbjct: 182 IQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWDRANGESLTWELESKMVES 361
Query: 1474 HPDLFVNP*VSRAKI 1488
+P+LF SR KI
Sbjct: 362 YPELFA*GKFSRTKI 406
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 105 bits (261), Expect = 2e-22
Identities = 58/166 (34%), Positives = 92/166 (54%), Gaps = 6/166 (3%)
Frame = +1
Query: 740 KCEFWMEEVKFLGHVISS-QGVAVDPSKIESIMSWEQPKIASDIRSFVGLAGYYRRFVKD 798
KCEF + VK++G ++++ +GV+ DP K+ +I W P RSF+G YY+ F+
Sbjct: 1030 KCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPG 1209
Query: 799 YAKLTSPLTQLTKKNQPFAWTEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNG 858
Y+++T PLT+LT+K+ PF W + E +F ++K+ +L + V D S
Sbjct: 1210 YSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFA 1389
Query: 859 LGCVMMQEKKV-----IAYASRQLKIHEKNYPTHDLELAAIVYALK 899
LG V+ QE +A+ S++L E NYP HD EL A+ L+
Sbjct: 1390 LGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAVWACLR 1527
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 66.6 bits (161), Expect = 7e-11
Identities = 34/85 (40%), Positives = 47/85 (55%)
Frame = +2
Query: 1200 SRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLP 1259
S S FW L + GT L +SSAY+P +DGQ+E + E LR + W + P
Sbjct: 20 SSLYSNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFP 199
Query: 1260 LIEFTYNNSFHASIGMAPYEALYGR 1284
E+ YN S++ S M P++ALYGR
Sbjct: 200 WAEYWYNTSYNISAAMTPFKALYGR 274
Score = 33.1 bits (74), Expect = 0.82
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Frame = +1
Query: 1328 KSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK--SKKLTPKFIGPYQITERVGPVAYR 1385
K AD +R+ EFQ G+HV +++ P A++ K +P F + A+
Sbjct: 403 KHQADKKRRHFEFQLGEHVLVKLQPYQQSSVALRKYQKFGSPNFGSLLTVCSL*VESAFH 582
Query: 1386 IALPPFLS 1393
PP+LS
Sbjct: 583 CKSPPYLS 606
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 66.2 bits (160), Expect = 9e-11
Identities = 68/261 (26%), Positives = 108/261 (41%), Gaps = 19/261 (7%)
Frame = +3
Query: 654 RLTSRRPHSELDMGIVSTS*CRLELPMHPQYLWTT*TVCSDHFWTSSWWCSLTTF*FTRR 713
R+ RR S L G +T *CRL L + + CS W + W + TT * +
Sbjct: 9 RMIWRRQRSSLIEGRTATK*CRLVLRTPARLTKDSSIECSQTNWGTRWRSTSTTC**SHS 188
Query: 714 VKKNTKSICVKC*EYCKGKELYANGSKCEFWME----EVKFLGHVISSQGVAVD---PSK 766
V++ +I S + W ++ + S Q D PSK
Sbjct: 189 VRRTI*TI*K---------------SDLKRWTNT**NSIRPNAPLASPQANFWDTSSPSK 323
Query: 767 IES-IMSWEQP-------KIASDIRSFVGLAGYYRRFVKDYAKLTSPLTQLTKKNQPFAW 818
I+S P +IA G RF+ P +L N+ F W
Sbjct: 324 ESR*ILSRSPPY*TSLVQRIAERSSDSRGRIAALNRFISRSTDKCLPFYKLLCGNKRFVW 503
Query: 819 TEKCEESFQEMKKRLTTALILALPKEGEPYDVYCDASHNGLGCVMMQ----EKKVIAYAS 874
EKCEE+F+++K+ LTT +L+ P+ G+ +Y S + V+++ E+K I Y S
Sbjct: 504 DEKCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIAISSTAVSSVLIREDRGEQKPIFYTS 683
Query: 875 RQLKIHEKNYPTHDLELAAIV 895
+++ E YPT + A++
Sbjct: 684 KRMTDPETRYPTLEKMAFAVI 746
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 63.5 bits (153), Expect = 6e-10
Identities = 30/73 (41%), Positives = 48/73 (65%), Gaps = 1/73 (1%)
Frame = +2
Query: 1344 DHVFLRVTPMT-GVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVS 1402
+ V L+V P G R K KL+ ++IGP+++ +R+G VAY +ALPP LS +H V HVS
Sbjct: 2 EQVLLKVLPTERGDCRFGKRGKLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVS 181
Query: 1403 QLRKYMADDSHVL 1415
++Y D ++++
Sbjct: 182 MFKRYHGDGNYII 220
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 51.2 bits (121), Expect = 3e-06
Identities = 48/194 (24%), Positives = 86/194 (43%), Gaps = 5/194 (2%)
Frame = +2
Query: 1039 SDNI-LRCKGRVCVPNDATMRRLILDEGHKSRLSIHPGMNKMYQDLK-LHFWWPGMKKQV 1096
+DNI +RC +P IL H S + H ++K ++ FWWP M K
Sbjct: 56 ADNIYIRCVAEEEIPG-------ILFHCHGSNYAGHFAVSKTVSKIQQAGFWWPTMFKDA 214
Query: 1097 AEYVSTCLTC*---NAKVEHQKPAGKLQSLDVPEWKWDSISMDFVTALPLTRRRFDAIWV 1153
++S C C N ++ P + ++V +D +DF+ P + I V
Sbjct: 215 HSFISKCDPCQRQGNIS*RNEMPQNFILEVEV----FDVWGIDFMGPFPSSYNN-KYILV 379
Query: 1154 VVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQA 1213
VD ++K + N + ++ + I GVP ++SD S F ++ + L +
Sbjct: 380 AVDYVSKWVEAIASPTN-DATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLLKK 556
Query: 1214 LGTKLRLSSAYNPQ 1227
G + ++++AY+PQ
Sbjct: 557 NGVRHKVATAYHPQ 598
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 41.6 bits (96), Expect = 0.002
Identities = 19/43 (44%), Positives = 28/43 (64%)
Frame = +1
Query: 870 IAYASRQLKIHEKNYPTHDLELAAIVYALKIWRHYLYGSMFTI 912
IAYASR L E+NY + E A ++A++ +RHYL+G F +
Sbjct: 64 IAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGPKFEL 192
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 38.1 bits (87), Expect = 0.026
Identities = 20/74 (27%), Positives = 38/74 (51%), Gaps = 1/74 (1%)
Frame = -2
Query: 893 AIVYALKIWRHYLYGSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQDYEFDLQ*HPG- 951
A+ +A K RHY+ + S +KY+F++ L R RW L +Y+ + +
Sbjct: 623 ALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRSQKAI 444
Query: 952 KTNVVADALSRKAM 965
K +++AD L+ + +
Sbjct: 443 KGSILADHLAHQPL 402
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 37.4 bits (85), Expect = 0.044
Identities = 24/71 (33%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = -2
Query: 1188 HGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACV 1247
HG+P IV+D S F S + + +L +S PQ++GQ E + + + D L+ +
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRL 506
Query: 1248 LDHKGSW-DEL 1257
KG W DEL
Sbjct: 505 DLKKGCWADEL 473
>BG644733 weakly similar to GP|15289942|db putative polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 174
Score = 36.2 bits (82), Expect = 0.097
Identities = 17/43 (39%), Positives = 28/43 (64%)
Frame = -3
Query: 1146 RRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLH 1188
+ +++I VVVDRLTK+ F+P +Y + A I + +IV +H
Sbjct: 130 KSYESI*VVVDRLTKSTLFIPFKTSYSAK*YARILLDEIVCIH 2
>AA660473
Length = 655
Score = 34.3 bits (77), Expect = 0.37
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = -3
Query: 449 WGLKSSM*FWEWIGWRSITFSW 470
W +S + FWEW+GW+S+ W
Sbjct: 305 WN*ESLICFWEWLGWKSLAIRW 240
>TC93746 weakly similar to GP|22830935|dbj|BAC15800. hypothetical
protein~similar to gag-pol polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (4%)
Length = 1019
Score = 32.3 bits (72), Expect = 1.4
Identities = 14/41 (34%), Positives = 26/41 (63%)
Frame = +2
Query: 1125 VPEWKWDSISMDFVTALPLTRRRFDAIWVVVDRLTKTAHFV 1165
VP+ W+ +++DF L T++ D+ VV D+ ++ AHF+
Sbjct: 476 VPKPPWEDVTIDFSLGLL*TQQLKDSKMVVGDKFSRMAHFI 598
>TC89832 homologue to GP|21618319|gb|AAM67369.1 unknown {Arabidopsis
thaliana}, partial (62%)
Length = 1347
Score = 31.6 bits (70), Expect = 2.4
Identities = 17/49 (34%), Positives = 24/49 (48%)
Frame = +1
Query: 1186 RLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTER 1234
+++G PS I R FWG++ LG+ R Y+P TDG R
Sbjct: 727 QIYGPPSKINKARQRTNVQVFWGSVH-GLGSFCRRCYCYSPDTDGIARR 870
>BQ137251 similar to GP|18026135|gb| cytochrome c oxidase subunit II
{Sphaerospira fraseri}, partial (12%)
Length = 1073
Score = 30.0 bits (66), Expect = 7.0
Identities = 21/63 (33%), Positives = 28/63 (44%)
Frame = -3
Query: 1219 RLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPLIEFTYNNSFHASIGMAPY 1278
RLS + P D R + L LLR SW + PL+EF ++ A G AP
Sbjct: 300 RLSCPFRPPPDTGL-RNLSFLPPLLRNFCFSVSLSWRQPSPLVEFRWSPRCAARAGRAPV 124
Query: 1279 EAL 1281
+L
Sbjct: 123 VSL 115
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.342 0.148 0.511
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,733,986
Number of Sequences: 36976
Number of extensions: 758454
Number of successful extensions: 5152
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2752
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 2272
Number of HSP's gapped (non-prelim): 3160
length of query: 1489
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1380
effective length of database: 4,984,343
effective search space: 6878393340
effective search space used: 6878393340
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0388a.4


