

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.1
(728 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
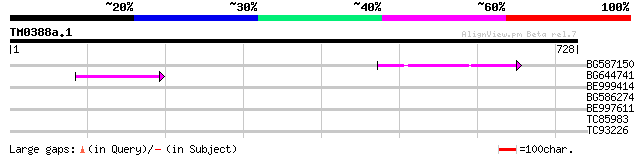
BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, part... 74 2e-13
BG644741 60 2e-09
BE999414 39 0.005
BG586274 33 0.51
BE997611 31 1.5
TC85983 weakly similar to GP|8777362|dbj|BAA96952.1 60S acidic r... 30 3.3
TC93226 weakly similar to PIR|B96737|B96737 hypothetical protein... 29 7.3
>BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, partial (1%)
Length = 754
Score = 73.9 bits (180), Expect = 2e-13
Identities = 56/187 (29%), Positives = 91/187 (47%), Gaps = 2/187 (1%)
Frame = +2
Query: 473 KALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETV- 531
K +AK K ++ F+K IP E+ A F + R + +E +
Sbjct: 113 KKVAKHLKRGANEKEMESFQKRVFMIPLEKPFEE----AYFTHRLWMFFRETRETEEDIR 280
Query: 532 -LMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNL 590
+ E + +R KK+ D G F I ++G+ ALC+ GAS+ ++P M + L L
Sbjct: 281 RMFCEAREKMKKRITLKKKSDSGKFAISWTMKGIEFPHALCNTGASVIILPRDMADHLGL 460
Query: 591 GEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFL 650
+V P+ S D S ++ GI+ D+ V + + PVDF VLD++ + L+L R FL
Sbjct: 461 -KVDPSKESFTFVDCSQRSSGGIIRDLEVQIGNALVPVDFHVLDIKINWNSSLLLRRAFL 637
Query: 651 ATGRAKI 657
+TGR +
Sbjct: 638 STGRGSV 658
>BG644741
Length = 735
Score = 60.5 bits (145), Expect = 2e-09
Identities = 30/114 (26%), Positives = 50/114 (43%)
Frame = -2
Query: 85 DTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQ 144
+ + L +FP SL A+ W P SI +W L + F R+ P + + + F
Sbjct: 599 NVLGLRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVA 420
Query: 145 STDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEF 198
E++ +W+RF LR P H + FY G +++ LD + G +
Sbjct: 419 LPGESVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGSY 258
>BE999414
Length = 613
Score = 39.3 bits (90), Expect = 0.005
Identities = 25/65 (38%), Positives = 31/65 (47%)
Frame = +3
Query: 62 DPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEK 121
DP HME I TYR ++ LF +LR A W + + SI SW DL +
Sbjct: 33 DPDEHMEN-IEVVLTYRSVR---GAVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGDLCHE 200
Query: 122 FTTRF 126
FTT F
Sbjct: 201 FTTHF 215
>BG586274
Length = 546
Score = 32.7 bits (73), Expect = 0.51
Identities = 23/105 (21%), Positives = 42/105 (39%), Gaps = 10/105 (9%)
Frame = -3
Query: 300 NDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQG----------QSLRQPQE 349
N ++N + N+ + ++ QG+ +Q PS G + + + Q
Sbjct: 340 NFQYNNYQQKSYPNNHQSGYPPRNNQQGSYQPQQNPSSGSSAPQESSTDTLLKQILESQT 161
Query: 350 RGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQL 394
R E K+L ++ N N + + + KNLENQF +
Sbjct: 160 RSEKHVGYELKNLHSKIDGSYNELNNKFSHLASTVKNLENQFASM 26
>BE997611
Length = 547
Score = 31.2 bits (69), Expect = 1.5
Identities = 14/59 (23%), Positives = 30/59 (50%)
Frame = -2
Query: 477 KKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTE 535
+K + KF+ + K +++P + +EQ+P KF++++L L + L +E
Sbjct: 444 EKEVPPKFNALHHILKGYKVNMPITETVEQIPCCIKFLQELLKTNANLSEEEFISLSSE 268
>TC85983 weakly similar to GP|8777362|dbj|BAA96952.1 60S acidic ribosomal
protein P3 {Arabidopsis thaliana}, partial (85%)
Length = 653
Score = 30.0 bits (66), Expect = 3.3
Identities = 17/61 (27%), Positives = 28/61 (45%)
Frame = +1
Query: 71 IRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRA 130
+RN N ++S ++I + +F F L T EW Q G I + + + + V A
Sbjct: 22 LRNFNIQLGFSLSPESITMGVFTFVLNKTGAEWTAKQYSGDIEASAESTFEIQRKLVQAA 201
Query: 131 L 131
L
Sbjct: 202 L 204
>TC93226 weakly similar to PIR|B96737|B96737 hypothetical protein F3I17.10
[imported] - Arabidopsis thaliana, partial (60%)
Length = 847
Score = 28.9 bits (63), Expect = 7.3
Identities = 15/49 (30%), Positives = 27/49 (54%)
Frame = -3
Query: 79 VQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFV 127
++N++ DTI +S++ +R + E L+ + SI S L TRF+
Sbjct: 746 IKNITKDTISVSIYKLGIRMVSVELLHH*SESSIVSSNHLIY*IHTRFL 600
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,131,146
Number of Sequences: 36976
Number of extensions: 245376
Number of successful extensions: 1206
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1201
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1204
length of query: 728
length of database: 9,014,727
effective HSP length: 103
effective length of query: 625
effective length of database: 5,206,199
effective search space: 3253874375
effective search space used: 3253874375
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0388a.1


