

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
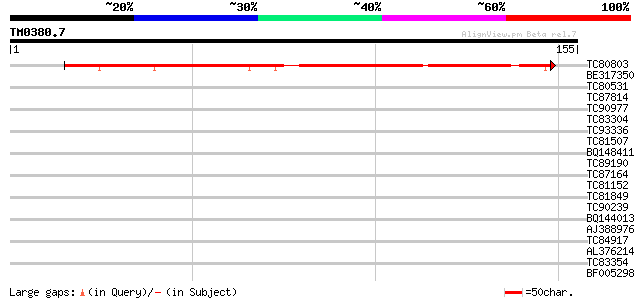
Query= TM0380.7
(155 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC80803 similar to GP|16323284|gb|AAL15397.1 AT4g35320/F23E12_12... 117 2e-27
BE317350 31 0.23
TC80531 similar to GP|3426042|gb|AAC32241.1| unknown protein {Ar... 30 0.30
TC87814 similar to PIR|E96663|E96663 probable RING zinc finger p... 29 0.67
TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {... 29 0.67
TC83304 homologue to GP|19386835|dbj|BAB86213. putative deoxygua... 29 0.87
TC93336 weakly similar to GP|10092180|gb|AAG12599.1 hypothetical... 29 0.87
TC81507 similar to GP|22831146|dbj|BAC16007. putative pre-mRNA s... 28 1.1
BQ148411 similar to GP|18491247|gb| At2g44600/F16B22.9 {Arabidop... 24 1.5
TC89190 28 1.5
TC87164 weakly similar to GP|21617895|gb|AAM66945.1 unknown {Ara... 28 1.5
TC81152 similar to GP|4741991|gb|AAD28791.1| calcium/calmodulin-... 28 1.5
TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {A... 28 1.5
TC90239 weakly similar to PIR|T00583|T00583 probable indole-3-ac... 28 1.9
BQ144013 similar to GP|16323087|gb| AT3g01490/F4P13_4 {Arabidops... 27 2.5
AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing fact... 27 2.5
TC84917 GP|12851094|dbj|BAB28940. evidence:NAS~hypothetical prot... 27 3.3
AL376214 homologue to GP|3204132|emb| extensin {Cicer arietinum}... 27 3.3
TC83354 similar to GP|6227001|gb|AAF06037.1| Contains 3 PF|00400... 27 3.3
BF005298 27 3.3
>TC80803 similar to GP|16323284|gb|AAL15397.1 AT4g35320/F23E12_120
{Arabidopsis thaliana}, partial (27%)
Length = 827
Score = 117 bits (293), Expect = 2e-27
Identities = 72/149 (48%), Positives = 98/149 (65%), Gaps = 15/149 (10%)
Frame = +1
Query: 16 PTTSNNKQ--LPVSHSRSVDSSEHG-------LRKRLSSFNTTPAAASASASAVWSSF-P 65
PT+ + K+ LPVS + + +R+RLS+ + ++ + + WSSF P
Sbjct: 94 PTSRSGKKGSLPVSTKTKLGDAADAVAAGGGFIRRRLSTSSLKNNTSTETTNNKWSSFIP 273
Query: 66 RSKSLS--SMGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWR 123
RSKSLS S+GD SI+NWW W+WAWILSRKPIFA DLE+N+QETKLL S ++G+W+
Sbjct: 274 RSKSLSFSSIGD----SIRNWWTWSWAWILSRKPIFATDLEMNEQETKLL-TSRDKGSWK 438
Query: 124 HVFYKLRSEIRRFIPPTNSHHH---LPRT 149
HV ++RSEIRR I TNS H+ LP+T
Sbjct: 439 HVICRMRSEIRRRI--TNSDHYYTPLPQT 519
>BE317350
Length = 631
Score = 30.8 bits (68), Expect = 0.23
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = +3
Query: 116 SHNRGTWRHVFYKLRSEIRRFIPPTNSHHHLPRTT 150
SH+ R + Y++ S IRR +PP+ SHH +P T
Sbjct: 180 SHHHSLQRPIPYQI-SLIRRSLPPSLSHHRIPLPT 281
>TC80531 similar to GP|3426042|gb|AAC32241.1| unknown protein {Arabidopsis
thaliana}, partial (27%)
Length = 999
Score = 30.4 bits (67), Expect = 0.30
Identities = 9/19 (47%), Positives = 12/19 (62%)
Frame = -3
Query: 78 GTSIKNWWDWTWAWILSRK 96
GT I +W W WAW+ R+
Sbjct: 202 GTMISGFWVWVWAWVWERE 146
>TC87814 similar to PIR|E96663|E96663 probable RING zinc finger protein
T12P18.14 [imported] - Arabidopsis thaliana, partial
(31%)
Length = 819
Score = 29.3 bits (64), Expect = 0.67
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 9/58 (15%)
Frame = +2
Query: 43 LSSFNTTPAAASASASAVWSSFPRS--------KSLSSMGDYAGTSIKNW-WDWTWAW 91
LSS N+ P+++ S S S F R SS Y S ++W W W+W W
Sbjct: 248 LSSKNSLPSSSFTSVSIPASPFRRVCLNFNQLIPCCSSDKSYRW*SSRSWSWSWSWRW 421
>TC90977 similar to GP|10178227|dbj|BAB11607. ZW10-like protein {Arabidopsis
thaliana}, partial (84%)
Length = 779
Score = 29.3 bits (64), Expect = 0.67
Identities = 14/42 (33%), Positives = 25/42 (59%)
Frame = -2
Query: 47 NTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWWDWT 88
N+TP AAS+S ++ SF ++ S S+ + + +NW+ T
Sbjct: 472 NSTPLAASSSFVSLTLSFHQTSSSLSISEKSTAGKRNWYSLT 347
>TC83304 homologue to GP|19386835|dbj|BAB86213. putative deoxyguanosine
kinase {Oryza sativa (japonica cultivar-group)}, partial
(15%)
Length = 919
Score = 28.9 bits (63), Expect = 0.87
Identities = 27/77 (35%), Positives = 42/77 (54%), Gaps = 5/77 (6%)
Frame = +2
Query: 1 MSSIGHQQHRSFFTIPTTSNNKQ----LPVSHSRSVDSSEHGLRKRLSSFNT-TPAAASA 55
M + H+ S IP S++ L +S+S S +S+ L L F++ T AAA++
Sbjct: 68 MHKLLHRPTSSSLPIPILSSSSSSYSPLLISNSNS-NSTPLLLSLPLKPFHSLTMAAATS 244
Query: 56 SASAVWSSFPRSKSLSS 72
S+++ SFP KSLSS
Sbjct: 245 SSTSSGVSFPLQKSLSS 295
>TC93336 weakly similar to GP|10092180|gb|AAG12599.1 hypothetical protein
3' partial; 20361-22062 {Arabidopsis thaliana}, partial
(16%)
Length = 855
Score = 28.9 bits (63), Expect = 0.87
Identities = 23/58 (39%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Frame = +3
Query: 20 NNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSS------FPRSKSLS 71
NNK P+SHS+S S H K +S N P + S +SA +S FP + SLS
Sbjct: 45 NNKTHPLSHSKS--SPVHATEKPTNS-NLDPPSKSVVSSAATTSPSKIQVFPPNTSLS 209
>TC81507 similar to GP|22831146|dbj|BAC16007. putative pre-mRNA splicing
factor SF2 (SR1 protein) {Oryza sativa (japonica
cultivar-group)}, partial (57%)
Length = 1287
Score = 28.5 bits (62), Expect = 1.1
Identities = 25/67 (37%), Positives = 29/67 (42%)
Frame = +3
Query: 5 GHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSF 64
G RS P S ++ S SRS+ S R L N +P SAS S S
Sbjct: 792 GKSSQRSPAKSPAKSVSRSRSRSRSRSLSGSRSRSRSPLPLRNKSPKKRSASRSPS-RSR 968
Query: 65 PRSKSLS 71
RSKSLS
Sbjct: 969 SRSKSLS 989
>BQ148411 similar to GP|18491247|gb| At2g44600/F16B22.9 {Arabidopsis
thaliana}, partial (7%)
Length = 659
Score = 24.3 bits (51), Expect(2) = 1.5
Identities = 9/29 (31%), Positives = 12/29 (41%)
Frame = -1
Query: 85 WDWTWAWILSRKPIFARDLELNDQETKLL 113
W W WAW + K + R Q +L
Sbjct: 125 WSWVWAWTWTSKISWRRSRRQEAQTPTVL 39
Score = 22.3 bits (46), Expect(2) = 1.5
Identities = 7/21 (33%), Positives = 9/21 (42%)
Frame = -1
Query: 71 SSMGDYAGTSIKNWWDWTWAW 91
S+ D S W W+W W
Sbjct: 173 SACNDGRFFSFSGGWAWSWVW 111
>TC89190
Length = 722
Score = 28.1 bits (61), Expect = 1.5
Identities = 6/10 (60%), Positives = 8/10 (80%)
Frame = +3
Query: 82 KNWWDWTWAW 91
K+WW+W W W
Sbjct: 291 KSWWNWNWKW 320
Score = 27.3 bits (59), Expect = 2.5
Identities = 11/24 (45%), Positives = 12/24 (49%), Gaps = 1/24 (4%)
Frame = +3
Query: 76 YAGTSIKNW-WDWTWAWILSRKPI 98
Y S NW W W W WIL+ I
Sbjct: 282 YPWKSWWNWNWKWNWCWILTTTSI 353
>TC87164 weakly similar to GP|21617895|gb|AAM66945.1 unknown {Arabidopsis
thaliana}, partial (68%)
Length = 861
Score = 28.1 bits (61), Expect = 1.5
Identities = 21/63 (33%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Frame = +3
Query: 11 SFFTIPTTSNNKQLP-VSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKS 69
S + S++ +P SH S S LSS TTP++ S+S+S+V+S S
Sbjct: 186 SVLAVHGNSSSTSVPSTSHPTSTTLSPVSKPTSLSSR*TTPSSFSSSSSSVFSGIQSHSS 365
Query: 70 LSS 72
SS
Sbjct: 366 SSS 374
>TC81152 similar to GP|4741991|gb|AAD28791.1| calcium/calmodulin-dependent
protein kinase {Nicotiana tabacum}, partial (49%)
Length = 783
Score = 28.1 bits (61), Expect = 1.5
Identities = 17/38 (44%), Positives = 23/38 (59%), Gaps = 3/38 (7%)
Frame = +3
Query: 42 RLSSFNTTPAAASASASAVWSS--FPRSKSLSSM-GDY 76
RL SFN +A+ ++VWSS F R+K L S+ G Y
Sbjct: 318 RLQSFNARRKLRAAAIASVWSSTIFLRTKKLKSLVGSY 431
>TC81849 similar to GP|19698885|gb|AAL91178.1 putative protein {Arabidopsis
thaliana}, partial (16%)
Length = 1477
Score = 28.1 bits (61), Expect = 1.5
Identities = 17/63 (26%), Positives = 28/63 (43%)
Frame = +1
Query: 1 MSSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV 60
+ G Q T T N LPV D++EHG +++S A++ +S S+
Sbjct: 490 IGGFGEQVQLESPTSTTPDRNNGLPVQSRTQTDNNEHGGSQQVSRLLEQGASSFSSLSSA 669
Query: 61 WSS 63
+S
Sbjct: 670 LNS 678
Score = 25.4 bits (54), Expect = 9.6
Identities = 13/33 (39%), Positives = 16/33 (48%)
Frame = +3
Query: 65 PRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKP 97
PRS S Y+ S NWW W + + RKP
Sbjct: 435 PRSIFFSYCPKYSAVSEPNWWIWRASSV--RKP 527
>TC90239 weakly similar to PIR|T00583|T00583 probable indole-3-acetate
beta-glucosyltransferase T27E13.11 - Arabidopsis
thaliana, partial (30%)
Length = 916
Score = 27.7 bits (60), Expect = 1.9
Identities = 8/18 (44%), Positives = 9/18 (49%)
Frame = -1
Query: 74 GDYAGTSIKNWWDWTWAW 91
G Y G + WW W W W
Sbjct: 67 GYYVGGAGGGWWRWRWIW 14
>BQ144013 similar to GP|16323087|gb| AT3g01490/F4P13_4 {Arabidopsis
thaliana}, partial (22%)
Length = 726
Score = 27.3 bits (59), Expect = 2.5
Identities = 9/28 (32%), Positives = 15/28 (53%)
Frame = +1
Query: 76 YAGTSIKNWWDWTWAWILSRKPIFARDL 103
Y+ W+++ W W+L P+FA L
Sbjct: 469 YSCLFFSTWFEFWWVWVLISMPMFAFQL 552
>AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing factor
[imported] - Arabidopsis thaliana, partial (62%)
Length = 508
Score = 27.3 bits (59), Expect = 2.5
Identities = 8/31 (25%), Positives = 15/31 (47%)
Frame = +1
Query: 61 WSSFPRSKSLSSMGDYAGTSIKNWWDWTWAW 91
W + S++ S + ++ + WW W W W
Sbjct: 214 W*KWLESRAFSQL*NWRWRGRQRWWRWWWWW 306
>TC84917 GP|12851094|dbj|BAB28940. evidence:NAS~hypothetical
protein~putative {Mus musculus}, partial (10%)
Length = 503
Score = 26.9 bits (58), Expect = 3.3
Identities = 8/17 (47%), Positives = 10/17 (58%)
Frame = +2
Query: 80 SIKNWWDWTWAWILSRK 96
S K W WTW+W S +
Sbjct: 263 STKETWTWTWSWAWSER 313
>AL376214 homologue to GP|3204132|emb| extensin {Cicer arietinum}, partial
(48%)
Length = 362
Score = 26.9 bits (58), Expect = 3.3
Identities = 6/11 (54%), Positives = 7/11 (63%)
Frame = -2
Query: 81 IKNWWDWTWAW 91
I+ WW W W W
Sbjct: 73 IRRWWGWRWRW 41
>TC83354 similar to GP|6227001|gb|AAF06037.1| Contains 3 PF|00400 WD40
G-beta repeat domains. {Arabidopsis thaliana}, partial
(41%)
Length = 531
Score = 26.9 bits (58), Expect = 3.3
Identities = 16/45 (35%), Positives = 21/45 (46%), Gaps = 8/45 (17%)
Frame = -2
Query: 116 SHNRGTWRHVFYKLRSEIRRFI--------PPTNSHHHLPRTTPQ 152
SH + RH+ +S RRF+ PPTN HH T P+
Sbjct: 410 SHKKRITRHI----KSHNRRFLIQSTTKASPPTNKIHHSKTTIPR 288
>BF005298
Length = 512
Score = 26.9 bits (58), Expect = 3.3
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = +1
Query: 105 LNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTNSHH 144
L + L ++ T RH L +RR+ PP +SHH
Sbjct: 172 LQSHQPAFLHKLRSQTTTRHFLPYLLRTLRRYFPP*SSHH 291
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.127 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,197,480
Number of Sequences: 36976
Number of extensions: 94813
Number of successful extensions: 1288
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 1163
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1252
length of query: 155
length of database: 9,014,727
effective HSP length: 88
effective length of query: 67
effective length of database: 5,760,839
effective search space: 385976213
effective search space used: 385976213
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0380.7


