

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
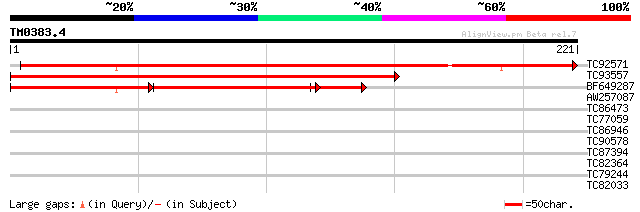
Query= TM0383.4
(221 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC92571 homologue to PIR|A96760|A96760 hypothetical protein T9L2... 332 5e-92
TC93557 homologue to PIR|A96760|A96760 hypothetical protein T9L2... 276 6e-75
BF649287 homologue to PIR|A96760|A96 hypothetical protein T9L24.... 125 5e-35
AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapi... 30 0.73
TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat prote... 29 1.6
TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 3.6
TC86946 homologue to GP|21539549|gb|AAM53327.1 putative guanine ... 27 4.8
TC90578 homologue to GP|10178144|dbj|BAB11589. gene_id:MPO12.6~u... 27 6.2
TC87394 similar to GP|3582779|gb|AAC69180.1| peroxisomal targeti... 27 8.1
TC82364 similar to GP|20268740|gb|AAM14073.1 unknown protein {Ar... 27 8.1
TC79244 weakly similar to PIR|B86193|B86193 hypothetical protein... 27 8.1
TC82033 similar to GP|2388586|gb|AAB71467.1| Similar to Saccharo... 27 8.1
>TC92571 homologue to PIR|A96760|A96760 hypothetical protein T9L24.52
[imported] - Arabidopsis thaliana, partial (56%)
Length = 951
Score = 332 bits (852), Expect = 5e-92
Identities = 179/231 (77%), Positives = 192/231 (82%), Gaps = 14/231 (6%)
Frame = +1
Query: 5 EVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAK-------------MQEVAGERGGY 51
EVREYTNL+DPKDKK GKGKDKIDDEDVTFQRMVAK MQEVAGERGGY
Sbjct: 1 EVREYTNLTDPKDKKLGKGKDKIDDEDVTFQRMVAKGSWLPVSAG*YKQMQEVAGERGGY 180
Query: 52 LHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRV 111
LHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASL DSSPASVPLPLRV
Sbjct: 181 LHGRGALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLADSSPASVPLPLRV 360
Query: 112 EPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSG 171
EPKPKSGIRQQDLLKKVVEIKPKRPRSE + A SD S+TNR+ D LK+ Q LSG
Sbjct: 361 EPKPKSGIRQQDLLKKVVEIKPKRPRSEGNKPMQAPSDASITNRQHDRDNLKDNKQCLSG 540
Query: 172 VKKVEEQSLSGSTKVEDLS-SGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
+KVEE+SLSG KVE+ S H+++ KPK++NS GGLLGLAYASSDDDE
Sbjct: 541 -QKVEERSLSGLKKVEEHPVSELHESETKPKVDNSAGGLLGLAYASSDDDE 690
>TC93557 homologue to PIR|A96760|A96760 hypothetical protein T9L24.52
[imported] - Arabidopsis thaliana, partial (58%)
Length = 497
Score = 276 bits (705), Expect = 6e-75
Identities = 140/152 (92%), Positives = 143/152 (93%)
Frame = +2
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 60
MAGREVREYTNL+DPKDKK GKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS
Sbjct: 41 MAGREVREYTNLTDPKDKKLGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDS 220
Query: 61 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIR 120
DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASL DSSPASVPLPLRVEPKPKSGIR
Sbjct: 221 DDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLADSSPASVPLPLRVEPKPKSGIR 400
Query: 121 QQDLLKKVVEIKPKRPRSESKQSTSASSDVSV 152
QQDLLKKVVEIKPKRPRSE + A SD S+
Sbjct: 401 QQDLLKKVVEIKPKRPRSEGNKPMQAPSDASI 496
>BF649287 homologue to PIR|A96760|A96 hypothetical protein T9L24.52
[imported] - Arabidopsis thaliana, partial (58%)
Length = 629
Score = 125 bits (313), Expect(2) = 5e-35
Identities = 64/65 (98%), Positives = 64/65 (98%)
Frame = +1
Query: 57 ALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPK 116
ALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASL DSSPASVPLPLRVEPKPK
Sbjct: 376 ALDSDDLLYLKEQMEAEEDAERLLRRTEKRAFAAFKRAASLADSSPASVPLPLRVEPKPK 555
Query: 117 SGIRQ 121
SGIRQ
Sbjct: 556 SGIRQ 570
Score = 104 bits (260), Expect = 2e-23
Identities = 54/69 (78%), Positives = 55/69 (79%), Gaps = 13/69 (18%)
Frame = +2
Query: 1 MAGREVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAK-------------MQEVAGE 47
MAGREVREYTNL+DPKDKK GKGKDKIDDEDVTFQRMVAK MQEVAGE
Sbjct: 41 MAGREVREYTNLTDPKDKKLGKGKDKIDDEDVTFQRMVAKGSWLPVSAG*YKQMQEVAGE 220
Query: 48 RGGYLHGRG 56
RGGYLHGRG
Sbjct: 221 RGGYLHGRG 247
Score = 39.7 bits (91), Expect(2) = 5e-35
Identities = 19/22 (86%), Positives = 19/22 (86%)
Frame = +2
Query: 118 GIRQQDLLKKVVEIKPKRPRSE 139
G QDLLKKVVEIKPKRPRSE
Sbjct: 560 GSGSQDLLKKVVEIKPKRPRSE 625
>AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapiens},
partial (2%)
Length = 466
Score = 30.0 bits (66), Expect = 0.73
Identities = 25/105 (23%), Positives = 43/105 (40%), Gaps = 9/105 (8%)
Frame = +1
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKK---------VE 176
KK V + K + +K ++S S ++ D T S+S VKK
Sbjct: 85 KKAVPVVTKTAPTHAKSASSKMEVDSDSDSDSDSSSSSSTSSSVSSVKKPAVSAAKATPA 264
Query: 177 EQSLSGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
++ S S D SS S + + ++S + ++SDD+E
Sbjct: 265 KKEESDSDSDSDSSSSSSSSSSSSSSSSSSSSSSSSSSSNSDDEE 399
>TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat protein
[imported] - Arabidopsis thaliana, partial (68%)
Length = 2654
Score = 28.9 bits (63), Expect = 1.6
Identities = 20/96 (20%), Positives = 39/96 (39%)
Frame = +2
Query: 126 KKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTK 185
K VVE + K+ +S K+ + D + +S + E S +
Sbjct: 44 KTVVESEMKKKKSNEKEEAPPPPSDHDDSDDDD----SDDSVEVSDYDSISEDEDSSADS 211
Query: 186 VEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
++D S D+D+K ++ + G + +SDD +
Sbjct: 212 IDDDDSSDSDSDSKSELEDEEGASPTTHHNASDDSD 319
>TC77059 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (96%)
Length = 1582
Score = 27.7 bits (60), Expect = 3.6
Identities = 18/79 (22%), Positives = 40/79 (49%), Gaps = 4/79 (5%)
Frame = +2
Query: 91 FKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPR----SESKQSTSA 146
F++ V + A P+P + +S + +D KKV + +P++P+ E+ ++ +
Sbjct: 677 FRKILGQVKQTEAMPPIPSAKQQPKESKPKTKDEPKKVAKSEPEKPKVEVEEEAPKAQTQ 856
Query: 147 SSDVSVTNRKPDHGRLKET 165
S +++ D G L++T
Sbjct: 857 ESS*PSSSK*DDFG*LEKT 913
>TC86946 homologue to GP|21539549|gb|AAM53327.1 putative guanine nucleotide
regulatory protein {Arabidopsis thaliana}, partial (35%)
Length = 892
Score = 27.3 bits (59), Expect = 4.8
Identities = 21/84 (25%), Positives = 44/84 (52%)
Frame = +1
Query: 4 REVREYTNLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDSDDL 63
+EV+E + ++ KDK+ +D+ D+ED+ ++K + + G++ + +
Sbjct: 274 QEVQEQSIKTEIKDKEIPPVQDEEDEEDI-----LSKKRHLNVVFIGHVDAGKSTTGGQI 438
Query: 64 LYLKEQMEAEEDAERLLRRTEKRA 87
LYL Q++ ER +++ EK A
Sbjct: 439 LYLSGQVD-----ERTIQKYEKEA 495
>TC90578 homologue to GP|10178144|dbj|BAB11589. gene_id:MPO12.6~unknown
protein {Arabidopsis thaliana}, partial (1%)
Length = 1100
Score = 26.9 bits (58), Expect = 6.2
Identities = 22/70 (31%), Positives = 30/70 (42%)
Frame = +2
Query: 132 KPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKVEDLSS 191
K K+P SE QS + S K + KE SLS VKK ++ + K D S
Sbjct: 449 KRKKPESELDQSDNISPSEDSGPAK----KSKERTTSLSKVKKRARETKTSGKKGTDEKS 616
Query: 192 GSHDADAKPK 201
+ + K K
Sbjct: 617 SAAERSKKSK 646
>TC87394 similar to GP|3582779|gb|AAC69180.1| peroxisomal targeting sequence
1 receptor {Nicotiana tabacum}, partial (65%)
Length = 1656
Score = 26.6 bits (57), Expect = 8.1
Identities = 30/99 (30%), Positives = 45/99 (45%)
Frame = +3
Query: 123 DLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSG 182
+ LK+ V K RS+S + SD++ P+ LKE GQ L + E L+
Sbjct: 285 EFLKEQVAAKQ---RSDSSRGVYVFSDLNPYVGHPNP--LKE-GQDLFRKGLLSEAVLAL 446
Query: 183 STKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
+V K N+ G LLG+A+A +DDD+
Sbjct: 447 EAEV-----------LKNPENSEGWRLLGIAHAENDDDQ 530
>TC82364 similar to GP|20268740|gb|AAM14073.1 unknown protein {Arabidopsis
thaliana}, partial (39%)
Length = 1174
Score = 26.6 bits (57), Expect = 8.1
Identities = 26/144 (18%), Positives = 56/144 (38%), Gaps = 17/144 (11%)
Frame = +3
Query: 68 EQMEAEEDAERLLRRTEKRAFAAFKRAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKK 127
++ + DA RR + K+ V+ VP PL ++PK K DL +
Sbjct: 378 DKAPVDVDAMETERRKIEAEDQLRKKMMKSVEDEKMKVPFPLFIKPKQKEKPPLFDLSEA 557
Query: 128 VVEIKPKRPRSESKQSTSASSDVSVTNRKPD----------HGRLKET-------GQSLS 170
+ ++K +++ ++ A + + +++ + HG K G
Sbjct: 558 IRQVK-ANAKAKFDETVEAHVRLGIDSKRTELAVRGTVILPHGAPKAVTVAVFAEGAQAE 734
Query: 171 GVKKVEEQSLSGSTKVEDLSSGSH 194
K + G +E+++SG++
Sbjct: 735 EAKAAGADIVGGKELIEEIASGNN 806
>TC79244 weakly similar to PIR|B86193|B86193 hypothetical protein
[imported] - Arabidopsis thaliana, partial (57%)
Length = 1114
Score = 26.6 bits (57), Expect = 8.1
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 67 KEQMEAEEDAERLLRRTEKRAFAAF 91
KE+ + +E ERL R+ EKR F F
Sbjct: 82 KEERQKKE*RERLTRKREKREFRGF 8
>TC82033 similar to GP|2388586|gb|AAB71467.1| Similar to Saccharomyces RAD16
(gb|X78993). {Arabidopsis thaliana}, partial (6%)
Length = 669
Score = 26.6 bits (57), Expect = 8.1
Identities = 19/52 (36%), Positives = 26/52 (49%), Gaps = 7/52 (13%)
Frame = +1
Query: 138 SESKQSTSASSDVSVTNRKPDHGRLKETGQS-------LSGVKKVEEQSLSG 182
S+ + +TS SSD S T+ K R ++ G LSGV+ E LSG
Sbjct: 22 SDGEDNTSDSSDFSGTSSKRRKTRSRKRGSKSKIESADLSGVESALEFDLSG 177
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.307 0.127 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,714,129
Number of Sequences: 36976
Number of extensions: 50088
Number of successful extensions: 263
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 257
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 259
length of query: 221
length of database: 9,014,727
effective HSP length: 92
effective length of query: 129
effective length of database: 5,612,935
effective search space: 724068615
effective search space used: 724068615
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0383.4


