

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
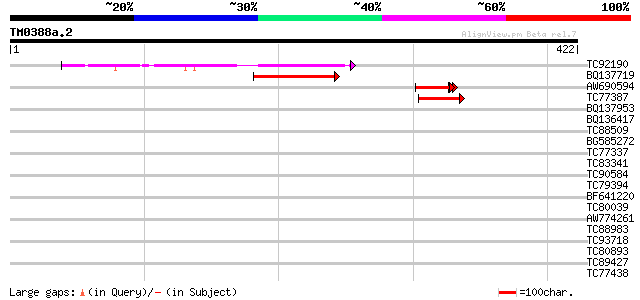
Query= TM0388a.2
(422 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92190 50 2e-06
BQ137719 45 7e-05
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 43 2e-04
TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19 ... 41 8e-04
BQ137953 weakly similar to GP|22136724|gb unknown protein {Arabi... 40 0.002
BQ136417 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 40 0.002
TC88509 similar to PIR|T01826|T01826 microfibril-associated prot... 40 0.002
BG585272 39 0.005
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 39 0.005
TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A... 39 0.005
TC90584 similar to GP|22136810|gb|AAM91749.1 unknown protein {Ar... 37 0.011
TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding... 37 0.019
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 36 0.032
TC80039 homologue to GP|6437558|gb|AAF08585.1| putative ATPase (... 36 0.032
AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis ... 35 0.054
TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protei... 35 0.054
TC93718 weakly similar to GP|10178129|dbj|BAB11541. gene_id:MOP1... 35 0.071
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 35 0.071
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 35 0.071
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 34 0.093
>TC92190
Length = 818
Score = 50.1 bits (118), Expect = 2e-06
Identities = 58/227 (25%), Positives = 95/227 (41%), Gaps = 8/227 (3%)
Frame = +2
Query: 39 HLAHKECVKERGVLYREGRRDLLGMQEAMEIRRWKSWVN--PIPFACEVMVREFYANTYC 96
+ A + + ER + E D+ + +E R W+ V A +V EFY N YC
Sbjct: 158 YFARRNIIHERVLSIPED--DMPQVPNQIEKRGWEYLVTYPENKEAYAELVVEFYCNAYC 331
Query: 97 DNQEDKSRPPVNSSWFRGDVIDYNPAVIRRHLG---LLTEDQE---KELFPEKATYHELL 150
Q+DK + S+ RG I ++ I L L T+DQ K+ + EL
Sbjct: 332 P-QKDKQK---EKSFVRGKQIPFDADTINSLLKSKLLRTKDQWPTFKQNVNTVSLLRELC 499
Query: 151 KEKSPEILEDMKAVIVRPGRDWEAVDDEIIFVRRKDMTPLARVWAEFLQDSLIPSSNKYE 210
K+++ IL+ P + PL ++WA ++ +L+P SN +
Sbjct: 500 KDETEWILDSTGKPKKFPS---------------SSLVPLPKIWAAYIGTNLVPCSNVSD 634
Query: 211 VREVTLVAIYCIVRGHPMDVATIIAGRIYSLYNRSKDKHRPIFPHLI 257
++ I I+ G +DV II +I + N + + FP LI
Sbjct: 635 IKVEKAALISAIMTGKVIDVGKIIQTQIQIIANSATNALE--FPRLI 769
>BQ137719
Length = 469
Score = 44.7 bits (104), Expect = 7e-05
Identities = 21/64 (32%), Positives = 39/64 (60%)
Frame = +2
Query: 182 VRRKDMTPLARVWAEFLQDSLIPSSNKYEVREVTLVAIYCIVRGHPMDVATIIAGRIYSL 241
+R D+TPLA+VWA F+ +L+P SN ++ + + I++G P++V ++A ++
Sbjct: 14 LRTSDLTPLAKVWAMFVLRTLLPCSNVSDLTILKANLLTAILKGEPVNVGRLLADDLWVT 193
Query: 242 YNRS 245
N S
Sbjct: 194 ANCS 205
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/28 (67%), Positives = 24/28 (84%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EE +D+EEEEEEEEEEE
Sbjct: 132 EEEEEEEEEEEDEHDDEEEEEEEEEEEE 215
Score = 42.4 bits (98), Expect = 3e-04
Identities = 20/31 (64%), Positives = 23/31 (73%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE D EEEEEEEEEEE E
Sbjct: 129 EEEEEEEEEEEEDEHDDEEEEEEEEEEEEEE 221
Score = 41.6 bits (96), Expect = 6e-04
Identities = 18/29 (62%), Positives = 25/29 (86%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E+EEEE+ +E E++EEEEEEEEEEE+
Sbjct: 138 EEEEEEEEEDEHDDEEEEEEEEEEEEEED 224
Score = 40.8 bits (94), Expect = 0.001
Identities = 18/29 (62%), Positives = 23/29 (79%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+E+E+E D EE D+EEEEEEEEEEE+
Sbjct: 81 EEEEDEHDEEEEDEHDEEEEEEEEEEEED 167
Score = 40.8 bits (94), Expect = 0.001
Identities = 18/31 (58%), Positives = 23/31 (74%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ DEEEE+ EE ED+ ++EEEEEEEE E
Sbjct: 120 EHDEEEEEEEEEEEEDEHDDEEEEEEEEEEE 212
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/46 (41%), Positives = 29/46 (62%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEED + E++EEEEEEEEE+++ E + + P P
Sbjct: 144 EEEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEEDEIDRISTLPDP 281
Score = 40.4 bits (93), Expect = 0.001
Identities = 18/31 (58%), Positives = 25/31 (80%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ E E++EEEEEEEEEE++ E
Sbjct: 141 EEEEEEEEDEHDDEEEEEEEEEEEEEEDDDE 233
Score = 39.3 bits (90), Expect = 0.003
Identities = 22/47 (46%), Positives = 26/47 (54%)
Frame = +3
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
K R + + E E+EE+E EE E EEEEEEEEEEE E
Sbjct: 30 KRRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDE 170
Score = 39.3 bits (90), Expect = 0.003
Identities = 16/29 (55%), Positives = 24/29 (82%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+EDE +E+ E+ E++EEEEEEEEE+E+
Sbjct: 87 EEDEHDEEEEDEHDEEEEEEEEEEEEDEH 173
Score = 37.4 bits (85), Expect = 0.011
Identities = 16/35 (45%), Positives = 24/35 (67%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+EDE +E+ EE E++E+E ++EEEEE E E
Sbjct: 111 EEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEE 215
Score = 37.4 bits (85), Expect = 0.011
Identities = 16/35 (45%), Positives = 24/35 (67%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E+E+E D EE E++EEE+E ++EEE E E
Sbjct: 105 EEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEE 209
Score = 37.0 bits (84), Expect = 0.014
Identities = 16/35 (45%), Positives = 23/35 (65%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+ DEEEED + E++EEEEEE+E ++ E E
Sbjct: 96 EHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEE 200
Score = 36.2 bits (82), Expect = 0.024
Identities = 17/34 (50%), Positives = 22/34 (64%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E +EEE+ EE E+ E ++EEEEEEE E E
Sbjct: 120 EHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEE 221
Score = 35.8 bits (81), Expect = 0.032
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = +3
Query: 297 WTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
W R +E EEED E E+ E +EEEE+E + E E
Sbjct: 27 WKRRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEE 149
Score = 34.3 bits (77), Expect = 0.093
Identities = 14/34 (41%), Positives = 24/34 (70%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++EEE++ +E E++EEEEE+E ++E E E
Sbjct: 102 DEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEE 203
>TC77387 similar to GP|15983396|gb|AAL11566.1 At1g51440/F5D21_19
{Arabidopsis thaliana}, partial (75%)
Length = 1793
Score = 41.2 bits (95), Expect = 8e-04
Identities = 19/34 (55%), Positives = 26/34 (75%)
Frame = +3
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
+EEEE+ EE E++EEEEEEEEEE + ++EV
Sbjct: 324 EEEEEEEEEDEDEEEEEEEEEEEEESPLPPLSEV 425
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/34 (55%), Positives = 26/34 (75%)
Frame = +3
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
++ M Q EEEE+ EE +++EEEEEEEEEEE+
Sbjct: 300 SSTMNQLHEEEEEEEEEDEDEEEEEEEEEEEEES 401
>BQ137953 weakly similar to GP|22136724|gb unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 657
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/60 (35%), Positives = 35/60 (58%)
Frame = +2
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
P L +R +L +E + P ++ E+E+ EE + E++EEEEEEE E+ + + E V
Sbjct: 182 PTTSYLEDKRKMKLQEEEKMK-PSDEAEKEEEEEQSSEEEEEEEEEESPVEDAKPIGEPV 358
>BQ136417 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (20%)
Length = 621
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/60 (35%), Positives = 35/60 (58%)
Frame = +2
Query: 280 PVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
P L +R +L +E + P ++ E+E+ EE + E++EEEEEEE E+ + + E V
Sbjct: 182 PTTSYLEDKRKMKLQEEEKMK-PSDEAEKEEEEEQSSEEEEEEEEEESPVEDAKPIGEPV 358
>TC88509 similar to PIR|T01826|T01826 microfibril-associated protein homolog
T15F16.8 - Arabidopsis thaliana, partial (83%)
Length = 1306
Score = 39.7 bits (91), Expect = 0.002
Identities = 21/55 (38%), Positives = 32/55 (58%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
KE++ Q +E +PQE+EEEE+ EE E+ + E + +EE V +V V P
Sbjct: 457 KEKLRQRDQE--EALPQEEEEEEEEEEEEEEESDYETDSDEEYTGVAMVKPVFVP 615
Score = 36.2 bits (82), Expect = 0.024
Identities = 16/43 (37%), Positives = 28/43 (64%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEE 329
++ +A+ + ++ Q D+EE P+E E++EEEEEEEE +
Sbjct: 424 EDAMAERRRRIKEKLRQRDQEEALPQEEEEEEEEEEEEEEESD 552
Score = 33.9 bits (76), Expect = 0.12
Identities = 16/31 (51%), Positives = 21/31 (67%)
Frame = +1
Query: 300 RMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
R +E + D EE +++EEEEEEEEEEE
Sbjct: 448 RRIKEKLRQRDQEEALPQEEEEEEEEEEEEE 540
>BG585272
Length = 827
Score = 38.5 bits (88), Expect = 0.005
Identities = 41/140 (29%), Positives = 65/140 (46%), Gaps = 11/140 (7%)
Frame = +1
Query: 1 MPVKIARTNTRAA-------ASSSTSR---SFDRSRFLSAIKEDFYRAHLAHKECVKERG 50
M K T++RAA ASSS SR FD RF ++ Y+ L ++
Sbjct: 184 MGPKKTNTSSRAASKKQKTVASSSRSRVQEPFDSLRFKGPYQQQRYKDLLERTFWSEKVF 363
Query: 51 VLYREGRRDLLGMQEAMEIRRWKSWVNPIPFACEVMVREFYANTY-CDNQEDKSRPPVNS 109
+ + G+ G+ + + R+W+ VNP ++REFYAN+ D ED S +
Sbjct: 364 QIKKNGQ--YRGIAQIICTRKWEILVNPPTLLNYDIIREFYANSIPTDPDEDFS----FT 525
Query: 110 SWFRGDVIDYNPAVIRRHLG 129
++ RG I ++ I +LG
Sbjct: 526 TFVRGRTIRFDRNAINAYLG 585
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 38.5 bits (88), Expect = 0.005
Identities = 17/42 (40%), Positives = 28/42 (66%), Gaps = 2/42 (4%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEE--EEEEEEEENVEVVNEVVTPPR 343
+++E++D EE GE+ EEE +EE+ EEE + E + PP+
Sbjct: 639 DEDEDDDDEEEGGEEDEEEGVDEEDNEEEEEDEDEEALQPPK 764
Score = 34.7 bits (78), Expect = 0.071
Identities = 15/31 (48%), Positives = 22/31 (70%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+ EED EE E+ EEEEE+E+EE ++
Sbjct: 663 EEEGGEEDEEEGVDEEDNEEEEEDEDEEALQ 755
Score = 34.3 bits (77), Expect = 0.093
Identities = 14/29 (48%), Positives = 21/29 (72%)
Frame = +3
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
D+++ED +E +D EEE EE+EEE V+
Sbjct: 618 DDQDEDDDEDEDDDDEEEGGEEDEEEGVD 704
Score = 32.3 bits (72), Expect = 0.35
Identities = 15/34 (44%), Positives = 23/34 (67%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+D++E+D E+ +D+EE EE+EEE E NE
Sbjct: 618 DDQDEDDDEDEDDDDEEEGGEEDEEEGVDEEDNE 719
Score = 31.6 bits (70), Expect = 0.60
Identities = 12/31 (38%), Positives = 23/31 (73%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
++DE+ +D +E ED+++++EEE EE+ E
Sbjct: 600 EDDEDGDDQDEDDDEDEDDDDEEEGGEEDEE 692
Score = 30.8 bits (68), Expect = 1.0
Identities = 13/36 (36%), Positives = 24/36 (66%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
E+E++ED ++ +D E+E++++EEE E E V
Sbjct: 594 EEEDDEDGDDQDEDDDEDEDDDDEEEGGEEDEEEGV 701
Score = 29.6 bits (65), Expect = 2.3
Identities = 13/37 (35%), Positives = 26/37 (70%), Gaps = 2/37 (5%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEE--EEENVEVVNE 337
+ED+E+ D ++ ++ E++++EEE EE+ E V+E
Sbjct: 597 EEDDEDGDDQDEDDDEDEDDDDEEEGGEEDEEEGVDE 707
>TC83341 weakly similar to GP|8978076|dbj|BAA98104.1 gene_id:K14A3.4~unknown
protein {Arabidopsis thaliana}, partial (80%)
Length = 1253
Score = 38.5 bits (88), Expect = 0.005
Identities = 20/55 (36%), Positives = 31/55 (56%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
+ERI EW ++EEEE+ EE +EEEEE++++ E E V ++ P
Sbjct: 532 QERIGVPRSEWIGNEGLQEEEEEEEEE-----EEEEEEDDDDVEKDETVEQIHVP 681
>TC90584 similar to GP|22136810|gb|AAM91749.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 686
Score = 37.4 bits (85), Expect = 0.011
Identities = 21/43 (48%), Positives = 28/43 (64%), Gaps = 3/43 (6%)
Frame = -2
Query: 299 ARMPQEDEEEEDP-EEPAGEDKEEEEEE--EEEEENVEVVNEV 338
A +P +EE +P EE A ED+EEEE+E EEEE+ E V +
Sbjct: 217 APLPVSEEESAEPLEEEAAEDEEEEEDEAAEEEEDEEEEVGPI 89
Score = 29.6 bits (65), Expect = 2.3
Identities = 22/48 (45%), Positives = 30/48 (61%), Gaps = 1/48 (2%)
Frame = -2
Query: 281 VAPL-LSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEE 327
+APL +S+E A+ +E A EDEEEE+ E +EEE+EEEE
Sbjct: 220 LAPLPVSEEESAEPLEEEAA----EDEEEEEDEAA----EEEEDEEEE 101
>TC79394 weakly similar to PIR|T49019|T49019 probable RNA binding protein -
Arabidopsis thaliana, partial (74%)
Length = 1909
Score = 36.6 bits (83), Expect = 0.019
Identities = 18/31 (58%), Positives = 22/31 (70%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ EEE + EE E++ EEEEEEEEEE VE
Sbjct: 266 ESSEEEVEYEEVEVEEEVEEEEEEEEEEEVE 358
Score = 35.8 bits (81), Expect = 0.032
Identities = 19/35 (54%), Positives = 22/35 (62%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
QE+ EE+ E E +EE EEEEEEEE EV E
Sbjct: 260 QEESSEEEVEYEEVEVEEEVEEEEEEEEEEEVEEE 364
Score = 32.7 bits (73), Expect = 0.27
Identities = 20/39 (51%), Positives = 22/39 (56%), Gaps = 2/39 (5%)
Frame = +2
Query: 305 DEEEEDPEEPAGEDKEEEEE--EEEEEENVEVVNEVVTP 341
D+EE EE E+ E EEE EEEEEE E V E P
Sbjct: 257 DQEESSEEEVEYEEVEVEEEVEEEEEEEEEEEVEEESKP 373
Score = 32.0 bits (71), Expect = 0.46
Identities = 17/35 (48%), Positives = 23/35 (65%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E E EE E E++EEEEEEEE EE + ++E
Sbjct: 278 EEVEYEEVEVEEEVEEEEEEEEEEEVEEESKPLDE 382
Score = 32.0 bits (71), Expect = 0.46
Identities = 22/54 (40%), Positives = 27/54 (49%), Gaps = 10/54 (18%)
Frame = +2
Query: 295 KEWTARMPQEDEEEEDPEEPA--GE--------DKEEEEEEEEEEENVEVVNEV 338
K+ TAR + +PE+P GE D+EE EEE E E VEV EV
Sbjct: 158 KDATARNSESPGGNSEPEKPTDSGEQVDLDGDNDQEESSEEEVEYEEVEVEEEV 319
Score = 30.0 bits (66), Expect = 1.8
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEV 338
++++EE EE ++ E EEE EEEE E EV
Sbjct: 251 DNDQEESSEEEVEYEEVEVEEEVEEEEEEEEEEEV 355
Score = 29.6 bits (65), Expect = 2.3
Identities = 14/44 (31%), Positives = 28/44 (62%), Gaps = 2/44 (4%)
Frame = +2
Query: 294 YKEWTARMPQEDEEEEDPEEPAGEDKE--EEEEEEEEEENVEVV 335
Y+E E+EEEE+ EE E+ + +EE+E +++++ E++
Sbjct: 290 YEEVEVEEEVEEEEEEEEEEEVEEESKPLDEEDEADKKKHAELL 421
Score = 28.9 bits (63), Expect = 3.9
Identities = 16/35 (45%), Positives = 24/35 (67%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E E EE+ EE E++EEEEE EEE + ++ +E
Sbjct: 293 EEVEVEEEVEEE--EEEEEEEEVEEESKPLDEEDE 391
Score = 27.7 bits (60), Expect = 8.7
Identities = 12/38 (31%), Positives = 22/38 (57%)
Frame = +2
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPP 342
+EEEE+ EE E++ + +EE+E + + + PP
Sbjct: 320 EEEEEEEEEEEVEEESKPLDEEDEADKKKHAELLALPP 433
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 35.8 bits (81), Expect = 0.032
Identities = 14/29 (48%), Positives = 22/29 (75%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENV 332
EDEE+++ ++ GED E+EE E+E+E V
Sbjct: 386 EDEEDDEEDDDEGEDDEDEEXEDEDEXXV 472
Score = 34.3 bits (77), Expect = 0.093
Identities = 13/34 (38%), Positives = 25/34 (73%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
ED+EEED E+ ++++++E E++E+E E +E
Sbjct: 362 EDDEEEDDEDEEDDEEDDDEGEDDEDEEXEDEDE 463
Score = 33.5 bits (75), Expect = 0.16
Identities = 15/35 (42%), Positives = 26/35 (73%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++DEEE+D +E ED EE+++E E++E+ E +E
Sbjct: 362 EDDEEEDDEDE---EDDEEDDDEGEDDEDEEXEDE 457
>TC80039 homologue to GP|6437558|gb|AAF08585.1| putative ATPase (ISW2-like)
{Arabidopsis thaliana}, partial (47%)
Length = 1840
Score = 35.8 bits (81), Expect = 0.032
Identities = 24/68 (35%), Positives = 40/68 (58%), Gaps = 4/68 (5%)
Frame = +3
Query: 271 PVHVI--EKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDK--EEEEEEE 326
P+H++ +KRL + SK+R + E AR D+E+ P+EPA +D +EE+E +
Sbjct: 114 PIHLLPKKKRLRT*TIRSKKRTKKKV-EAVARSAGSDDEDA-PDEPAADDDGVDEEDENQ 287
Query: 327 EEEENVEV 334
+E N E+
Sbjct: 288 DESVNPEI 311
>AW774261 weakly similar to EGAD|146423|156 vitellogenin {Anolis pulchellus},
partial (40%)
Length = 495
Score = 35.0 bits (79), Expect = 0.054
Identities = 17/28 (60%), Positives = 19/28 (67%)
Frame = +1
Query: 306 EEEEDPEEPAGEDKEEEEEEEEEEENVE 333
EEE DPEE E + EE EEEEE E +E
Sbjct: 298 EEENDPEEEMEEIEYEEVEEEEEVEEIE 381
Score = 33.9 bits (76), Expect = 0.12
Identities = 19/37 (51%), Positives = 23/37 (61%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
+E EEEE+ EE E +E EE+ EEEEE E E V
Sbjct: 343 EEVEEEEEVEEIEEEVEENEEDAEEEEEE*EKEEEEV 453
Score = 33.5 bits (75), Expect = 0.16
Identities = 18/47 (38%), Positives = 28/47 (59%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E + ++ E + +E EE+ EE + +EEEEE E+EEE VE
Sbjct: 316 EEEMEEIEYEEVEEEEEVEEIEEEVEENEEDAEEEEEE*EKEEEEVE 456
Score = 32.7 bits (73), Expect = 0.27
Identities = 20/44 (45%), Positives = 27/44 (60%), Gaps = 1/44 (2%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEE-EEEEEEEEENVEVVNEVVTPPRAD 345
+E+EE E+ EE E++E+ EEEEEE E+ E V E T D
Sbjct: 352 EEEEEVEEIEEEVEENEEDAEEEEEE*EKEEEEVEEDDTMQNLD 483
Score = 32.3 bits (72), Expect = 0.35
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 4/41 (9%)
Frame = +1
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEE----EEEEEENVEVVNE 337
+ +E++ EE+ EE E+ EEEEE EEE EEN E E
Sbjct: 295 LEEENDPEEEMEEIEYEEVEEEEEVEEIEEEVEENEEDAEE 417
Score = 29.3 bits (64), Expect = 3.0
Identities = 15/36 (41%), Positives = 21/36 (57%)
Frame = +1
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
P+E+ EE + EE E++ EE EEE EE + E
Sbjct: 313 PEEEMEEIEYEEVEEEEEVEEIEEEVEENEEDAEEE 420
>TC88983 similar to SP|P93231|VP41_LYCES Vacuolar assembly protein VPS41
homolog. [Tomato] {Lycopersicon esculentum}, partial
(17%)
Length = 668
Score = 35.0 bits (79), Expect = 0.054
Identities = 14/27 (51%), Positives = 21/27 (76%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+DE EED E+ ED++EE EE+++EE
Sbjct: 146 DDEREEDEEDEEEEDEDEEVEEDDDEE 226
Score = 31.2 bits (69), Expect = 0.79
Identities = 13/21 (61%), Positives = 16/21 (75%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEE 323
+EDEEEED +E ED +EEE
Sbjct: 167 EEDEEEEDEDEEVEEDDDEEE 229
Score = 27.7 bits (60), Expect = 8.7
Identities = 10/16 (62%), Positives = 15/16 (93%)
Frame = +2
Query: 316 GEDKEEEEEEEEEEEN 331
G+D+ EE+EE+EEEE+
Sbjct: 143 GDDEREEDEEDEEEED 190
>TC93718 weakly similar to GP|10178129|dbj|BAB11541. gene_id:MOP10.6~unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 687
Score = 34.7 bits (78), Expect = 0.071
Identities = 15/30 (50%), Positives = 20/30 (66%)
Frame = +1
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+ Q + +E P +P D EE+EEEEEEEE
Sbjct: 49 LSQMETTQEQPSKPNNIDDEEDEEEEEEEE 138
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 34.7 bits (78), Expect = 0.071
Identities = 15/27 (55%), Positives = 23/27 (84%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+D +EED E+ GED+E+EE+EEE++E
Sbjct: 486 DDGDEEDEED--GEDEEDEEDEEEDDE 560
Score = 31.2 bits (69), Expect = 0.79
Identities = 11/28 (39%), Positives = 22/28 (78%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
ED++++ EE + ++EE+EE+EEE++
Sbjct: 474 EDDDDDGDEEDEEDGEDEEDEEDEEEDD 557
Score = 30.4 bits (67), Expect = 1.3
Identities = 21/74 (28%), Positives = 39/74 (52%)
Frame = +3
Query: 260 LIHAVRDRARRPVHVIEKRLPVAPLLSKERIAQLYKEWTARMPQEDEEEEDPEEPAGEDK 319
LI+A A + + ++ R P+ LL ++ YKE ++DE+++D ++ ED
Sbjct: 210 LINAANSVAPK-LTAMDSRFPLEHLLGRK---PAYKE-NKSDTEDDEDDDDDDDVQDEDD 374
Query: 320 EEEEEEEEEEENVE 333
+ EEE+ +E E
Sbjct: 375 DGEEEDYSGDEGEE 416
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 34.7 bits (78), Expect = 0.071
Identities = 11/27 (40%), Positives = 23/27 (84%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
ED++++D + GED++E+EEE+++E+
Sbjct: 460 EDDDDDDDDNDDGEDEDEDEEEDDDED 540
Score = 32.3 bits (72), Expect = 0.35
Identities = 10/28 (35%), Positives = 23/28 (81%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+D+E++D ++ +D E+E+E+EEE+++
Sbjct: 451 DDDEDDDDDDDDNDDGEDEDEDEEEDDD 534
Score = 30.8 bits (68), Expect = 1.0
Identities = 11/30 (36%), Positives = 22/30 (72%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+D++E+D ++ D E+E+E+EEE++ E
Sbjct: 448 DDDDEDDDDDDDDNDDGEDEDEDEEEDDDE 537
Score = 28.9 bits (63), Expect = 3.9
Identities = 10/34 (29%), Positives = 23/34 (67%), Gaps = 6/34 (17%)
Frame = +1
Query: 304 EDEEEEDPEE------PAGEDKEEEEEEEEEEEN 331
ED+E+ DPE+ G D ++E++++++++N
Sbjct: 388 EDDEDADPEDDPVPNGAGGSDDDDEDDDDDDDDN 489
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 - garden
pea, partial (93%)
Length = 1607
Score = 34.3 bits (77), Expect = 0.093
Identities = 12/33 (36%), Positives = 24/33 (72%)
Frame = +2
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ ED++E+D E ED++E+E+E+++E+ E
Sbjct: 1034 LDDEDDDEDDDAEDDDEDEDEDEDEDDDEDEEE 1132
Score = 28.9 bits (63), Expect = 3.9
Identities = 9/31 (29%), Positives = 23/31 (74%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
++D+E+ED +E +D++EEE + +++ + +
Sbjct: 1070 EDDDEDEDEDEDEDDDEDEEETKTKKKSSAK 1162
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,039,254
Number of Sequences: 36976
Number of extensions: 193328
Number of successful extensions: 3486
Number of sequences better than 10.0: 177
Number of HSP's better than 10.0 without gapping: 1583
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2482
length of query: 422
length of database: 9,014,727
effective HSP length: 99
effective length of query: 323
effective length of database: 5,354,103
effective search space: 1729375269
effective search space used: 1729375269
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0388a.2


