

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0383.12
(339 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
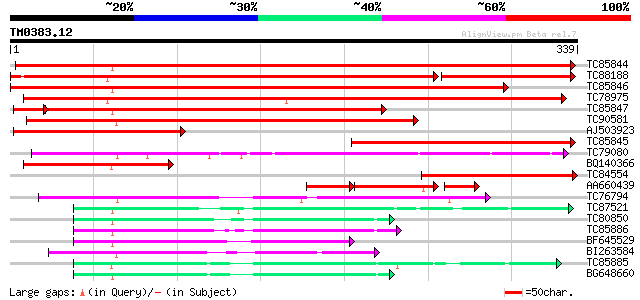
Score E
Sequences producing significant alignments: (bits) Value
TC85844 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 ... 373 e-104
TC88188 homologue to SP|P80572|ADHX_PEA Alcohol dehydrogenase cl... 294 9e-96
TC85846 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 ... 345 1e-95
TC78975 similar to GP|21592656|gb|AAM64605.1 alcohol dehydrogena... 342 1e-94
TC85847 homologue to SP|P13603|ADH1_TRIRP Alcohol dehydrogenase ... 258 9e-73
TC90581 similar to PIR|E86357|E86357 alcohol dehydrogenase homol... 228 3e-60
AJ503923 similar to GP|10129655|emb alcohol dehydrogenase-like p... 185 2e-47
TC85845 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 ... 127 6e-30
TC79080 similar to GP|10177043|dbj|BAB10455. alcohol dehydrogena... 126 1e-29
BQ140366 weakly similar to SP|P41681|ADH2_ Alcohol dehydrogenase... 96 2e-20
TC84554 weakly similar to GP|7705214|gb|AAB33480.2| alcohol dehy... 87 1e-17
AA660439 similar to PIR|S52036|S520 probable alcohol dehydrogena... 48 7e-17
TC76794 similar to GP|7416846|dbj|BAA94084.1 NAD-dependent sorbi... 84 8e-17
TC87521 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydro... 60 9e-10
TC80850 homologue to SP|O82515|MTD_MEDSA Probable mannitol dehyd... 58 6e-09
TC85886 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydro... 57 8e-09
BF645529 weakly similar to GP|22475166|gb putative sinapyl alcoh... 57 1e-08
BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1... 57 1e-08
TC85885 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydro... 54 7e-08
BG648660 similar to SP|Q9ZRF1|MTD_ Probable mannitol dehydrogena... 52 3e-07
>TC85844 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 (EC
1.1.1.1). [Garden pea] {Pisum sativum}, complete
Length = 1502
Score = 373 bits (958), Expect = e-104
Identities = 175/339 (51%), Positives = 244/339 (71%), Gaps = 4/339 (1%)
Frame = +2
Query: 4 STSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH--- 60
S + Q+I C+AAVAW AG+PLV+EEV+V+PPQ E+R+K++ TSLC +D+ WE+
Sbjct: 92 SNTAGQIIKCRAAVAWEAGKPLVMEEVEVAPPQAGEVRLKILFTSLCHTDVYFWEAKGQT 271
Query: 61 PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLER 120
P+FPRIFGHEA GIVES+G GVT K GDH L VF GEC +C C S +SN+C +L +
Sbjct: 272 PLFPRIFGHEAGGIVESIGEGVTHLKPGDHALPVFTGECGDCPHCKSEESNMCDLLRINT 451
Query: 121 N-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVA 179
+ G+M +D ++RFS+KG+P++H+ G S+FSEYTVVH+GC K++P+APL+K+C+LSCG+
Sbjct: 452 DRGVMINDNQSRFSLKGQPIHHFVGTSTFSEYTVVHAGCVAKINPDAPLDKVCILSCGIC 631
Query: 180 AGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGI 239
GLGA NVA GS+V IFGLG VGL+ A+GA++ GASRIIGVD + E AK FG+
Sbjct: 632 TGLGATINVAKPKPGSSVAIFGLGAVGLAAAEGARISGASRIIGVDLVSSRFELAKKFGV 811
Query: 240 TEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKV 299
E V+P +++P+ QVI +T+GG D + EC G + +A + DGWG+ V +GVP
Sbjct: 812 NEFVNPKDHDKPVQQVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNK 991
Query: 300 KPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYV 338
H LL RTLKG+ +G +KP++DLP++V+KY+
Sbjct: 992 DDAFKTHPMNLLNERTLKGTFYGNYKPRTDLPNVVEKYM 1108
>TC88188 homologue to SP|P80572|ADHX_PEA Alcohol dehydrogenase class III (EC
1.1.1.1) (Glutathione-dependent formaldehyde
dehydrogenase), complete
Length = 1597
Score = 294 bits (752), Expect(2) = 9e-96
Identities = 146/260 (56%), Positives = 188/260 (72%), Gaps = 4/260 (1%)
Frame = +1
Query: 1 MSSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAW--- 57
M+SST QVITCKAAVAW +PL IE+V+V+PPQ E+R++++ T+LC +D W
Sbjct: 223 MASSTEG-QVITCKAAVAWEPNKPLTIEDVEVAPPQANEVRIQILFTALCHTDAYTWSGK 399
Query: 58 ESHPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVC-QVL 116
+ +FP I GHEA+GIVESVG GVT+ K GDHV+ + EC ECK C SGK+N+C +V
Sbjct: 400 DPEGLFPCILGHEAAGIVESVGEGVTDVKPGDHVIPCYQAECGECKFCKSGKTNLCGKVR 579
Query: 117 GLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC 176
G+M +D+K RFSIKGKP+YH+ G S+FS+YTVVH K+ P+A L+K+CLL C
Sbjct: 580 AATGVGVMMNDRKPRFSIKGKPIYHFMGTSTFSQYTVVHDVSVAKIHPDAALDKVCLLGC 759
Query: 177 GVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKA 236
GV GLGA WN A V GS V IFGLGTVGL+VA+GAK GASRIIG+D + K + AK
Sbjct: 760 GVPTGLGAVWNTAKVEPGSIVAIFGLGTVGLAVAEGAKSAGASRIIGIDIDSNKFDTAKN 939
Query: 237 FGITEVVDPNSYEEPIAQVI 256
FG+TE ++P +E+PI QVI
Sbjct: 940 FGVTEFINPKDHEKPIQQVI 999
Score = 74.3 bits (181), Expect(2) = 9e-96
Identities = 34/80 (42%), Positives = 52/80 (64%)
Frame = +2
Query: 259 ITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKG 318
+TDGG D+SFEC+G+ ++ +AL+ GWG +V +GV +S L+ GR KG
Sbjct: 1007 LTDGGVDYSFECIGNVSVMRSALECTHKGWGTSVVVGVAASGQEISTRPFQLVTGRVWKG 1186
Query: 319 SMFGGWKPKSDLPSLVDKYV 338
+ FGG+K +S +P LV+KY+
Sbjct: 1187 TAFGGFKSRSQVPWLVEKYL 1246
>TC85846 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 (EC
1.1.1.1). [Garden pea] {Pisum sativum}, partial (78%)
Length = 955
Score = 345 bits (885), Expect = 1e-95
Identities = 163/302 (53%), Positives = 222/302 (72%), Gaps = 4/302 (1%)
Frame = +3
Query: 1 MSSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH 60
++ S +T QVI C+AAVAW AG+PLV+EEV+V+PPQ E+R+K++ TSLC +D+ WE+
Sbjct: 39 LTMSNTTVQVIKCRAAVAWEAGKPLVMEEVEVAPPQAGEVRLKILFTSLCHTDVYFWEAK 218
Query: 61 ---PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLG 117
P+FPRIFGHEA GIVESVG GVT K GDH L VF GEC +C C S +SN+C +L
Sbjct: 219 GQTPLFPRIFGHEAGGIVESVGEGVTHLKPGDHALPVFTGECGDCPHCKSEESNMCDLLR 398
Query: 118 LERN-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC 176
+ + G+M +D ++RFSIKG+P++H+ G S+FSEYTVVH+GC K++P+APL+K+C+LSC
Sbjct: 399 INTDRGVMINDNQSRFSIKGQPIHHFVGTSTFSEYTVVHAGCVAKINPDAPLDKVCILSC 578
Query: 177 GVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKA 236
G+ GLGA NVA GS+V IFGLG VGL+ A+GA++ GASRIIGVD + E AK
Sbjct: 579 GICTGLGATINVAKPKPGSSVAIFGLGAVGLAAAEGARISGASRIIGVDLVSSRFELAKK 758
Query: 237 FGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGV 296
FG+ E V+P +++P+ QVI +T+GG D + EC G + +A + DGWG+ V +GV
Sbjct: 759 FGVNEFVNPKDHDKPVQQVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGV 938
Query: 297 PK 298
PK
Sbjct: 939 PK 944
>TC78975 similar to GP|21592656|gb|AAM64605.1 alcohol dehydrogenase
putative {Arabidopsis thaliana}, partial (77%)
Length = 1691
Score = 342 bits (877), Expect = 1e-94
Identities = 171/333 (51%), Positives = 234/333 (69%), Gaps = 8/333 (2%)
Frame = +2
Query: 9 QVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAW----ESHPIFP 64
+ ITCKAAVA G GEP V+E++ V PPQ ME+R+K++ TS+C +D+S W E +P
Sbjct: 119 KTITCKAAVAHGPGEPFVVEQILVQPPQKMEVRIKILYTSICHTDLSGWKGECEPQRAYP 298
Query: 65 RIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNGLM 124
RIFGHEASGIVESVG GV + KE D V+ +F GEC ECK C K+N+C+ G++ +
Sbjct: 299 RIFGHEASGIVESVGEGVNDMKENDKVVLIFNGECGECKCCMCKKTNMCEKSGVDPMKKL 478
Query: 125 HSDQKTRFS-IKGKPVYHYCGVSSFSEYTVVHSGCAVKVSP--NAPLEKICLLSCGVAAG 181
D +RFS I GKP++H+ S+F+EYTVV S CAVK++ N L+K+ LLSCGV+ G
Sbjct: 479 MCDGTSRFSSIDGKPIFHFLNTSTFTEYTVVDSACAVKLNTEDNLSLKKLTLLSCGVSTG 658
Query: 182 LGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCEN-AKAFGIT 240
+GAAWN A+V GS+V IFGLG VGL+VA+GA+ +GAS+IIGVD NP K N A+ GIT
Sbjct: 659 IGAAWNNANVHAGSSVAIFGLGAVGLAVAEGARARGASKIIGVDINPDKFNNTAETMGIT 838
Query: 241 EVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVK 300
E ++P E+P+ ++I +TDGG D+SFEC G+ +++ + S +GWGLTV LG+
Sbjct: 839 EFINPKDEEKPVYEIIREMTDGGVDYSFECTGNLNVLRDSFLSVHEGWGLTVLLGIHGSP 1018
Query: 301 PVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSL 333
++ H L GR ++GS+FGG+K KS LP+L
Sbjct: 1019KLLPIHPMELFDGRRIEGSVFGGFKGKSQLPNL 1117
>TC85847 homologue to SP|P13603|ADH1_TRIRP Alcohol dehydrogenase 1 (EC
1.1.1.1). [Creeping white clover] {Trifolium repens},
partial (60%)
Length = 751
Score = 258 bits (660), Expect(2) = 9e-73
Identities = 121/208 (58%), Positives = 161/208 (77%), Gaps = 4/208 (1%)
Frame = +1
Query: 22 GEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESV 78
G+PLV+EEV+V+PPQ E+R+K++ TSLC +D+ WE+ P+FPRIFGHEA GIVESV
Sbjct: 106 GKPLVMEEVEVAPPQAGEVRLKILFTSLCHTDVYFWEAKGQTPLFPRIFGHEAGGIVESV 285
Query: 79 GLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERN-GLMHSDQKTRFSIKGK 137
G GVT GDH L VF GEC +C C S +SN+C +L + + G+M +D ++RFSIKG+
Sbjct: 286 GEGVTHLTPGDHALLVFTGECGDCPHCMSEESNMCDLLRINTDRGVMINDNQSRFSIKGQ 465
Query: 138 PVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTV 197
P++H+ G S+FSEYTVVH+GC K++P+APL+K+C+LSCG+ GLGA NVA GS+V
Sbjct: 466 PIHHFVGTSTFSEYTVVHAGCVAKINPDAPLDKVCILSCGICTGLGATINVAKPKPGSSV 645
Query: 198 VIFGLGTVGLSVAQGAKLKGASRIIGVD 225
IFGLG VGL+ A+GA++ GASRIIGVD
Sbjct: 646 AIFGLGAVGLAAAEGARISGASRIIGVD 729
Score = 33.1 bits (74), Expect(2) = 9e-73
Identities = 15/21 (71%), Positives = 17/21 (80%)
Frame = +3
Query: 3 SSTSTPQVITCKAAVAWGAGE 23
S+T+T QVI C AAVAW AGE
Sbjct: 48 SNTTTGQVIKCTAAVAWEAGE 110
>TC90581 similar to PIR|E86357|E86357 alcohol dehydrogenase homolog
[imported] - Arabidopsis thaliana, partial (52%)
Length = 1289
Score = 228 bits (580), Expect = 3e-60
Identities = 126/242 (52%), Positives = 162/242 (66%), Gaps = 8/242 (3%)
Frame = +1
Query: 11 ITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHPI-----FPR 65
I CKAA+ +GEPLVIEEV++ PP+ E+R+K++CTSLC SD++ W+ + FPR
Sbjct: 286 IRCKAAICKKSGEPLVIEEVELDPPKSWEVRIKILCTSLCHSDVTFWKMNSSAPTARFPR 465
Query: 66 IFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNGLMH 125
I GHEA G+VESVG V E KEGD V+ VF+ C EC C S KSN C G + M
Sbjct: 466 ILGHEAVGLVESVGENVEEVKEGDLVVPVFLPNCGECIDCGSTKSNNCTKFGNKPIRDMP 645
Query: 126 SDQKTRF-SIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGA 184
D +RF +KG+ V+H GVSSFSEYTVV VK++ + PL+K CLLSCGV+ G+GA
Sbjct: 646 RDGTSRFRDMKGEVVHHLLGVSSFSEYTVVDVTHVVKITHDIPLDKACLLSCGVSTGIGA 825
Query: 185 AWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRII-GVDNNPQKCENAK-AFGITEV 242
AW VADV KG+TV IFGLG VGL+VA K +GAS+I VD N + N + GIT+
Sbjct: 826 AWKVADVEKGTTVAIFGLGAVGLAVAVATKQRGASKIYRRVDLNHDEI*NEENKCGITDC 1005
Query: 243 VD 244
++
Sbjct: 1006ME 1011
>AJ503923 similar to GP|10129655|emb alcohol dehydrogenase-like protein
{Arabidopsis thaliana}, partial (25%)
Length = 350
Score = 185 bits (470), Expect = 2e-47
Identities = 88/103 (85%), Positives = 96/103 (92%)
Frame = +2
Query: 3 SSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHPI 62
+S+STPQVITCKAAVAWGAGE LV+EEV+VSPPQP+EIR+KVV TSLCRSD+SAWE+H I
Sbjct: 41 ASSSTPQVITCKAAVAWGAGEALVMEEVEVSPPQPLEIRIKVVSTSLCRSDLSAWETHAI 220
Query: 63 FPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPC 105
FPRI GHEASGIVESVGLGVTEFKEGD VLTVFIGECM CK C
Sbjct: 221 FPRISGHEASGIVESVGLGVTEFKEGDQVLTVFIGECMSCKLC 349
>TC85845 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 (EC
1.1.1.1). [Garden pea] {Pisum sativum}, partial (45%)
Length = 734
Score = 127 bits (319), Expect = 6e-30
Identities = 60/134 (44%), Positives = 88/134 (64%)
Frame = +1
Query: 205 VGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGA 264
VGL+ A+GA++ GASRIIGVD + E AK FG+ E V+P +++P+ QVI +T+GG
Sbjct: 1 VGLAAAEGARISGASRIIGVDLVSSRFELAKKFGVNEFVNPKDHDKPVQQVIAEMTNGGV 180
Query: 265 DFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGW 324
D + EC G + +A + DGWG+ V +GVP H LL RTLKG+ +G +
Sbjct: 181 DRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNKDDAFQTHPMNLLNERTLKGTFYGNY 360
Query: 325 KPKSDLPSLVDKYV 338
KP++DLP++V+KY+
Sbjct: 361 KPRTDLPNVVEKYM 402
>TC79080 similar to GP|10177043|dbj|BAB10455. alcohol dehydrogenase-like
protein {Arabidopsis thaliana}, partial (96%)
Length = 1524
Score = 126 bits (317), Expect = 1e-29
Identities = 98/333 (29%), Positives = 153/333 (45%), Gaps = 12/333 (3%)
Frame = +3
Query: 14 KAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHPIF--PRIFGHEA 71
+ AV +PL IEE + P+ E+ +K +C SD+ + F P + GHE
Sbjct: 234 RGAVFREPNKPLTIEEFHIPRPKAGELLIKTKGCGVCHSDLHVMKGEIPFSSPCVVGHEI 413
Query: 72 SGIVESVGLG-----VTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGL--ERNGLM 124
+G V G + G V+ FI C C C+ G ++C+ G +
Sbjct: 414 TGEVVEHGQHTDSKTIERLPIGSRVVGAFIMPCGNCSYCSKGHDDLCEAFFAYNRAKGTL 593
Query: 125 HSDQKTRFSIKGK--PVYHYCGVSSFSEYTVVHSGCAVKVSPNA-PLEKICLLSCGVAAG 181
+ D +TR ++G P+Y Y + +EY VV + A+ V PN+ P + +L C V
Sbjct: 594 Y-DGETRLFLRGSGNPIYMY-SMGGLAEYCVVPAN-ALAVLPNSMPYTESAILGCAVFTA 764
Query: 182 LGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITE 241
GA + A+V G TV + G G VG S Q A+ GAS II VD +K E AK G T
Sbjct: 765 YGAMAHAAEVRPGDTVAVIGTGGVGSSCLQIARAFGASDIIAVDVQDEKLEKAKTLGATH 944
Query: 242 VVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKP 301
++ ++ E+PI +++ G D + E +G QS DG G V +G+ +
Sbjct: 945 TIN-SAKEDPIEKILEITGGKGVDVAVEALGRPQTFAQCTQSVKDG-GKAVMIGLAQAGS 1118
Query: 302 VMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLV 334
+ L+ + +GG + + DLP L+
Sbjct: 1119LGEVDINRLVRRKIKVIGSYGG-RARQDLPKLI 1214
>BQ140366 weakly similar to SP|P41681|ADH2_ Alcohol dehydrogenase 2 (EC
1.1.1.1). [Deer mouse] {Peromyscus maniculatus}, partial
(11%)
Length = 327
Score = 95.9 bits (237), Expect = 2e-20
Identities = 43/92 (46%), Positives = 63/92 (67%), Gaps = 2/92 (2%)
Frame = +3
Query: 9 QVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWES--HPIFPRI 66
+++ K AV WG G+P+ +EE+QV PP+ E+RVK++C+S+C +DIS+ + H FP
Sbjct: 51 RLLHAKPAVCWGLGKPVTVEEIQVDPPKAKEVRVKMLCSSVCHTDISSLKGFPHNQFPLX 230
Query: 67 FGHEASGIVESVGLGVTEFKEGDHVLTVFIGE 98
GHE G+VES+G VT KEGD + +IGE
Sbjct: 231 LGHEGIGVVESIGDQVTNLKEGDFIXPTYIGE 326
>TC84554 weakly similar to GP|7705214|gb|AAB33480.2| alcohol dehydrogenase
ADH {Lycopersicon esculentum}, partial (27%)
Length = 550
Score = 86.7 bits (213), Expect = 1e-17
Identities = 41/93 (44%), Positives = 62/93 (66%)
Frame = +2
Query: 247 SYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAH 306
S E+ +++VI +T+GGAD+ FEC+G ++T A S +GWG TV +GV ++ +
Sbjct: 20 SNEKSVSEVIKDMTNGGADYCFECIGLASLMTEAFNSSREGWGKTVIIGVEMHGSPLTLN 199
Query: 307 YGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
+L G+T+ GS+FGG KPKSDLP L KY++
Sbjct: 200 PYDILKGKTITGSLFGGLKPKSDLPLLAQKYLD 298
>AA660439 similar to PIR|S52036|S520 probable alcohol dehydrogenase (EC
1.1.1.1) ADH3b - tomato, partial (21%)
Length = 403
Score = 48.1 bits (113), Expect(3) = 7e-17
Identities = 27/52 (51%), Positives = 37/52 (70%), Gaps = 2/52 (3%)
Frame = +2
Query: 207 LSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPN--SYEEPIAQVI 256
L+VA AK +GAS+IIGVD N K E K FGIT+ V+P+ S E+ +++VI
Sbjct: 89 LAVAVAAKQRGASKIIGVDLNHDKFEIGKQFGITDFVNPSSTSNEKSVSEVI 244
Score = 48.1 bits (113), Expect(3) = 7e-17
Identities = 21/29 (72%), Positives = 24/29 (82%)
Frame = +3
Query: 178 VAAGLGAAWNVADVSKGSTVVIFGLGTVG 206
V+ G+GAAW VADV KG+TV IFGLG VG
Sbjct: 3 VSTGIGAAWKVADVEKGTTVAIFGLGAVG 89
Score = 27.7 bits (60), Expect(3) = 7e-17
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +1
Query: 261 DGGADFSFECVGDTDMITTAL 281
+GGAD+ FEC+G ++T L
Sbjct: 256 NGGADYCFECIGLATLMTEXL 318
>TC76794 similar to GP|7416846|dbj|BAA94084.1 NAD-dependent sorbitol
dehydrogenase {Prunus persica}, partial (93%)
Length = 1691
Score = 84.0 bits (206), Expect = 8e-17
Identities = 72/283 (25%), Positives = 118/283 (41%), Gaps = 13/283 (4%)
Frame = +2
Query: 18 AWGAG-EPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHPIF------PRIFGHE 70
AW G L I+ + P ++R+K+ +C SD+ ++ P + GHE
Sbjct: 269 AWLVGLNTLKIQPFNLPSLGPHDVRIKMKAVGICGSDVHYLKTLRCADFIVKEPMVIGHE 448
Query: 71 ASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKT 130
+GI+E VG V GD V C C C G+ N+C + +H
Sbjct: 449 CAGIIEEVGSQVKTLVPGDRVAIEPGISCWRCDHCKLGRYNLCPDMKFFATPPVH----- 613
Query: 131 RFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICL---LSCGVAAGLGAAWN 187
S + V + K+ N LE+ + LS GV A
Sbjct: 614 ---------------GSLANQIVHPADLCFKLPENVSLEEGAMCEPLSVGV-----HACR 733
Query: 188 VADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNS 247
A++ + V+I G G +GL A+ GA RI+ VD + + AK+ G ++V ++
Sbjct: 734 RANIGPETNVLIMGAGPIGLVTMLSARAFGAPRIVVVDVDDHRLSVAKSLGADDIVKVST 913
Query: 248 YEEPIAQVINRITD---GGADFSFECVGDTDMITTALQSCCDG 287
+ +A+ + +I + G D +F+C G +TTAL + G
Sbjct: 914 NIQDVAEEVKQIHNVLGAGVDVTFDCAGFNKTMTTALTATQPG 1042
>TC87521 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (98%)
Length = 1433
Score = 60.5 bits (145), Expect = 9e-10
Identities = 75/307 (24%), Positives = 112/307 (36%), Gaps = 8/307 (2%)
Frame = +2
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ KV+ +C SD+ ++ +P + GHE +GIV VG V +FK GD V +
Sbjct: 182 DVAFKVLYCGICHSDLHMIKNEWGMSTYPLVPGHEIAGIVTEVGSKVEKFKIGDKVGVGC 361
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIK---GKPVYHYCGVSSFSEY 151
+ C C+ C N C Q +S K G Y +S+
Sbjct: 362 LVDSCRACQNCEENLENYC------------PKQTNTYSAKYSDGSITY-----GGYSDS 490
Query: 152 TVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQ 211
V V + PLE L C G + I GLG +G +
Sbjct: 491 MVADEHFIVHIPDGLPLESAAPLLCAGITVYSPLRYFGLDKPGMNIGIVGLGGLGHLGVK 670
Query: 212 GAKLKGAS-RIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFEC 270
AK GA+ +I N +K E + G + S+++ Q DG D
Sbjct: 671 FAKAFGANVTVISTSPNKEK-EAIENLGADSFL--ISHDQDKMQAAMGTLDGIID----- 826
Query: 271 VGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDL 330
D L G V +G P KP H L++ +T+ GS GG K ++
Sbjct: 827 TVSADHPLLPLVGLLKYHGKLVMVGAPD-KPPELPHIPLIMGRKTISGSGIGGMKETQEM 1003
Query: 331 PSLVDKY 337
K+
Sbjct: 1004IDFAAKH 1024
>TC80850 homologue to SP|O82515|MTD_MEDSA Probable mannitol dehydrogenase
(EC 1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Alfalfa], partial (61%)
Length = 710
Score = 57.8 bits (138), Expect = 6e-09
Identities = 46/196 (23%), Positives = 78/196 (39%), Gaps = 4/196 (2%)
Frame = +1
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ VK++ +C SD+ ++ +P + GHE G+V VG+ V +F+ GD+V + V
Sbjct: 166 DVSVKILYCGVCHSDLHTLKNDWGFTTYPVVPGHEIVGVVTKVGINVKKFRVGDNVGVGV 345
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVV 154
+ C C+ C C L N KG + +S++ VV
Sbjct: 346 IVESCQTCENCNQDLEQYCPKLVFTYNS----------PYKGTRTH-----GGYSDFVVV 480
Query: 155 HSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAK 214
H V+ N PL+ L C G + + GLG +G + K
Sbjct: 481 HQRYVVQFPDNLPLDAGAPLLCAGITVYSPMKYYGMTEPGKHLGVAGLGGLGHVAIKFGK 660
Query: 215 LKGASRIIGVDNNPQK 230
G ++ + +P K
Sbjct: 661 AFGL-KVTVITTSPNK 705
>TC85886 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (94%)
Length = 1395
Score = 57.4 bits (137), Expect = 8e-09
Identities = 49/200 (24%), Positives = 82/200 (40%), Gaps = 4/200 (2%)
Frame = +2
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ KV+ +C SD+ ++ +P + GHE +G+V VG V +FK GD V +
Sbjct: 185 DVAFKVLYCGICHSDLHIMQNEWGKTTYPLVPGHELTGVVTEVGSKVKKFKVGDKVGVGY 364
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVV 154
+ C C+ C N C NG K+R G Y FS+ V
Sbjct: 365 MVDSCRSCENCADDIENYCTKYTQTFNG------KSR---DGTITY-----GGFSDSMVA 502
Query: 155 HSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAK 214
+++ + PL+ L C + G + + GLG +G + AK
Sbjct: 503 DEHFVIRIPDSLPLDGAGPLLCAGVTVYSPLRHFGLDKPGMNIGVVGLGGLGHMAVKFAK 682
Query: 215 LKGASRIIGVDNNPQKCENA 234
GA ++ + +P+K + A
Sbjct: 683 AFGA-KVTVISTSPKKEKEA 739
>BF645529 weakly similar to GP|22475166|gb putative sinapyl alcohol
dehydrogenase {Populus tremula x Populus tremuloides},
partial (40%)
Length = 658
Score = 57.0 bits (136), Expect = 1e-08
Identities = 38/173 (21%), Positives = 74/173 (41%), Gaps = 5/173 (2%)
Frame = +3
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ +K++ +C +D+ + ++P + GHE +G++ VG V FK+GD V +
Sbjct: 141 DVTIKILYCGICHTDVHHAKDDWGITMYPVVPGHEITGVITKVGSDVEGFKDGDKVGVGC 320
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVV 154
C++C+ C + + N C+ L NG+ T +S+ VV
Sbjct: 321 LAASCLDCEYCKTDQENYCEKLQFVYNGIFWDGSIT--------------YGGYSQMLVV 458
Query: 155 HSGCAVKVSPNAPLEKICLLSC-GVAAGLGAAWNVADVSKGSTVVIFGLGTVG 206
V + + P++ L C G+ + + G + + GLG +G
Sbjct: 459 DYRYVVHIPESLPMDAAAPLLCAGITVFSPLKDHXLVSTAGKRIGVVGLGXLG 617
>BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1.1.1.1).
{Neurospora crassa}, partial (57%)
Length = 682
Score = 56.6 bits (135), Expect = 1e-08
Identities = 49/205 (23%), Positives = 87/205 (41%), Gaps = 7/205 (3%)
Frame = +2
Query: 24 PLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH-PI---FPRIFGHEASGIVESVG 79
P+ +++ V P P E+ V V + +C +D+ AW+ P+ P + GHE +G+V + G
Sbjct: 122 PIEYKQIPVQKPGPDEVLVNVKFSGVCHTDLHAWQGDWPLDTKLPLVGGHEGAGVVVARG 301
Query: 80 LGVTEFKEGDHVLTVFI-GECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKP 138
V + K G+ V ++ G C+ C C + ++C L + +++ G
Sbjct: 302 DLVKDVKIGEKVGIKWLNGSCLSCSYCQNADESLCAEALL-----------SGYTVDG-- 442
Query: 139 VYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC-GVAAGLGAAWNVADVSKGSTV 197
SF +Y + + ++ LE I + C G+ G + V G ++
Sbjct: 443 --------SFQQYAIAKAIHVARIPEECDLESISPILCAGITVYKGL--KESGVKAGQSI 592
Query: 198 VIFGL-GTVGLSVAQGAKLKGASRI 221
I G G +G Q K G I
Sbjct: 593 AIVGAGGGLGSIAXQYCKAMGIHAI 667
>TC85885 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (97%)
Length = 1330
Score = 54.3 bits (129), Expect = 7e-08
Identities = 65/300 (21%), Positives = 115/300 (37%), Gaps = 8/300 (2%)
Frame = +3
Query: 39 EIRVKVVCTSLCRSDISAWES---HPIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ KV+ +C +D+ ++ + I+P + GHE +GIV VG V +FK GD V +
Sbjct: 183 DVAFKVLYCGICHTDLHMMKNEWGNSIYPLVPGHELAGIVTEVGSKVEKFKVGDKVGVGY 362
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVV 154
+ C C+ C N C + G + D + +S+ V
Sbjct: 363 MVDSCRSCENCAEDLENYCPQQTV-TCGAKYRDGSVTY-------------GGYSDSMVA 500
Query: 155 HSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAK 214
+++ + PL+ L C + G + + GLG +G + AK
Sbjct: 501 DEHFVIRIPDSLPLDVAGPLLCAGVTVYSPLRHFQLDKPGMNIGVVGLGGLGHMAVKFAK 680
Query: 215 LKGASRIIGVDNNPQK----CENAKAFGITEVVDPNSYEEPIAQVINRITDGGADFSFEC 270
GA+ + + +P K E+ A DP+ + + +N I D
Sbjct: 681 AFGAN-VTVISTSPSKEKEAIEHLGADSFLVSRDPDQMQAAMG-TLNGIID--------T 830
Query: 271 VGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDL 330
V + I + L + GV KP+ + LL + + GS+ GG K ++
Sbjct: 831 VSASHPILPLIGLLKSNGKLVMVGGV--AKPLELPVFSLLGGRKLVAGSLIGGIKETQEM 1004
>BG648660 similar to SP|Q9ZRF1|MTD_ Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (64%)
Length = 721
Score = 52.4 bits (124), Expect = 3e-07
Identities = 46/196 (23%), Positives = 78/196 (39%), Gaps = 4/196 (2%)
Frame = +1
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ ++V+ +C SD+ ++ +P + GHE GIV VG V +FK GD V +
Sbjct: 130 DVALRVLYCGICHSDLHKAKNEWGTTNYPLVPGHEIVGIVTEVGSKVKKFKVGDRVGVGC 309
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVV 154
+ C C+ C N C L L +G +SD + FS+ V
Sbjct: 310 LVDSCHSCQNCVDNLENYCPQLTL-TDGSKYSDGTSTH-------------GGFSDSMVS 447
Query: 155 HSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAK 214
++ PL+ L C + + + GLG +G + AK
Sbjct: 448 DEHFVFRIPDQLPLDAAAPLLCAGITVYSPLRHFGLDKPDMNIGVVGLGGLGHMAVKFAK 627
Query: 215 LKGASRIIGVDNNPQK 230
GA+ + + +P+K
Sbjct: 628 AFGAN-VTVISTSPKK 672
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,708,465
Number of Sequences: 36976
Number of extensions: 171464
Number of successful extensions: 741
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 713
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 718
length of query: 339
length of database: 9,014,727
effective HSP length: 97
effective length of query: 242
effective length of database: 5,428,055
effective search space: 1313589310
effective search space used: 1313589310
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0383.12


