

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0377b.5
(372 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
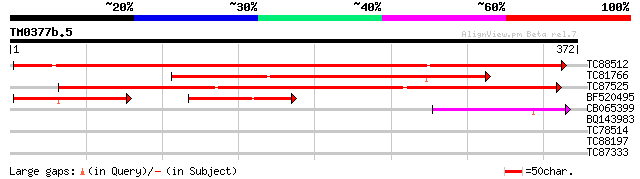
Score E
Sequences producing significant alignments: (bits) Value
TC88512 weakly similar to PIR|T01933|T01933 probable aldose 1-ep... 565 e-162
TC81766 weakly similar to GP|15824565|gb|AAL09397.1 plasmodesmal... 344 3e-95
TC87525 similar to GP|9294498|dbj|BAB02717.1 aldose 1-epimerase-... 332 1e-91
BF520495 weakly similar to PIR|T01933|T019 probable aldose 1-epi... 120 7e-48
CB065399 OMNI|NTL01EC00 galactose-1-epimerase (mutarotase) {Esch... 54 7e-08
BQ143983 26 0.20
TC78514 homologue to SP|Q9SMJ3|ZDS_CAPAN Zeta-carotene desaturas... 29 3.3
TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-act... 28 4.4
TC87333 similar to SP|P49972|SR52_LYCES Signal recognition parti... 28 7.4
>TC88512 weakly similar to PIR|T01933|T01933 probable aldose 1-epimerase (EC
5.1.3.3) - common tobacco, partial (76%)
Length = 1323
Score = 565 bits (1457), Expect = e-162
Identities = 274/363 (75%), Positives = 311/363 (85%)
Frame = +3
Query: 3 TKIFVLLCLLFLAASGFVNGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHG 62
TKIFVLL L + A GFVNGS +KEK + +FELKKG LSLKV+NWGA++VSLVLPDK+G
Sbjct: 120 TKIFVLLLCLLIVAFGFVNGSDEKEK--VGIFELKKGGLSLKVTNWGASIVSLVLPDKNG 293
Query: 63 KLGDVVLGYDSLKQYTNDSTYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGP 122
KL D+VLGYDS+K+YTNDSTYFG T GRVANRIGGAQF LNG HYKL+ANEGNNTLHGG
Sbjct: 294 KLADIVLGYDSIKEYTNDSTYFGATVGRVANRIGGAQFNLNGKHYKLVANEGNNTLHGGS 473
Query: 123 RGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAK 182
RGFSDV+WKV+RY KEG P +TF+YHSFDGEEGFPGDLL TVSYIL KN L IIMKAK
Sbjct: 474 RGFSDVIWKVKRYQKEGPSPSVTFTYHSFDGEEGFPGDLLATVSYILTGKNQLVIIMKAK 653
Query: 183 ALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYD 242
ALNKPTPVNL NHAYWN+G NSGNIL EV+QIFGS+ T++D LIPTG+ ASVKGT YD
Sbjct: 654 ALNKPTPVNLANHAYWNIGGQNSGNILNEVIQIFGSKTTIVDDKLIPTGKFASVKGTAYD 833
Query: 243 FLKPQVVGARINQLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQ 302
FLKPQ++G+RI+QL + GYDINYVLDG K K IKLAA V DKKSGR+++L+TNAPGLQ
Sbjct: 834 FLKPQIIGSRISQLAQHKGYDINYVLDGEKGK-KIKLAAKVHDKKSGRILELYTNAPGLQ 1010
Query: 303 FYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
FYT N +K+ KGK G+VY+ AGLCLESQ FPDSVNHPNFPS+IVTPEKPYKH ML KFS
Sbjct: 1011FYTGNYIKDVKGKSGYVYKAHAGLCLESQAFPDSVNHPNFPSTIVTPEKPYKHYMLFKFS 1190
Query: 363 TKA 365
TK+
Sbjct: 1191TKS 1199
>TC81766 weakly similar to GP|15824565|gb|AAL09397.1 plasmodesmal receptor
{Nicotiana tabacum}, partial (48%)
Length = 635
Score = 344 bits (883), Expect = 3e-95
Identities = 171/211 (81%), Positives = 190/211 (90%), Gaps = 2/211 (0%)
Frame = +2
Query: 107 YKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVS 166
YKLIANEGNNTLHGGPRGFSDVLWKVE+YV+EGD+P I F YHSFDGEEGFPGDL VTVS
Sbjct: 5 YKLIANEGNNTLHGGPRGFSDVLWKVEKYVREGDQPLIKFFYHSFDGEEGFPGDLKVTVS 184
Query: 167 YILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGEVVQIFGSRITVLDSH 226
YIL KNSL+IIM+AKALNKPTP+NLINHAYWNLGNHNSGNIL EVVQIFGS+ T D++
Sbjct: 185 YIL-RKNSLTIIMQAKALNKPTPINLINHAYWNLGNHNSGNILDEVVQIFGSKFTPFDNN 361
Query: 227 LIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGK--SKGLIKLAAIVM 284
LIPTG+ +SVKGTP DFLKPQ+VGARINQL KTNGY++NYVL+ GK +K +K+AAIVM
Sbjct: 362 LIPTGKFSSVKGTPNDFLKPQIVGARINQLQKTNGYNVNYVLNKGKLENKEGLKVAAIVM 541
Query: 285 DKKSGRVMKLFTNAPGLQFYTANKVKNEKGK 315
DKKSGRVMKL TNAPGLQFY AN VKN+KGK
Sbjct: 542 DKKSGRVMKLSTNAPGLQFYRANCVKNDKGK 634
>TC87525 similar to GP|9294498|dbj|BAB02717.1 aldose 1-epimerase-like
protein {Arabidopsis thaliana}, partial (94%)
Length = 1331
Score = 332 bits (852), Expect = 1e-91
Identities = 170/331 (51%), Positives = 217/331 (65%), Gaps = 1/331 (0%)
Frame = +3
Query: 33 LFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND-STYFGTTAGRV 91
+FEL G + L V+N G T+ S +P K G L DVVLG DS++ Y + YFG GRV
Sbjct: 63 IFELNNGTMQLLVTNLGCTITSFSVPAKDGVLSDVVLGLDSVESYQKGLAPYFGCIVGRV 242
Query: 92 ANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSF 151
ANRI +FTL+G+ Y L N+ NTLHGG GF +W V Y K+G+ P ITF Y S
Sbjct: 243 ANRIKNGKFTLDGVEYSLPLNKAPNTLHGGNVGFDKKVWDVLEY-KKGETPSITFKYDSH 419
Query: 152 DGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGE 211
DGEEG+PGD+ VT +Y L S ++ + M+ A NKPT +NL H YWNL H+SGNIL
Sbjct: 420 DGEEGYPGDITVTATYTLTSSTTMRLDMEGVAKNKPTIINLAQHTYWNLAGHSSGNILDH 599
Query: 212 VVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGG 271
++I + +T +D + +PTGEI VKGTP+DF + +G INQ+ GYD NYVLD G
Sbjct: 600 SIKISANHVTPVDQNTVPTGEIVPVKGTPFDFTSEKRIGDTINQVGL--GYDHNYVLDCG 773
Query: 272 KSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQ 331
+ K ++ AA V D S RV+ L+TNAPG+QFYT N V N GKGG VY AGLCLE+Q
Sbjct: 774 EEKAGLRHAAKVRDPSSSRVLNLWTNAPGVQFYTGNYVDNVTGKGGAVYGKHAGLCLETQ 953
Query: 332 VFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
FPD+VN PNFPS +V P + Y+H ML +FS
Sbjct: 954 GFPDAVNKPNFPSVVVKPGEKYQHSMLFEFS 1046
>BF520495 weakly similar to PIR|T01933|T019 probable aldose 1-epimerase (EC
5.1.3.3) - common tobacco, partial (13%)
Length = 462
Score = 120 bits (300), Expect(2) = 7e-48
Identities = 63/81 (77%), Positives = 71/81 (86%), Gaps = 3/81 (3%)
Frame = +1
Query: 3 TKIFVLLCLLFLAASGFVNGSA-KKEKDH--IRLFELKKGDLSLKVSNWGATLVSLVLPD 59
TKIF+ LCLLFLA SGFVNGS KKEK+H I+LFELKKGDL+LKV+NWGATLVSLVLPD
Sbjct: 22 TKIFMPLCLLFLAFSGFVNGSMNKKEKNHDDIKLFELKKGDLTLKVTNWGATLVSLVLPD 201
Query: 60 KHGKLGDVVLGYDSLKQYTND 80
K+G LGD+VLGYD+ K YT D
Sbjct: 202 KNGNLGDIVLGYDTPKAYTVD 264
Score = 88.6 bits (218), Expect(2) = 7e-48
Identities = 45/71 (63%), Positives = 54/71 (75%)
Frame = +3
Query: 118 LHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSI 177
++ GPRGFSDVLWKVE+YV+EGD+P I F YHSFDGEEGFP + S + K +I
Sbjct: 252 IYCGPRGFSDVLWKVEKYVREGDQPLIKFGYHSFDGEEGFPW-*PQSNSELHSKKKFPTI 428
Query: 178 IMKAKALNKPT 188
IM+AKALNKPT
Sbjct: 429 IMQAKALNKPT 461
>CB065399 OMNI|NTL01EC00 galactose-1-epimerase (mutarotase) {Escherichia coli
K12-MG1655}, partial (25%)
Length = 571
Score = 54.3 bits (129), Expect = 7e-08
Identities = 32/93 (34%), Positives = 46/93 (49%), Gaps = 2/93 (2%)
Frame = -2
Query: 278 KLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQVFPDSV 337
K+AA V +K++T AP LQFY+ N + +G Y GL LES+ PDS
Sbjct: 570 KVAAHVWSADEKLQLKVYTTAPALQFYSGNFLGGTPSRGTEPYADWQGLALESEFLPDSP 391
Query: 338 NHPNF--PSSIVTPEKPYKHVMLLKFSTKAPYA 368
NHP + P + P + Y + +F + YA
Sbjct: 390 NHPEWPQPDCFLRPGEEYSSLTEYQFIAE*CYA 292
>BQ143983
Length = 738
Score = 26.2 bits (56), Expect(2) = 0.20
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +1
Query: 139 GDKPRITFSYHSFDGEEG 156
G+ P ITF Y S +G+EG
Sbjct: 355 GETPSITFKYDSREGDEG 408
Score = 25.4 bits (54), Expect(2) = 0.20
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = +3
Query: 185 NKPTPVNLINHAYWNL 200
N PT +NL + YWNL
Sbjct: 498 NMPTIINLAHFTYWNL 545
>TC78514 homologue to SP|Q9SMJ3|ZDS_CAPAN Zeta-carotene desaturase
chloroplast precursor (EC 1.14.99.30) (Carotene 7
8-desaturase)., partial (88%)
Length = 2063
Score = 28.9 bits (63), Expect = 3.3
Identities = 23/62 (37%), Positives = 28/62 (45%), Gaps = 4/62 (6%)
Frame = -2
Query: 305 TANKVKNEKGKGGF---VYQPRAGL-CLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLK 360
+A K EK G + P A CL+S F +SV P + S VT P MLLK
Sbjct: 1465 SARSAKQEKSASGV*RRLSNPVAFFSCLDSSRFCNSVTQPLYLSCTVTTGTPTSSYMLLK 1286
Query: 361 FS 362
S
Sbjct: 1285 NS 1280
>TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-activated
protein kinase {Vicia faba}, partial (96%)
Length = 1158
Score = 28.5 bits (62), Expect = 4.4
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = -2
Query: 202 NHNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPY 241
NH+ NI G V+ GS+ +LD +G+I + GT Y
Sbjct: 896 NHSFFNISGMVILFAGSKTKILDITSRHSGDI*TKSGTEY 777
>TC87333 similar to SP|P49972|SR52_LYCES Signal recognition particle 54 kDa
protein 2 (SRP54). [Tomato] {Lycopersicon esculentum},
partial (58%)
Length = 1236
Score = 27.7 bits (60), Expect = 7.4
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 1/70 (1%)
Frame = +1
Query: 37 KKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSL-KQYTNDSTYFGTTAGRVANRI 95
KKGD++ N AT +S +LP + K + G SL K ++ G G
Sbjct: 709 KKGDMNSMTRNMNATHMSKILPPQMLKQIGGMGGLQSLMKSMGSNKDMMGMFGG------ 870
Query: 96 GGAQFTLNGI 105
GG Q LNG+
Sbjct: 871 GGEQ*VLNGL 900
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,044,079
Number of Sequences: 36976
Number of extensions: 147391
Number of successful extensions: 707
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 696
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 698
length of query: 372
length of database: 9,014,727
effective HSP length: 98
effective length of query: 274
effective length of database: 5,391,079
effective search space: 1477155646
effective search space used: 1477155646
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0377b.5


