

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0381.8
(460 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
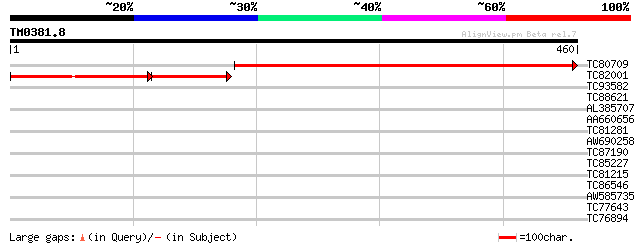
Sequences producing significant alignments: (bits) Value
TC80709 similar to PIR|B84616|B84616 hypothetical protein At2g22... 460 e-130
TC82001 similar to GP|14334566|gb|AAK59462.1 unknown protein {Ar... 200 1e-82
TC93582 similar to PIR|B84616|B84616 hypothetical protein At2g22... 36 0.027
TC88621 similar to PIR|H84698|H84698 hypothetical protein At2g29... 34 0.14
AL385707 homologue to GP|16755592|gb| small GTPase Rab2 {Nicotia... 33 0.30
AA660656 weakly similar to GP|6498428|dbj| Similar to Arabidopsi... 32 0.52
TC81281 similar to PIR|D96589|D96589 hypothetical protein T22H22... 30 2.6
AW690258 GP|22900886|gb amt2-like protein {Medicago truncatula},... 28 7.4
TC87190 amt2-like protein [Medicago truncatula] 28 7.4
TC85227 similar to GP|15983773|gb|AAL10483.1 AT4g29060/F19B15_90... 28 7.4
TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Ar... 28 7.4
TC86546 weakly similar to PIR|T45944|T45944 laccase-like protein... 28 7.4
AW585735 similar to GP|13958313|gb| NADPH oxidase 1 {Podospora a... 28 9.7
TC77643 similar to GP|21105742|gb|AAM34770.1 nam-like protein 7 ... 28 9.7
TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protea... 28 9.7
>TC80709 similar to PIR|B84616|B84616 hypothetical protein At2g22730
[imported] - Arabidopsis thaliana, partial (40%)
Length = 1250
Score = 460 bits (1184), Expect = e-130
Identities = 227/279 (81%), Positives = 256/279 (91%), Gaps = 1/279 (0%)
Frame = +3
Query: 183 PLESKKTISSIETNVSEHGDDGVLAQDQAFIRGTRSTSKLRNQFTRFSKDMQELLYDQVY 242
P+ESK+T +S ETNVSE+GDDG+LA+DQAFIRG++ TSKL NQFTRF DMQELL+++VY
Sbjct: 6 PMESKQTRTSNETNVSENGDDGILAEDQAFIRGSKLTSKLGNQFTRFLNDMQELLHERVY 185
Query: 243 IINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVCGILGTLGGGLILDRM 302
+INVLGYI+YNFVIGAYSYWGPKAGYSIYHMSN DLLFGGITIVCGI GTL GGLILD+M
Sbjct: 186 VINVLGYIAYNFVIGAYSYWGPKAGYSIYHMSNADLLFGGITIVCGIFGTLAGGLILDKM 365
Query: 303 TSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIVFFSIGELLIFATQAPVNYVSLRCV 362
+STISNAFK+LSGAT LGAIFCLVAFLFK L GFI+ FS+GELLIFATQAPVNYVSLRCV
Sbjct: 366 SSTISNAFKILSGATFLGAIFCLVAFLFKGLFGFIILFSVGELLIFATQAPVNYVSLRCV 545
Query: 363 KPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDRINDWRKSALCLTSVFFLAAGIWFIGI 422
KPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQD INDWRK+++CLTS+FFLAAG+WFIG
Sbjct: 546 KPSLRPLSMAISTVSIHIFGDVPSAPLVGVLQDHINDWRKTSICLTSIFFLAAGVWFIGT 725
Query: 423 FLRTVDLSTEDD-EDQSATSLRGKMKPLLEGNSDASSQA 460
FL++ DL +DD ED+S T+LRG KPLLEG +DASSQA
Sbjct: 726 FLKSDDLFNKDDEEDESTTTLRGVRKPLLEGINDASSQA 842
>TC82001 similar to GP|14334566|gb|AAK59462.1 unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 623
Score = 200 bits (509), Expect(2) = 1e-82
Identities = 99/116 (85%), Positives = 107/116 (91%)
Frame = +3
Query: 1 MASEPGQTPNPSWFTPKRLLLIFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGS 60
M + GQ PNPSWFTPKRLL+IFC+IN INYVDRGAIASNGVNG L T T+ SGVCTAG+
Sbjct: 54 MVPKSGQPPNPSWFTPKRLLIIFCIINLINYVDRGAIASNGVNGTLETRTE-SGVCTAGT 230
Query: 61 GIQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVA 116
GIQGDFNL+NFQDGVLSSAFMVGLLIASPIFASLAKSH+PFRLIGVGLSVWTFAV+
Sbjct: 231 GIQGDFNLSNFQDGVLSSAFMVGLLIASPIFASLAKSHNPFRLIGVGLSVWTFAVS 398
Score = 124 bits (312), Expect(2) = 1e-82
Identities = 58/65 (89%), Positives = 59/65 (90%)
Frame = +1
Query: 116 AGCGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTAL 175
AGCG SFDFWSI ICRML+GV EASFISLAAPFIDDNAP AQKTAWLATFYMCIP GTAL
Sbjct: 397 AGCGSSFDFWSIAICRMLIGVSEASFISLAAPFIDDNAPAAQKTAWLATFYMCIPDGTAL 576
Query: 176 GYVYG 180
GYV G
Sbjct: 577 GYVNG 591
>TC93582 similar to PIR|B84616|B84616 hypothetical protein At2g22730
[imported] - Arabidopsis thaliana, partial (4%)
Length = 591
Score = 36.2 bits (82), Expect = 0.027
Identities = 20/41 (48%), Positives = 23/41 (55%)
Frame = +3
Query: 411 FFLAAGIWFIGIFLRTVDLSTEDDEDQSATSLRGKMKPLLE 451
FF AA IWFIGIFL + D E+ E Q + PLLE
Sbjct: 9 FFPAAVIWFIGIFLNSKDKFNEESEHQVSRVEGTTTAPLLE 131
>TC88621 similar to PIR|H84698|H84698 hypothetical protein At2g29650
[imported] - Arabidopsis thaliana, partial (49%)
Length = 1074
Score = 33.9 bits (76), Expect = 0.14
Identities = 30/127 (23%), Positives = 54/127 (41%), Gaps = 6/127 (4%)
Frame = +1
Query: 62 IQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAV------ 115
+ ++N N G++ S+F G L+ A + +++ G+ W+ A
Sbjct: 415 MSAEYNWNPTTVGLVQSSFFWGYLLTQIAGGIWADTVGGKQVLAFGVIWWSVATILTPVA 594
Query: 116 AGCGISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTAL 175
A G+ F + + R +G+GE + + PVA+++ LA Y +G L
Sbjct: 595 AKLGLPF----LLVARAFMGIGEGVAMPAMNNILSKWVPVAERSRSLALVY----SGMYL 750
Query: 176 GYVYGFA 182
G V G A
Sbjct: 751 GSVTGLA 771
>AL385707 homologue to GP|16755592|gb| small GTPase Rab2 {Nicotiana tabacum},
partial (10%)
Length = 540
Score = 32.7 bits (73), Expect = 0.30
Identities = 14/21 (66%), Positives = 17/21 (80%)
Frame = -3
Query: 288 GILGTLGGGLILDRMTSTISN 308
G GTL GGL+LD MT+T+SN
Sbjct: 64 GTRGTLAGGLVLDYMTNTLSN 2
>AA660656 weakly similar to GP|6498428|dbj| Similar to Arabidopsis thaliana
chromosome II BAC T27A16 sequence; hypothetical protein.
(AC005496), partial (14%)
Length = 634
Score = 32.0 bits (71), Expect = 0.52
Identities = 23/91 (25%), Positives = 42/91 (45%), Gaps = 1/91 (1%)
Frame = +3
Query: 66 FNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFW 125
F N+ G++ S+F G ++ LAK ++ VG+ +W+ A A +
Sbjct: 306 FGWNSSTAGLVQSSFFWGYALSQLPGGWLAKIFGGXNVLVVGVLIWSVATALVPFLAGYM 485
Query: 126 -SITICRMLVGVGEASFISLAAPFIDDNAPV 155
+ + R+LVG+ E+ S A I + P+
Sbjct: 486 PGLILTRILVGIXESVSPSAATDLIAXSIPL 578
>TC81281 similar to PIR|D96589|D96589 hypothetical protein T22H22.15
[imported] - Arabidopsis thaliana, partial (24%)
Length = 585
Score = 29.6 bits (65), Expect = 2.6
Identities = 28/125 (22%), Positives = 48/125 (38%), Gaps = 2/125 (1%)
Frame = +2
Query: 58 AGSGIQGDFNLNNFQDGVLSSAFMVGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAG 117
A SGI D NL + + S +G ++ + + SLA +G
Sbjct: 164 AQSGITDDLNLGVAEYSLFGSILTIGAMVGAIVSGSLADYAGRRAAMGFSELFCILGWLA 343
Query: 118 CGISFDFWSITICRMLVGVGEASFISLAAPFID--DNAPVAQKTAWLATFYMCIPAGTAL 175
+S W + + R+L+G G +S P +N + + L + I G +L
Sbjct: 344 IAVSKVAWWLYVGRLLLGCG-MGILSYVVPIYIA*NNTQGSSRGIHLQFIKLMICFGVSL 520
Query: 176 GYVYG 180
Y+ G
Sbjct: 521 TYLIG 535
>AW690258 GP|22900886|gb amt2-like protein {Medicago truncatula}, partial
(23%)
Length = 450
Score = 28.1 bits (61), Expect = 7.4
Identities = 23/77 (29%), Positives = 37/77 (47%), Gaps = 6/77 (7%)
Frame = +1
Query: 249 YISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFG-GITIVCGILG-----TLGGGLILDRM 302
++ +++ +GA+S WG G +YH D G I + GI G +G + DR
Sbjct: 139 WLIFSYTVGAFSLWG---GGFLYHWGVIDYSGGYVIHLSSGIAGFTAAYWVGPRIKSDRE 309
Query: 303 TSTISNAFKLLSGATLL 319
+N +L+GA LL
Sbjct: 310 RFPPNNVLLMLAGAGLL 360
>TC87190 amt2-like protein [Medicago truncatula]
Length = 1912
Score = 28.1 bits (61), Expect = 7.4
Identities = 23/77 (29%), Positives = 37/77 (47%), Gaps = 6/77 (7%)
Frame = +2
Query: 249 YISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFG-GITIVCGILG-----TLGGGLILDRM 302
++ +++ +GA+S WG G +YH D G I + GI G +G + DR
Sbjct: 584 WLIFSYTVGAFSLWG---GGFLYHWGVIDYSGGYVIHLSSGIAGFTAAYWVGPRIKSDRE 754
Query: 303 TSTISNAFKLLSGATLL 319
+N +L+GA LL
Sbjct: 755 RFPPNNVLLMLAGAGLL 805
>TC85227 similar to GP|15983773|gb|AAL10483.1 AT4g29060/F19B15_90 {Arabidopsis
thaliana}, partial (63%)
Length = 3731
Score = 28.1 bits (61), Expect = 7.4
Identities = 28/132 (21%), Positives = 49/132 (36%)
Frame = -2
Query: 22 IFCVINTINYVDRGAIASNGVNGALPTCTDDSGVCTAGSGIQGDFNLNNFQDGVLSSAFM 81
I ++ + D+ S+ + A + D+GVC+ S + G + S F
Sbjct: 2044 IVALLEPSSEADKLLFVSDAIATASSSTCADTGVCSCTSSVAGLSGEGT*LFSSVKSLFK 1865
Query: 82 VGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFDFWSITICRMLVGVGEASF 141
+ SP P +L + L T A G+S ++ + V G S
Sbjct: 1864 LSSGDKSPF--------CPVKLASISLFSGTAATLSSGVS*GAGALNVISDSVTTGPGST 1709
Query: 142 ISLAAPFIDDNA 153
SL+A + N+
Sbjct: 1708 ASLSASVVATNS 1673
>TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 938
Score = 28.1 bits (61), Expect = 7.4
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +2
Query: 322 IFCLVAFLFKNLSGFIVFFS 341
++CL+ FLFK +SGF+ S
Sbjct: 230 VYCLLLFLFKTVSGFVAELS 289
>TC86546 weakly similar to PIR|T45944|T45944 laccase-like protein -
Arabidopsis thaliana, partial (64%)
Length = 2066
Score = 28.1 bits (61), Expect = 7.4
Identities = 18/76 (23%), Positives = 35/76 (45%)
Frame = +1
Query: 254 FVIGAYSYWGPKAGYSIYHMSNPDLLFGGITIVCGILGTLGGGLILDRMTSTISNAFKLL 313
F++ + +W +G+SI + N +++ + +L GL+LD
Sbjct: 1456 FMV*IFMFWRKTSGFSILLLINSSIIW*ILQFATRLLCRWEDGLLLD------------- 1596
Query: 314 SGATLLGAIFCLVAFL 329
S T+LG FC+V ++
Sbjct: 1597 SEQTILGYGFCIVMWM 1644
>AW585735 similar to GP|13958313|gb| NADPH oxidase 1 {Podospora anserina},
partial (4%)
Length = 496
Score = 27.7 bits (60), Expect = 9.7
Identities = 25/91 (27%), Positives = 42/91 (45%), Gaps = 3/91 (3%)
Frame = +1
Query: 82 VGLLIASPIFASLAKSHSPFRLIGVGLSVWTFAVAGCGISFD--FWSITICRMLVGVGEA 139
+GL I F H L +GLSVW+ AG +++D + +CR L+ +
Sbjct: 142 IGLFIYG--FLKQKTDHELDNLNVIGLSVWSSRGAGLCLAYDGALILLPVCRNLIKQFRS 315
Query: 140 -SFISLAAPFIDDNAPVAQKTAWLATFYMCI 169
F++ PF D+N ++TA+ + I
Sbjct: 316 FRFLNNIIPF-DENLWFHRQTAYAMLIFTLI 405
>TC77643 similar to GP|21105742|gb|AAM34770.1 nam-like protein 7 {Petunia x
hybrida}, partial (27%)
Length = 1611
Score = 27.7 bits (60), Expect = 9.7
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = +1
Query: 322 IFCLVAFLFKNLSGFIVFFSIGELLIFATQAPV 354
+FC +LFK L F++ F GE F + +
Sbjct: 325 VFCYTTYLFKKLDFFVLLFLWGEKFDFLIRVSI 423
>TC76894 homologue to SP|P35100|CLPA_PEA ATP-dependent clp protease
ATP-binding subunit clpA homolog chloroplast precursor.
[Garden pea], partial (71%)
Length = 2413
Score = 27.7 bits (60), Expect = 9.7
Identities = 15/31 (48%), Positives = 21/31 (67%)
Frame = -2
Query: 189 TISSIETNVSEHGDDGVLAQDQAFIRGTRST 219
T+SSI TN+SEH +LA+ Q I+ +ST
Sbjct: 444 TLSSISTNLSEHLRMKILARSQH*IQRIQST 352
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,817,723
Number of Sequences: 36976
Number of extensions: 216602
Number of successful extensions: 991
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 982
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 988
length of query: 460
length of database: 9,014,727
effective HSP length: 99
effective length of query: 361
effective length of database: 5,354,103
effective search space: 1932831183
effective search space used: 1932831183
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0381.8


