

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
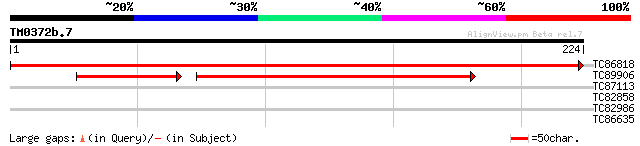
Query= TM0372b.7
(224 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86818 similar to GP|21593515|gb|AAM65482.1 unknown {Arabidopsi... 415 e-117
TC89906 weakly similar to GP|12083292|gb|AAG48805.1 unknown prot... 107 5e-37
TC87113 weakly similar to GP|8649004|emb|CAB94852.1 lipoxygenase... 28 2.2
TC82858 weakly similar to PIR|E96803|E96803 probable lipase 416... 28 2.8
TC82986 similar to GP|21554086|gb|AAM63167.1 unknown {Arabidopsi... 27 8.2
TC86635 weakly similar to GP|9758600|dbj|BAB09233.1 gene_id:MDF2... 27 8.2
>TC86818 similar to GP|21593515|gb|AAM65482.1 unknown {Arabidopsis
thaliana}, partial (80%)
Length = 949
Score = 415 bits (1066), Expect = e-117
Identities = 192/224 (85%), Positives = 208/224 (92%)
Frame = +1
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MVFWVFGYGSLVWNPGF+YDEK++G+IKDYRRVFDLACIDHRGTPE+PARTCTLEEKEGE
Sbjct: 130 MVFWVFGYGSLVWNPGFDYDEKMLGFIKDYRRVFDLACIDHRGTPENPARTCTLEEKEGE 309
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG YCVRGGPEKEKL MQYLERRECEYD KTLV+FYKEGDS HPALTGVIVFTSTPD
Sbjct: 310 ICWGVAYCVRGGPEKEKLAMQYLERRECEYDQKTLVNFYKEGDSLHPALTGVIVFTSTPD 489
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
K NN YYLGPAPL+ MARQIATA+GPCGNNRDYLFLLEKAMHN+GHEDD+VIELA+EVRK
Sbjct: 490 KENNIYYLGPAPLEDMARQIATANGPCGNNRDYLFLLEKAMHNLGHEDDLVIELANEVRK 669
Query: 181 VLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEPIALDS 224
VLGV NV+ N+KKL GP Q+ H PH PIP+LQL P+PEPIALDS
Sbjct: 670 VLGVVNVVLNEKKLVGPTQLPHQPHVPIPSLQLHPLPEPIALDS 801
>TC89906 weakly similar to GP|12083292|gb|AAG48805.1 unknown protein
{Arabidopsis thaliana}, partial (79%)
Length = 739
Score = 107 bits (268), Expect(2) = 5e-37
Identities = 53/109 (48%), Positives = 77/109 (70%)
Frame = +3
Query: 74 EKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVNNKYYLGPAPL 133
E +++ + YL RE +YD K VD + E ++ PA++G +V+ ++PDK N YLGPA +
Sbjct: 141 EDQEIALTYLGVREKQYDRKEYVDVFTELTATTPAISGALVYIASPDKKVNVNYLGPASV 320
Query: 134 DVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVL 182
+ +ARQI A GP G NR+YLFLLEKA+ IG +D VI+LA+EVR++L
Sbjct: 321 EEIARQIVQAEGPTGPNREYLFLLEKALLQIGCQDKHVIDLANEVRRIL 467
Score = 63.5 bits (153), Expect(2) = 5e-37
Identities = 28/41 (68%), Positives = 29/41 (70%)
Frame = +2
Query: 27 IKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICWGAVY 67
IKDYRRVF DHRGTPE P R TLE +GEICWGA Y
Sbjct: 2 IKDYRRVFYQGSTDHRGTPEFPGRAVTLEPAQGEICWGAAY 124
>TC87113 weakly similar to GP|8649004|emb|CAB94852.1 lipoxygenase {Prunus
dulcis}, partial (32%)
Length = 1032
Score = 28.5 bits (62), Expect = 2.2
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +1
Query: 22 KVIGYIKDYRRVFDLACIDHRGTPEH 47
K IG + ++ RV+D AC + GTP++
Sbjct: 625 KGIGKLNEWDRVYDYACYNDLGTPDN 702
>TC82858 weakly similar to PIR|E96803|E96803 probable lipase 4162-5963
[imported] - Arabidopsis thaliana, partial (31%)
Length = 561
Score = 28.1 bits (61), Expect = 2.8
Identities = 9/19 (47%), Positives = 16/19 (83%)
Frame = -3
Query: 74 EKEKLVMQYLERRECEYDH 92
+K++ +QYL+RR+C+Y H
Sbjct: 493 QKQQFSLQYLQRRKCKYCH 437
>TC82986 similar to GP|21554086|gb|AAM63167.1 unknown {Arabidopsis
thaliana}, complete
Length = 747
Score = 26.6 bits (57), Expect = 8.2
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = -1
Query: 116 TSTPDKVNNKYYLGPAPL 133
TS P+++NN++YL PL
Sbjct: 444 TSFPERINNRFYLSMIPL 391
>TC86635 weakly similar to GP|9758600|dbj|BAB09233.1
gene_id:MDF20.10~pir||T08929~similar to unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1963
Score = 26.6 bits (57), Expect = 8.2
Identities = 15/40 (37%), Positives = 23/40 (57%)
Frame = -2
Query: 98 FYKEGDSSHPALTGVIVFTSTPDKVNNKYYLGPAPLDVMA 137
F DSS V++F STP++ + + L P PL+V+A
Sbjct: 924 FSSPSDSSS---VSVVLFFSTPEESSFFFCLFPEPLEVLA 814
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,994,902
Number of Sequences: 36976
Number of extensions: 117443
Number of successful extensions: 578
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 569
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 578
length of query: 224
length of database: 9,014,727
effective HSP length: 93
effective length of query: 131
effective length of database: 5,575,959
effective search space: 730450629
effective search space used: 730450629
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0372b.7


