

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
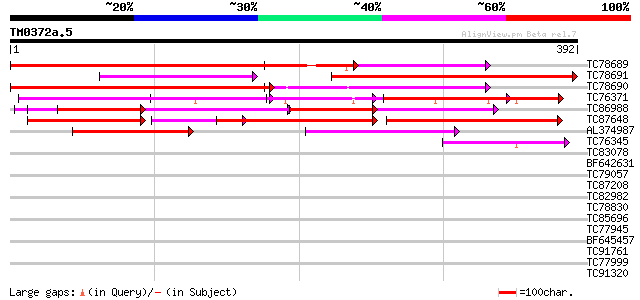
Query= TM0372a.5
(392 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78689 weakly similar to GP|17380978|gb|AAL36301.1 unknown prot... 348 2e-96
TC78691 similar to GP|17380978|gb|AAL36301.1 unknown protein {Ar... 321 3e-88
TC78690 weakly similar to GP|17380978|gb|AAL36301.1 unknown prot... 298 2e-81
TC76371 weakly similar to GP|23304737|emb|CAC87937. PDI-like pro... 171 7e-72
TC86988 weakly similar to GP|17529294|gb|AAL38874.1 unknown prot... 241 4e-64
TC87648 similar to GP|17529294|gb|AAL38874.1 unknown protein {Ar... 124 1e-56
AL374987 weakly similar to GP|21436483|gb| thioredoxin {Brugia m... 85 5e-17
TC76345 similar to GP|3928152|emb|CAA10290.1 ribulose 1 5-bispho... 67 1e-11
TC83078 homologue to GP|23325349|gb|AAN24013.1 very narrowly con... 40 0.001
BF642631 homologue to GP|6560635|gb| thioredoxin-like protein {M... 39 0.005
TC79057 similar to GP|14581677|gb|AAK64512.1 Hsp70 interacting p... 37 0.017
TC87208 similar to SP|Q43636|THIH_RICCO Thioredoxin H-type (TRX-... 35 0.038
TC82982 weakly similar to GP|1388082|gb|AAC49355.1| thioredoxin ... 35 0.050
TC78830 similar to GP|12322718|gb|AAG51342.1 thioredoxin-like pr... 35 0.050
TC85696 similar to SP|Q43636|THIH_RICCO Thioredoxin H-type (TRX-... 33 0.25
TC77945 similar to SP|O24296|GSHY_PEA Phospholipid hydroperoxide... 33 0.25
BF645457 similar to PIR|T52301|T5 GYMNOS/PICKLE protein [importe... 32 0.32
TC91761 similar to GP|21554313|gb|AAM63418.1 thioredoxin-like pr... 32 0.32
TC77999 similar to SP|P29450|THIF_PEA Thioredoxin F-type chloro... 32 0.42
TC91320 similar to GP|22135912|gb|AAM91538.1 unknown protein {Ar... 31 0.94
>TC78689 weakly similar to GP|17380978|gb|AAL36301.1 unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1037
Score = 348 bits (894), Expect = 2e-96
Identities = 172/246 (69%), Positives = 201/246 (80%), Gaps = 5/246 (2%)
Frame = +2
Query: 1 MAGINFEAKHIDSHDVLKILAAEGAEYLLSCEGKVPLSECDGKVICLFFSANWCRPCRAF 60
MAG+NFEA++ DS DVLK+ AAEG E+LLSCE KVPLS+C+GK+ICLFFSANWCRPCR F
Sbjct: 314 MAGLNFEAEYPDSFDVLKVFAAEGVEFLLSCERKVPLSDCNGKIICLFFSANWCRPCRLF 493
Query: 61 VPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQV 120
+P LV LYETLRKRGIN+EIIF+SFD +EDGF EH+KSMPWLAVPFD L+RRLIDRY+V
Sbjct: 494 IPHLVGLYETLRKRGINIEIIFISFDHDEDGFKEHIKSMPWLAVPFDAKLNRRLIDRYRV 673
Query: 121 DQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLA 180
D+IPSFIPL S+ L V+KN+IE IEDYGADAFPFTRKRHEELKAIDKRKREE NL+ELL
Sbjct: 674 DRIPSFIPLCSDALTVDKNMIEWIEDYGADAFPFTRKRHEELKAIDKRKREEVNLDELLT 853
Query: 181 HEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNL-----KATK 235
H GR+FLISGDDRKV VSEL GK G Y+S P+ + ++ + + TK
Sbjct: 854 HGGRNFLISGDDRKVLVSELTGKL*G-----YFSVPTGLLLVMPSTIHLQMHTTISRDTK 1018
Query: 236 GNCFEI 241
G+CFEI
Sbjct: 1019GHCFEI 1036
Score = 100 bits (248), Expect = 1e-21
Identities = 57/156 (36%), Positives = 85/156 (53%)
Frame = +2
Query: 177 ELLAHEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKG 236
++ A EG +FL+S + RKVP+S+ GK I L+F A W P R F L +Y L+ +G
Sbjct: 365 KVFAAEGVEFLLSCE-RKVPLSDCNGKIICLFFSANWCRPCRLFIPHLVGLYETLRK-RG 538
Query: 237 NCFEIVLISTDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKV 296
EI+ IS D D F + S PWLA+P++ + L Y + IP+ + + D
Sbjct: 539 INIEIIFISFDHDEDGFKEHIKSMPWLAVPFDAKLNRRLIDRYRVDRIPSFIPLCSDALT 718
Query: 297 ISLNGKFMVTSYGADAFPFTESRIRDLEAALRKEGE 332
+ N + YGADAFPFT R +L+A +++ E
Sbjct: 719 VDKNMIEWIEDYGADAFPFTRKRHEELKAIDKRKRE 826
>TC78691 similar to GP|17380978|gb|AAL36301.1 unknown protein {Arabidopsis
thaliana}, partial (38%)
Length = 815
Score = 321 bits (822), Expect = 3e-88
Identities = 150/170 (88%), Positives = 160/170 (93%)
Frame = +3
Query: 223 QLTDVYNNLKATKGNCFEIVLISTDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRIYDIK 282
+L D YNNLK TKG+CFEIVL+STDRD KEFNVNR+S PWLAIPYEDRTRHDLCRI+DIK
Sbjct: 6 RLADAYNNLKDTKGHCFEIVLVSTDRDLKEFNVNRTSMPWLAIPYEDRTRHDLCRIFDIK 185
Query: 283 GIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTESRIRDLEAALRKEGEALPQQVEDVK 342
IPALV IGPDGKVISLNG+FMV+SYGA+AFPFTESRIRDLEAALRKEGEALPQQVEDVK
Sbjct: 186 KIPALVFIGPDGKVISLNGQFMVSSYGAEAFPFTESRIRDLEAALRKEGEALPQQVEDVK 365
Query: 343 HEHVLKLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHPSCLEKVNKDQDW 392
HEH+LKLDMAKAYVCDSCK+QGKFW FSCDVCDYDLHPSCLEKVN DQ W
Sbjct: 366 HEHLLKLDMAKAYVCDSCKKQGKFWTFSCDVCDYDLHPSCLEKVNNDQCW 515
Score = 68.6 bits (166), Expect = 4e-12
Identities = 36/110 (32%), Positives = 61/110 (54%), Gaps = 1/110 (0%)
Frame = +3
Query: 63 RLVELYETLRK-RGINLEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQVD 121
RL + Y L+ +G EI+ VS DR+ FN + SMPWLA+P++ L + +
Sbjct: 6 RLADAYNNLKDTKGHCFEIVLVSTDRDLKEFNVNRTSMPWLAIPYEDRTRHDLCRIFDIK 185
Query: 122 QIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKRE 171
+IP+ + + + ++ N + YGA+AFPFT R +L+A +++ E
Sbjct: 186 KIPALVFIGPDGKVISLNGQFMVSSYGAEAFPFTESRIRDLEAALRKEGE 335
>TC78690 weakly similar to GP|17380978|gb|AAL36301.1 unknown protein
{Arabidopsis thaliana}, partial (21%)
Length = 676
Score = 298 bits (763), Expect = 2e-81
Identities = 140/183 (76%), Positives = 160/183 (86%)
Frame = +2
Query: 1 MAGINFEAKHIDSHDVLKILAAEGAEYLLSCEGKVPLSECDGKVICLFFSANWCRPCRAF 60
MAG+NFEA++ DS DVLK+ AAEG E+LLSCE KVPLS+C+GK+ICLFFSANWCRPCR F
Sbjct: 125 MAGLNFEAEYPDSFDVLKVFAAEGVEFLLSCERKVPLSDCNGKIICLFFSANWCRPCRLF 304
Query: 61 VPRLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQV 120
+P LV LYETLRKRGIN+EIIF+SFD +EDGF EH+KSMPWLAVPFD L+RRLIDRY+V
Sbjct: 305 IPHLVGLYETLRKRGINIEIIFISFDHDEDGFKEHIKSMPWLAVPFDAKLNRRLIDRYRV 484
Query: 121 DQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLA 180
D+IPSFIPL S+ L V+KN+IE IEDYGADAFPFTRKRH ELKAI RKR E NL+ELL
Sbjct: 485 DRIPSFIPLCSDALTVDKNMIEWIEDYGADAFPFTRKRHXELKAIXXRKRXEVNLDELLT 664
Query: 181 HEG 183
H G
Sbjct: 665 HGG 673
Score = 99.0 bits (245), Expect = 3e-21
Identities = 59/157 (37%), Positives = 84/157 (52%), Gaps = 1/157 (0%)
Frame = +2
Query: 177 ELLAHEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKG 236
++ A EG +FL+S + RKVP+S+ GK I L+F A W P R F L +Y L+ +G
Sbjct: 176 KVFAAEGVEFLLSCE-RKVPLSDCNGKIICLFFSANWCRPCRLFIPHLVGLYETLRK-RG 349
Query: 237 NCFEIVLISTDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKV 296
EI+ IS D D F + S PWLA+P++ + L Y + IP+ + + D
Sbjct: 350 INIEIIFISFDHDEDGFKEHIKSMPWLAVPFDAKLNRRLIDRYRVDRIPSFIPLCSDALT 529
Query: 297 ISLNGKFMVTSYGADAFPFTESRIRDLEA-ALRKEGE 332
+ N + YGADAFPFT R +L+A RK E
Sbjct: 530 VDKNMIEWIEDYGADAFPFTRKRHXELKAIXXRKRXE 640
>TC76371 weakly similar to GP|23304737|emb|CAC87937. PDI-like protein
{Quercus suber}, partial (21%)
Length = 1710
Score = 171 bits (434), Expect(2) = 7e-72
Identities = 101/263 (38%), Positives = 150/263 (56%), Gaps = 15/263 (5%)
Frame = +2
Query: 7 EAKHIDSHDVLKILAAEGAEYLLSCEG-KVPLSECDGKVICLFFSANWCRPCRAFVPRLV 65
E + + S +LA++ ++LLS G +V +SE +GKV+ L F+ANW PCR F L+
Sbjct: 164 EEEAVSSFKFSYLLASKDRDFLLSSTGTQVKISELEGKVVGLLFAANWYPPCRGFTQLLI 343
Query: 66 ELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPF-DVNLHRRLIDRYQVDQIP 124
+YE L+ EI++VS D + D FN +MPWLA+PF D+ + L +Y V+ IP
Sbjct: 344 GIYEQLKSNIPQFEIVYVSSDEDLDAFNGFYGNMPWLAIPFSDLETKKALNRKYDVEGIP 523
Query: 125 SFI---PLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLAH 181
+ P +S+ ++ +E I YG A+PF+++R E+L ++ K E L LLA+
Sbjct: 524 CLVMLQPDHSKGEATLRDGVELIYRYGVQAYPFSKERLEQLHVAEREKLENQTLANLLAN 703
Query: 182 EGRDFLIS----GDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKG- 236
RD+++S G +VPV+ L GKTIGLYF A W P FT +L +VY +K
Sbjct: 704 NHRDYVLSHTGTGLLTQVPVASLVGKTIGLYFSAGWCVPCTKFTPKLINVYQIIKQELAE 883
Query: 237 -----NCFEIVLISTDRDHKEFN 254
FEIVL+S DRD + F+
Sbjct: 884 KQDPHEDFEIVLVSNDRDQESFD 952
Score = 123 bits (309), Expect = 1e-28
Identities = 71/176 (40%), Positives = 101/176 (57%), Gaps = 6/176 (3%)
Frame = +2
Query: 178 LLAHEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGN 237
LLA + RDFL+S +V +SEL GK +GL F A W PP R FT L +Y LK+
Sbjct: 200 LLASKDRDFLLSSTGTQVKISELEGKVVGLLFAANWYPPCRGFTQLLIGIYEQLKSNIPQ 379
Query: 238 CFEIVLISTDRDHKEFNVNRSSTPWLAIPYED-RTRHDLCRIYDIKGIPALVLIGPD--- 293
FEIV +S+D D FN + PWLAIP+ D T+ L R YD++GIP LV++ PD
Sbjct: 380 -FEIVYVSSDEDLDAFNGFYGNMPWLAIPFSDLETKKALNRKYDVEGIPCLVMLQPDHSK 556
Query: 294 GKVISLNGKFMVTSYGADAFPFTESRIRDLEAALRK--EGEALPQQVEDVKHEHVL 347
G+ +G ++ YG A+PF++ R+ L A R+ E + L + + ++VL
Sbjct: 557 GEATLRDGVELIYRYGVQAYPFSKERLEQLHVAEREKLENQTLANLLANNHRDYVL 724
Score = 117 bits (293), Expect(2) = 7e-72
Identities = 53/129 (41%), Positives = 79/129 (61%), Gaps = 4/129 (3%)
Frame = +3
Query: 259 STPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTES 318
S PWLA+P+ D +L R +D++GIP LV+IGPDGK I+++G+ ++ Y +A+PFT S
Sbjct: 966 SCPWLALPFGDPEIKNLARHFDVQGIPCLVIIGPDGKTITIHGRNLINLYQENAYPFTAS 1145
Query: 319 RIRDLEAALRKEGEALPQQVEDVKHEHVLKL----DMAKAYVCDSCKEQGKFWAFSCDVC 374
++ LE L +E + LP V H H L L + ++C C EQG WA+ C C
Sbjct: 1146 KVEQLEKQLEEEAKDLPNLVHHEGHHHGLNLVSDGNGGGPFICCVCDEQGSNWAYQCLQC 1325
Query: 375 DYDLHPSCL 383
Y++HP C+
Sbjct: 1326 GYEVHPKCV 1352
Score = 41.6 bits (96), Expect = 5e-04
Identities = 28/87 (32%), Positives = 44/87 (50%), Gaps = 1/87 (1%)
Frame = +3
Query: 98 SMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRK 157
S PWLA+PF + L + V IP + + + + + I Y +A+PFT
Sbjct: 966 SCPWLALPFGDPEIKNLARHFDVQGIPCLVIIGPDGKTITIHGRNLINLYQENAYPFTAS 1145
Query: 158 RHEELKAIDKRKREEA-NLEELLAHEG 183
+ E+L +K+ EEA +L L+ HEG
Sbjct: 1146 KVEQL---EKQLEEEAKDLPNLVHHEG 1217
>TC86988 weakly similar to GP|17529294|gb|AAL38874.1 unknown protein
{Arabidopsis thaliana}, partial (66%)
Length = 1378
Score = 241 bits (615), Expect = 4e-64
Identities = 126/329 (38%), Positives = 191/329 (57%), Gaps = 3/329 (0%)
Frame = +2
Query: 13 SHDVLKILAAEGAEYLLSCEG-KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETL 71
+H+V IL++ ++LL G +V + GK + +FSA+WC PCR F P+LVE+Y+ L
Sbjct: 59 THNVHSILSSSDRDFLLRNTGDQVKIDSLKGKKLGFYFSASWCGPCRGFTPKLVEVYDEL 238
Query: 72 RKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPF-DVNLHRRLIDRYQVDQIPSFIPLY 130
G E++FVS D++++ F + MPWLA+PF D RL + + V+ IP L
Sbjct: 239 SPNG-EFEVVFVSADKDDEAFKSYFSKMPWLAIPFSDSETRGRLDELFHVNGIPHLALLD 415
Query: 131 SEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLAHEGRDFLISG 190
++ ++ ++ I YGA+A+PFT KR +ELK I++ + +L +LA RDFLIS
Sbjct: 416 EAGKVITEDGVDIIRVYGAEAYPFTSKRVQELKDIEEEAKRNQSLRSILASRSRDFLISS 595
Query: 191 DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDH 250
D ++P+SEL GKT+GL+FCA FT +L +VY LK G FE+V I D +
Sbjct: 596 DGNEIPISELEGKTVGLHFCATSYRACTLFTQKLKEVYKKLK-ENGENFEVVFIPLDDEE 772
Query: 251 KEFNVNRSSTPWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGA 310
F S PWL++P +D+T L + +++ +P LV+IGPDGK + N + +G
Sbjct: 773 DAFKKELESAPWLSLPLKDKTCAKLIQYFELSELPTLVIIGPDGKTLHPNAAEAIEDHGV 952
Query: 311 DAFPFTESRIRDL-EAALRKEGEALPQQV 338
DA+PFT + +L E A KE + V
Sbjct: 953 DAYPFTPEKFSELDEIAKAKEASQTLESV 1039
Score = 132 bits (333), Expect(2) = 7e-47
Identities = 69/193 (35%), Positives = 111/193 (56%), Gaps = 1/193 (0%)
Frame = +2
Query: 4 INFEAKHIDSHDVLKILAAEGAEYLLSCEG-KVPLSECDGKVICLFFSANWCRPCRAFVP 62
I EAK S + ILA+ ++L+S +G ++P+SE +GK + L F A R C F
Sbjct: 518 IEEEAKRNQS--LRSILASRSRDFLISSDGNEIPISELEGKTVGLHFCATSYRACTLFTQ 691
Query: 63 RLVELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQVDQ 122
+L E+Y+ L++ G N E++F+ D EED F + L+S PWL++P +LI +++ +
Sbjct: 692 KLKEVYKKLKENGENFEVVFIPLDDEEDAFKKELESAPWLSLPLKDKTCAKLIQYFELSE 871
Query: 123 IPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLAHE 182
+P+ + + + + N E IED+G DA+PFT ++ EL I K K LE +L
Sbjct: 872 LPTLVIIGPDGKTLHPNAAEAIEDHGVDAYPFTPEKFSELDEIAKAKEASQTLESVLVSG 1051
Query: 183 GRDFLISGDDRKV 195
+DF+I D +KV
Sbjct: 1052DQDFVIDKDGKKV 1090
Score = 73.2 bits (178), Expect = 2e-13
Identities = 29/62 (46%), Positives = 48/62 (76%), Gaps = 1/62 (1%)
Frame = +3
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGIN-LEIIFVSFDREEDGF 92
++P+SE GK + L+FSA+WC PCRAF+P+L+E Y ++ + + LE++F+S DR+++ F
Sbjct: 1191 QIPVSELVGKTVLLYFSAHWCPPCRAFLPKLIEAYHKIKAQNNDALEVVFISSDRDQESF 1370
Query: 93 NE 94
NE
Sbjct: 1371 NE 1376
Score = 72.8 bits (177), Expect(2) = 7e-47
Identities = 31/61 (50%), Positives = 43/61 (69%)
Frame = +3
Query: 194 KVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEF 253
++PVSEL GKT+ LYF A+W PP RAF +L + Y+ +KA + E+V IS+DRD + F
Sbjct: 1191 QIPVSELVGKTVLLYFSAHWCPPCRAFLPKLIEAYHKIKAQNNDALEVVFISSDRDQESF 1370
Query: 254 N 254
N
Sbjct: 1371 N 1373
>TC87648 similar to GP|17529294|gb|AAL38874.1 unknown protein {Arabidopsis
thaliana}, partial (33%)
Length = 990
Score = 124 bits (311), Expect(2) = 1e-56
Identities = 53/122 (43%), Positives = 76/122 (61%)
Frame = +1
Query: 261 PWLAIPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTESRI 320
PWLA+P+ D + L R + + GIP LV IGP G+ ++ + +V YGADA+PFTE RI
Sbjct: 358 PWLALPFGDTRKEFLSRKFKVSGIPKLVAIGPSGQTVTKEARGLVGLYGADAYPFTEKRI 537
Query: 321 RDLEAALRKEGEALPQQVEDVKHEHVLKLDMAKAYVCDSCKEQGKFWAFSCDVCDYDLHP 380
+++EA + P++V HEH L L Y CD CK++G W++ C CD+DLHP
Sbjct: 538 KEIEAQKDDIAKGWPEKVTHETHEHELVLSRRNVYCCDGCKDEGDTWSYLCAECDFDLHP 717
Query: 381 SC 382
+C
Sbjct: 718 NC 723
Score = 114 bits (284), Expect(2) = 1e-56
Identities = 53/111 (47%), Positives = 72/111 (64%)
Frame = +3
Query: 144 IEDYGADAFPFTRKRHEELKAIDKRKREEANLEELLAHEGRDFLISGDDRKVPVSELAGK 203
IED+G DA+PFT ++ EL I K K LE +L +DF+I D +K+PVSEL GK
Sbjct: 6 IEDHGVDAYPFTPEKFSELDEIAKAKEASQTLESVLVSGDQDFVIDKDGKKIPVSELVGK 185
Query: 204 TIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEFN 254
T+ LYF A+W PP RAF +L + Y+ +KA + E+V IS+DRD + FN
Sbjct: 186 TVLLYFSAHWCPPCRAFLPKLIEAYHKIKAQNNDALEVVFISSDRDQESFN 338
Score = 77.0 bits (188), Expect(2) = 3e-23
Identities = 33/84 (39%), Positives = 59/84 (69%), Gaps = 2/84 (2%)
Frame = +3
Query: 13 SHDVLKILAAEGAEYLLSCEGK-VPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETL 71
S + +L + ++++ +GK +P+SE GK + L+FSA+WC PCRAF+P+L+E Y +
Sbjct: 90 SQTLESVLVSGDQDFVIDKDGKKIPVSELVGKTVLLYFSAHWCPPCRAFLPKLIEAYHKI 269
Query: 72 RKRGIN-LEIIFVSFDREEDGFNE 94
+ + + LE++F+S DR+++ FNE
Sbjct: 270 KAQNNDALEVVFISSDRDQESFNE 341
Score = 49.3 bits (116), Expect(2) = 3e-23
Identities = 25/66 (37%), Positives = 36/66 (53%)
Frame = +1
Query: 99 MPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIEDYGADAFPFTRKR 158
MPWLA+PF L +++V IP + + V K + YGADA+PFT KR
Sbjct: 355 MPWLALPFGDTRKEFLSRKFKVSGIPKLVAIGPSGQTVTKEARGLVGLYGADAYPFTEKR 534
Query: 159 HEELKA 164
+E++A
Sbjct: 535 IKEIEA 552
>AL374987 weakly similar to GP|21436483|gb| thioredoxin {Brugia malayi},
partial (46%)
Length = 473
Score = 84.7 bits (208), Expect = 5e-17
Identities = 40/107 (37%), Positives = 62/107 (57%)
Frame = -3
Query: 205 IGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCFEIVLISTDRDHKEFNVNRSSTPWLA 264
I +YF A+W PP R FT L + Y ++A N EI+ +S+D + +F+ + PW A
Sbjct: 429 IAIYFSAHWCPPCRQFTPILGEFYKEIQANHPNDLEIIFVSSDSNPPQFDEYFKTMPWTA 250
Query: 265 IPYEDRTRHDLCRIYDIKGIPALVLIGPDGKVISLNGKFMVTSYGAD 311
+P+ D +L + + + GIP LV+ P+G VIS G+ VT GA+
Sbjct: 249 VPFGDPIIQNLKQKFSVSGIPFLVVCSPNGDVISTTGRADVTGQGAE 109
Score = 75.5 bits (184), Expect = 3e-14
Identities = 37/85 (43%), Positives = 52/85 (60%), Gaps = 1/85 (1%)
Frame = -3
Query: 44 VICLFFSANWCRPCRAFVPRLVELYETLRKRGIN-LEIIFVSFDREEDGFNEHLKSMPWL 102
+I ++FSA+WC PCR F P L E Y+ ++ N LEIIFVS D F+E+ K+MPW
Sbjct: 432 LIAIYFSAHWCPPCRQFTPILGEFYKEIQANHPNDLEIIFVSSDSNPPQFDEYFKTMPWT 253
Query: 103 AVPFDVNLHRRLIDRYQVDQIPSFI 127
AVPF + + L ++ V IP +
Sbjct: 252 AVPFGDPIIQNLKQKFSVSGIPFLV 178
>TC76345 similar to GP|3928152|emb|CAA10290.1 ribulose 1 5-bisphosphate
carboxylase small subunit {Cicer arietinum}, partial
(98%)
Length = 1243
Score = 67.0 bits (162), Expect = 1e-11
Identities = 31/92 (33%), Positives = 47/92 (50%), Gaps = 4/92 (4%)
Frame = +1
Query: 300 NGKFMVTSYGADAFPFTESRIRDLEAALRKEGEALPQQVEDVKHEHVLKL----DMAKAY 355
N + ++ Y +A+PFT S++ LE L + LP V H H L L + +
Sbjct: 841 NSRNLINLYQXNAYPFTASKVEQLEKQLEXXAKDLPNLVHHEGHHHGLNLVSDGNGGGPF 1020
Query: 356 VCDSCKEQGKFWAFSCDVCDYDLHPSCLEKVN 387
+C C EQG WA+ C C Y++HP C+ V+
Sbjct: 1021ICCVCDEQGSNWAYQCLQCGYEVHPKCVTTVH 1116
>TC83078 homologue to GP|23325349|gb|AAN24013.1 very narrowly conserved
hypothetical protein {Bifidobacterium longum NCC2705},
partial (0%)
Length = 1167
Score = 40.4 bits (93), Expect = 0.001
Identities = 29/110 (26%), Positives = 51/110 (46%), Gaps = 14/110 (12%)
Frame = +3
Query: 224 LTDVYNNLKATKGNCFEIVLIS-----TDRDHKEFNVNRSSTPWLAIPYEDRTRHDLCRI 278
L +Y+++K T G+ +IV + D+ K+F+ +S PW + H I
Sbjct: 276 LIPIYDHIKKT-GSQHKIVWVPIVEEWNDKLKKKFDSLKSKMPWYVL-------HHFAPI 431
Query: 279 YDIKGI---------PALVLIGPDGKVISLNGKFMVTSYGADAFPFTESR 319
IK I P V++ P GK++ N M+ +G FP+++S+
Sbjct: 432 KGIKYIKEELHFKQKPLFVVLSPQGKILHHNAFHMIXVWGVKGFPYSKSK 581
>BF642631 homologue to GP|6560635|gb| thioredoxin-like protein {Manduca
sexta}, complete
Length = 518
Score = 38.5 bits (88), Expect = 0.005
Identities = 22/68 (32%), Positives = 35/68 (51%), Gaps = 1/68 (1%)
Frame = +3
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREEDGFN 93
K LSE K++ + F A WC PC+ P+L E+ + ++ ++ V D ED
Sbjct: 78 KTRLSEAGDKLVVIDFMATWCGPCKMIGPKLDEMANEMAD---SIVVLKVDVDECEDIAT 248
Query: 94 EH-LKSMP 100
E+ + SMP
Sbjct: 249 EYNINSMP 272
>TC79057 similar to GP|14581677|gb|AAK64512.1 Hsp70 interacting
protein/thioredoxin chimera {Vitis labrusca}, partial
(84%)
Length = 1335
Score = 36.6 bits (83), Expect = 0.017
Identities = 19/69 (27%), Positives = 32/69 (45%)
Frame = +1
Query: 7 EAKHIDSHDVLKILAAEGAEYLLSCEGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVE 66
+A+ D+ LK G + E K+ + +++ L+F+A WC PCR P
Sbjct: 769 QAQDKDALSALKDGQVIGVHSVGELETKLSAASKTSRLLVLYFTATWCGPCRYISPLYTS 948
Query: 67 LYETLRKRG 75
L E ++ G
Sbjct: 949 LAEKYQRSG 975
>TC87208 similar to SP|Q43636|THIH_RICCO Thioredoxin H-type (TRX-H). [Castor
bean] {Ricinus communis}, partial (94%)
Length = 682
Score = 35.4 bits (80), Expect = 0.038
Identities = 17/47 (36%), Positives = 27/47 (57%)
Frame = +3
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+I + F+A+WC PCR P L E+ + + E+IF+ D +E
Sbjct: 129 KLIVVDFTASWCGPCRFIAPILAEIAKKIP------EVIFLKVDIDE 251
>TC82982 weakly similar to GP|1388082|gb|AAC49355.1| thioredoxin h
{Arabidopsis thaliana}, partial (38%)
Length = 481
Score = 35.0 bits (79), Expect = 0.050
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +1
Query: 42 GKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEI 80
GK++ + F+A+WC PC+ P L E +K I L++
Sbjct: 85 GKLVIVDFTASWCPPCKMIAPIFKNLAEEYKKVAIFLKV 201
>TC78830 similar to GP|12322718|gb|AAG51342.1 thioredoxin-like protein;
56513-57227 {Arabidopsis thaliana}, partial (88%)
Length = 975
Score = 35.0 bits (79), Expect = 0.050
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = +3
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYE 69
K+ ++ DGK++ FSA WC PC+ P E+ E
Sbjct: 321 KLEEAKKDGKIVIANFSAVWCGPCKVIAPYYCEMSE 428
>TC85696 similar to SP|Q43636|THIH_RICCO Thioredoxin H-type (TRX-H). [Castor
bean] {Ricinus communis}, partial (94%)
Length = 714
Score = 32.7 bits (73), Expect = 0.25
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +2
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETL 71
K+I + F+A+WC PCR P L E+ + L
Sbjct: 185 KLIVVDFTASWCGPCRFIAPILAEIAKKL 271
>TC77945 similar to SP|O24296|GSHY_PEA Phospholipid hydroperoxide
glutathione peroxidase chloroplast precursor (EC
1.11.1.9) (PHGPx)., complete
Length = 1006
Score = 32.7 bits (73), Expect = 0.25
Identities = 21/64 (32%), Positives = 31/64 (47%), Gaps = 1/64 (1%)
Frame = +1
Query: 35 VPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGIN-LEIIFVSFDREEDGFN 93
VPLS+ GKV+ + A+ C + L LYE + +G+ L F +E G N
Sbjct: 361 VPLSKFKGKVLLIVNVASRCGLTSSNYTELSHLYENFKDKGLEVLAFPCNQFGMQEPGSN 540
Query: 94 EHLK 97
E +K
Sbjct: 541 EEIK 552
>BF645457 similar to PIR|T52301|T5 GYMNOS/PICKLE protein [imported] -
Arabidopsis thaliana, partial (9%)
Length = 560
Score = 32.3 bits (72), Expect = 0.32
Identities = 28/89 (31%), Positives = 38/89 (42%)
Frame = +2
Query: 295 KVISLNGKFMVTSYGADAFPFTESRIRDLEAALRKEGEALPQQVEDVKHEHVLKLDMAKA 354
K+ SL + V S + ES D L+K G + K E + + D AK
Sbjct: 95 KMSSLVERLRVRSDRKPVYNLDESDDEDF--LLKKPGAS------QEKFERIDRSD-AKE 247
Query: 355 YVCDSCKEQGKFWAFSCDVCDYDLHPSCL 383
+C +C E G SC C+Y H SCL
Sbjct: 248 DLCQACGESGDL--LSCATCNYAYHSSCL 328
>TC91761 similar to GP|21554313|gb|AAM63418.1 thioredoxin-like protein
{Arabidopsis thaliana}, partial (55%)
Length = 613
Score = 32.3 bits (72), Expect = 0.32
Identities = 16/54 (29%), Positives = 28/54 (51%)
Frame = +2
Query: 37 LSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED 90
LS+ + +++ + F WC CRA P+L E + EIIF+ + +E+
Sbjct: 311 LSQAEDRLVIVEFYGTWCASCRALFPKLCRTAEE------HPEIIFLKVNFDEN 454
>TC77999 similar to SP|P29450|THIF_PEA Thioredoxin F-type chloroplast
precursor (TRX-F). [Garden pea] {Pisum sativum},
complete
Length = 835
Score = 32.0 bits (71), Expect = 0.42
Identities = 14/48 (29%), Positives = 22/48 (45%)
Frame = +2
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED 90
K + L WC PC+ P+ EL E L+++F+ D +D
Sbjct: 311 KTVVLDMYTQWCGPCKVIAPKYKELAEKY------LDVVFLKLDCNQD 436
>TC91320 similar to GP|22135912|gb|AAM91538.1 unknown protein {Arabidopsis
thaliana}, partial (66%)
Length = 1372
Score = 30.8 bits (68), Expect = 0.94
Identities = 49/206 (23%), Positives = 75/206 (35%), Gaps = 20/206 (9%)
Frame = -3
Query: 88 EEDGFNEHLKSMPWLAVPFDVNLHRRLIDRYQVDQIPSFIPLYSEDLIVEKNLIECIEDY 147
+ED F+E KS+ VP+ L ++ + V ++ L +L K L + +D
Sbjct: 785 DEDDFDEE-KSVQQTKVPYKNALVPKITMKTSVKELDLEAALAEREL--HKKLWKEAKDR 615
Query: 148 GADAFPFTRKRHEELKAID------KRKREEANLEEL---------LAHEGR--DFLISG 190
G D T KR+ E+ D + EA + ++ L GR D G
Sbjct: 614 GEDYKIATLKRNIEMDEYDFMHWRRSFEEREALIRDISCRKTLGLPLEEPGRYVDASFFG 435
Query: 191 DDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQ---LTDVYNNLKATKGNCFEIVLISTD 247
D+ P S L Y YW P + ++ +TD +N KG
Sbjct: 434 KDQYDPESPL-------YRYDYWGEPKNSEKSRKERMTDTHNKSIVGKG----------- 309
Query: 248 RDHKEFNVNRSSTPWLAIPYEDRTRH 273
T W +PYED +H
Sbjct: 308 ------------TVWCEMPYEDSVKH 267
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.140 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,163,275
Number of Sequences: 36976
Number of extensions: 192008
Number of successful extensions: 950
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 907
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 929
length of query: 392
length of database: 9,014,727
effective HSP length: 98
effective length of query: 294
effective length of database: 5,391,079
effective search space: 1584977226
effective search space used: 1584977226
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0372a.5


