

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0370.4
(170 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
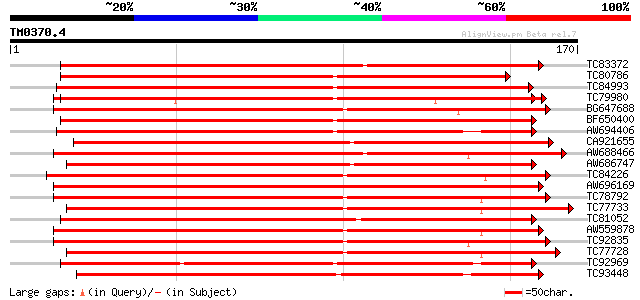
Score E
Sequences producing significant alignments: (bits) Value
TC83372 weakly similar to GP|20339360|gb|AAM19353.1 TIR-similar-... 169 5e-43
TC80786 similar to GP|20339360|gb|AAM19353.1 TIR-similar-domain-... 163 3e-41
TC84993 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanu... 163 4e-41
TC79980 weakly similar to GP|15787901|gb|AAL07542.1 resistance g... 161 1e-40
BG647688 similar to GP|9965109|gb| resistance protein MG13 {Glyc... 159 5e-40
BF650400 weakly similar to GP|9965109|gb| resistance protein MG1... 156 4e-39
AW694406 weakly similar to GP|3947735|emb NL27 {Solanum tuberosu... 155 1e-38
CA921655 similar to GP|20339360|gb TIR-similar-domain-containing... 155 1e-38
AW688466 similar to GP|9965109|gb| resistance protein MG13 {Glyc... 154 2e-38
AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein ... 154 2e-38
TC84226 similar to GP|22037313|gb|AAM89998.1 disease resistance-... 153 3e-38
AW696169 similar to GP|9965109|gb| resistance protein MG13 {Glyc... 150 2e-37
TC78792 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 150 3e-37
TC77733 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 149 4e-37
TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 149 5e-37
AW559878 weakly similar to GP|18033111|g functional candidate re... 149 7e-37
TC92835 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 148 9e-37
TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance... 146 3e-36
TC92969 weakly similar to GP|20339360|gb|AAM19353.1 TIR-similar-... 145 1e-35
TC93448 similar to GP|9858478|gb|AAG01052.1| resistance protein ... 143 4e-35
>TC83372 weakly similar to GP|20339360|gb|AAM19353.1
TIR-similar-domain-containing protein TSDC {Pisum
sativum}, partial (21%)
Length = 1151
Score = 169 bits (428), Expect = 5e-43
Identities = 79/145 (54%), Positives = 107/145 (73%)
Frame = +1
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
YD+F+SF+GEDTR++F L+ L R+G +TF+DDE LK GN IS +L KAIE S + I+
Sbjct: 475 YDIFLSFKGEDTRYSFTGFLYNILCREGFKTFMDDEELKGGNEISSSLIKAIEASRISIV 654
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
V S+N+A S WCLDELV +L+C R NQQ++ PIFY +EPS VRHQ+ YG+AM HE++
Sbjct: 655 VFSENFADSPWCLDELVTMLKCKERKNQQIL-PIFYKIEPSWVRHQRNSYGKAMTKHEEE 831
Query: 136 YGINSDRIRTWRSTLNEVCQLVGHH 160
+G +S+++ WRS L EV L G H
Sbjct: 832 FGNDSEKVNKWRSALCEVANLSGEH 906
>TC80786 similar to GP|20339360|gb|AAM19353.1 TIR-similar-domain-containing
protein TSDC {Pisum sativum}, partial (20%)
Length = 831
Score = 163 bits (413), Expect = 3e-41
Identities = 78/135 (57%), Positives = 104/135 (76%)
Frame = +1
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+ +F+SFRGEDTR++F L + L + G +TF+DDE L G+ +SP L AIE S + II
Sbjct: 325 WQMFLSFRGEDTRYSFTGSLFQALSQGGFKTFMDDEGLHTGDRVSPCLRNAIEASRLSII 504
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
VLS+NYA+STWCLDELVKILEC ++ N QLV+PIFY VEPS +RH + YG+ MA HEKK
Sbjct: 505 VLSENYANSTWCLDELVKILEC-KKWNNQLVWPIFYKVEPSDIRHLRKSYGKDMAQHEKK 681
Query: 136 YGINSDRIRTWRSTL 150
+GI+S+R++ W+S L
Sbjct: 682 FGIDSERVQKWKSAL 726
>TC84993 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum
tuberosum}, partial (21%)
Length = 672
Score = 163 bits (412), Expect = 4e-41
Identities = 77/143 (53%), Positives = 104/143 (71%)
Frame = +2
Query: 15 NYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLI 74
++DVFISFRG+DTR F L++ LK+ G++TF+DD LK G+ IS AL KAIEES I
Sbjct: 113 SFDVFISFRGDDTRRKFTSHLNEALKKSGLKTFIDDNELKKGDEISSALIKAIEESCASI 292
Query: 75 IVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEK 134
++LS+NYASS WCL+ELVKILEC ++ N Q+V PIFY ++PSHVR+Q G YG+A A +EK
Sbjct: 293 VILSENYASSKWCLNELVKILEC-KKDNGQIVIPIFYEIDPSHVRYQIGSYGQAFAKYEK 469
Query: 135 KYGINSDRIRTWRSTLNEVCQLV 157
D ++ W+ L E Q++
Sbjct: 470 NLRHKKDNLQKWKDALTEALQII 538
>TC79980 weakly similar to GP|15787901|gb|AAL07542.1 resistance gene analog
NBS7 {Helianthus annuus}, partial (33%)
Length = 1258
Score = 161 bits (407), Expect = 1e-40
Identities = 77/146 (52%), Positives = 107/146 (72%)
Frame = +3
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
YDVF+SF GEDTR++F L+ L+ +G + F+DDE L+ GN IS L KAIE+S + I+
Sbjct: 579 YDVFLSFCGEDTRYSFTGFLYHALRLEGFKIFMDDEGLEGGNQISQTLLKAIEKSRLSIV 758
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
VLS+NY STWCLDELVKI+EC ++ N +LV+P+FY +E S + ++K YG+AMAAHE +
Sbjct: 759 VLSENYGYSTWCLDELVKIMEC-KKTNNKLVWPLFYKIEQSDLSYKKSSYGKAMAAHEDR 935
Query: 136 YGINSDRIRTWRSTLNEVCQLVGHHI 161
+G S+ ++ WRS L+EV L HI
Sbjct: 936 FGKESENVQKWRSALSEVALLKADHI 1013
Score = 116 bits (290), Expect = 5e-27
Identities = 69/155 (44%), Positives = 98/155 (62%), Gaps = 10/155 (6%)
Frame = +3
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFV-------DDEMLKAGNVISPALSKA 66
+ YDVF+SFRGEDT TF L+ L+ K I+TF DDE L+ +SP++ KA
Sbjct: 54 YKYDVFLSFRGEDTYCTFAGNLYHALRNKKIKTFFPHDQIQNDDEELQ----LSPSILKA 221
Query: 67 IEESMVLIIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYG 126
I+ES + I+VLSKNYA+ST CL+ELV IL+C + N QLV+PIFY V S V+ Q+ KYG
Sbjct: 222 IQESRISIVVLSKNYATSTRCLNELVIILQCMKMKN-QLVWPIFYEVHSSDVKLQRCKYG 398
Query: 127 ---EAMAAHEKKYGINSDRIRTWRSTLNEVCQLVG 158
+A+ +++ R+ W+ L++V + G
Sbjct: 399 SSSKAILKFRERFKDYPRRMWEWQQALSQVTSIAG 503
>BG647688 similar to GP|9965109|gb| resistance protein MG13 {Glycine max},
partial (38%)
Length = 719
Score = 159 bits (402), Expect = 5e-40
Identities = 80/154 (51%), Positives = 108/154 (69%), Gaps = 5/154 (3%)
Frame = +3
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F YDVF+SFRG DTR+ F L++ L+ KGI TF+DD L+ G+ I+P+L KAI+ES ++
Sbjct: 54 FTYDVFLSFRGTDTRYGFTGNLYEALRVKGIHTFIDDRELQRGDQITPSLLKAIQESKIV 233
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
IIV S +YASS++CLDELV I+ CS+ N LV PIFY VEPSHVR+Q G YGEA+A HE
Sbjct: 234 IIVFSNHYASSSFCLDELVHIIHCSKE-NGCLVLPIFYGVEPSHVRYQTGSYGEALAEHE 410
Query: 134 -----KKYGINSDRIRTWRSTLNEVCQLVGHHIS 162
+KY N ++++ W L + L G+H +
Sbjct: 411 EARKKEKYKDNMEKLQKWEMALKQAANLSGYHFN 512
>BF650400 weakly similar to GP|9965109|gb| resistance protein MG13 {Glycine
max}, partial (36%)
Length = 644
Score = 156 bits (394), Expect = 4e-39
Identities = 74/143 (51%), Positives = 102/143 (70%)
Frame = +1
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
YDVF+SFRG+DTR F L+ L RK + TF+D+ L+ G+ ISPAL KAIEES V II
Sbjct: 151 YDVFLSFRGDDTRRNFTSHLYDTLSRKKVETFIDNNELEKGDEISPALIKAIEESHVSII 330
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
+ S+NYASS WCL+EL KIL+C ++ QQ+V P+FY+++PSHVR Q G Y +A H +
Sbjct: 331 IFSENYASSKWCLNELKKILQC-KKYMQQIVIPVFYNIDPSHVRKQTGSYEQAFTKHMRD 507
Query: 136 YGINSDRIRTWRSTLNEVCQLVG 158
+N+D+++ W++ L E LVG
Sbjct: 508 LKLNNDKLQEWKAALAEAASLVG 576
>AW694406 weakly similar to GP|3947735|emb NL27 {Solanum tuberosum}, partial
(14%)
Length = 628
Score = 155 bits (391), Expect = 1e-38
Identities = 77/144 (53%), Positives = 102/144 (70%)
Frame = +3
Query: 15 NYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLI 74
++DVFISFRG+DTR F L++ LK+ G++TF+DD LK G+ IS AL KAIEES I
Sbjct: 108 SFDVFISFRGDDTRRKFTSHLNEALKKSGVKTFIDDSELKKGDEISSALIKAIEESCASI 287
Query: 75 IVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEK 134
++ S++YASS WCL+ELVKILEC ++ N Q+V PIFY ++PSHVR+Q G YG+A A HEK
Sbjct: 288 VIFSEDYASSKWCLNELVKILEC-KKDNGQIVIPIFYEIDPSHVRNQIGSYGQAFAKHEK 464
Query: 135 KYGINSDRIRTWRSTLNEVCQLVG 158
+ + W+ L EV L G
Sbjct: 465 NL-----KQQKWKDALTEVSNLSG 521
>CA921655 similar to GP|20339360|gb TIR-similar-domain-containing protein
TSDC {Pisum sativum}, partial (32%)
Length = 763
Score = 155 bits (391), Expect = 1e-38
Identities = 77/144 (53%), Positives = 104/144 (71%)
Frame = -1
Query: 20 ISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVLSK 79
+ FRGEDTR++F L+K L + G +TF+DD L G+ ISP L AIEES + II+LS+
Sbjct: 763 LKFRGEDTRYSFTGNLYKALCQGGFKTFMDDGGLHTGDKISPTLLNAIEESRLSIIILSE 584
Query: 80 NYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGIN 139
NYA+S+WCL+ELVKI+EC + N QLV+PIFY V+PS +RH + YGE MA HE +GI+
Sbjct: 583 NYANSSWCLEELVKIMECMKLKN-QLVWPIFYKVKPSDIRHLRNCYGEDMAQHENNFGID 407
Query: 140 SDRIRTWRSTLNEVCQLVGHHIST 163
S+R++ W+S L EV L G +T
Sbjct: 406 SERVQKWKSALFEVSNLSGKAYTT 335
>AW688466 similar to GP|9965109|gb| resistance protein MG13 {Glycine max},
partial (31%)
Length = 668
Score = 154 bits (389), Expect = 2e-38
Identities = 77/157 (49%), Positives = 107/157 (68%), Gaps = 3/157 (1%)
Frame = +2
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F++DVFISFRG DTR+ F L+K L KGIRTF+DD+ L+ G+ I+P+L K+IE+S +
Sbjct: 80 FSFDVFISFRGTDTRYGFTGNLYKALSDKGIRTFIDDKELQRGDEITPSLLKSIEDSRIA 259
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
IIV SK+YASS++CL ELV I++CS ++ P+FY EPS VRHQ YGEA+A HE
Sbjct: 260 IIVFSKDYASSSFCLXELVHIIQCSNEKGTTVI-PVFYGTEPSQVRHQNDSYGEALAKHE 436
Query: 134 KKY---GINSDRIRTWRSTLNEVCQLVGHHISTTG*V 167
+ + N +R+ W+ LN+ L GHH + *+
Sbjct: 437 EGFQNSKENMERLLKWKKALNQAANLSGHHFQSRK*I 547
>AW686747 weakly similar to PIR|A54810|A5 TMV resistance protein N - tobacco
(Nicotiana glutinosa), partial (10%)
Length = 644
Score = 154 bits (388), Expect = 2e-38
Identities = 79/141 (56%), Positives = 97/141 (68%)
Frame = +3
Query: 18 VFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVL 77
VF+SFRGEDTR F L L+R+GI+TF DD L+ G VIS L+KAIEESM II+L
Sbjct: 126 VFLSFRGEDTRQGFTDHLFASLERRGIKTFKDDHDLERGEVISYELNKAIEESMFAIIIL 305
Query: 78 SKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYG 137
S NYASSTWCLDEL KI+ECS+ Q V+PIFY V+PS VRHQ+G + EA HE+K+
Sbjct: 306 SPNYASSTWCLDELKKIVECSKSFG-QAVFPIFYGVDPSDVRHQRGSFDEAFRKHEEKFR 482
Query: 138 INSDRIRTWRSTLNEVCQLVG 158
+ ++ WR L EV G
Sbjct: 483 KDRTKVERWRDALREVAGYSG 545
>TC84226 similar to GP|22037313|gb|AAM89998.1 disease resistance-like
protein GS0-1 {Glycine max}, partial (72%)
Length = 665
Score = 153 bits (387), Expect = 3e-38
Identities = 76/154 (49%), Positives = 106/154 (68%), Gaps = 3/154 (1%)
Frame = +3
Query: 12 HIFNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESM 71
++F Y VF+SFRG DTR+ F L+K L KGI TF+DD L+ G+ I+P+L AIEES
Sbjct: 18 YVFKYQVFLSFRGSDTRYGFTGNLYKALTDKGIHTFIDDSELQRGDEITPSLDNAIEESR 197
Query: 72 VLIIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAA 131
+ I V S NYASS++CLDELV I+ ++ N +LV P+F+ V+PSHVRH +G YGEA+A
Sbjct: 198 IFIPVFSANYASSSFCLDELVHIIHLYKQ-NGRLVLPVFFGVDPSHVRHHRGSYGEALAK 374
Query: 132 HEKKYGINSD---RIRTWRSTLNEVCQLVGHHIS 162
HE+++ N+D R++ W+ L + L G H S
Sbjct: 375 HEERFQHNTDHMERLQKWKIALTQAANLSGDHRS 476
>AW696169 similar to GP|9965109|gb| resistance protein MG13 {Glycine max},
partial (42%)
Length = 653
Score = 150 bits (380), Expect = 2e-37
Identities = 76/147 (51%), Positives = 100/147 (67%)
Frame = +2
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F YDVF+SFRGEDTR+ F L K L KG+RTF+DDE LK G+ I+P+L KAIE+SM+
Sbjct: 86 FKYDVFLSFRGEDTRYGFTGNLKKALDDKGVRTFIDDEKLKKGDEITPSLLKAIEDSMMA 265
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
IIVLS+NYASS++CL EL IL+ + + V P+FY V+PSHVR K YGEAM H+
Sbjct: 266 IIVLSENYASSSFCLQELSHILDTMKDKAGRYVLPVFYKVDPSHVRKLKRSYGEAMKKHD 445
Query: 134 KKYGINSDRIRTWRSTLNEVCQLVGHH 160
+ + W+ +L++V L G H
Sbjct: 446 VASSSSHNMNNKWKDSLHQVANLSGSH 526
>TC78792 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (23%)
Length = 1344
Score = 150 bits (378), Expect = 3e-37
Identities = 77/152 (50%), Positives = 102/152 (66%), Gaps = 3/152 (1%)
Frame = +3
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F YDVF+SFRG DTR+ F L+ +L +KGI TF DD L+ G+ I+ +L K IEES +
Sbjct: 21 FTYDVFLSFRGIDTRYGFTGNLYSDLCKKGIHTFFDDRELQGGDEITSSLFKVIEESRIF 200
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
I VLS NYASS++CLDELV I+ C + N++LV PIFY VEPSHVRH KG YG+A+ H
Sbjct: 201 IPVLSINYASSSFCLDELVHIIHCFKE-NRRLVLPIFYDVEPSHVRHHKGSYGKALDDHI 377
Query: 134 KKYGINS---DRIRTWRSTLNEVCQLVGHHIS 162
+ + N DR++ W+ L + GH I+
Sbjct: 378 ETFQNNKHSMDRLQKWKMALTQTANFSGHQIN 473
>TC77733 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (12%)
Length = 632
Score = 149 bits (377), Expect = 4e-37
Identities = 76/155 (49%), Positives = 103/155 (66%), Gaps = 3/155 (1%)
Frame = +3
Query: 18 VFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVL 77
VF+SFRG DTR+ F L+K L KGIRTF+DD L+ G+ I+P+L KAIEES + I +
Sbjct: 54 VFLSFRGSDTRNNFTGNLYKALVDKGIRTFIDDNDLERGDEITPSLVKAIEESRIFIPIF 233
Query: 78 SKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYG 137
S NYASS++CLDELV I+ C + LV P+FY VEP+H+RHQ G YGE + HE+++
Sbjct: 234 SANYASSSFCLDELVHIIHCYKT-KSCLVLPVFYDVEPTHIRHQSGSYGEHLTKHEERFQ 410
Query: 138 INS---DRIRTWRSTLNEVCQLVGHHISTTG*VSI 169
N +R+R W+ L + L G+H S G + I
Sbjct: 411 NNEKNMERLRQWKIALIQAANLSGYHYSPHGELGI 515
>TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 656
Score = 149 bits (376), Expect = 5e-37
Identities = 76/143 (53%), Positives = 98/143 (68%)
Frame = +2
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
YDVF++FRGEDTR F+ L L+RKGI F DD L+ G I P L +AIE S V I
Sbjct: 110 YDVFVTFRGEDTRFNFIDHLFAALQRKGIFAFRDDANLQKGESIPPELIRAIEGSQVFIA 289
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
VLSKNY+SSTWCL ELV IL+CS+ ++ V P+FY V+PS VRHQKG YGEA + HE+
Sbjct: 290 VLSKNYSSSTWCLRELVHILDCSQVSGRR-VLPVFYDVDPSEVRHQKGIYGEAFSKHEQT 466
Query: 136 YGINSDRIRTWRSTLNEVCQLVG 158
+ +S +++WR L +V + G
Sbjct: 467 FQHDSHVVQSWREALTQVGNISG 535
>AW559878 weakly similar to GP|18033111|g functional candidate resistance
protein KR1 {Glycine max}, partial (12%)
Length = 586
Score = 149 bits (375), Expect = 7e-37
Identities = 79/150 (52%), Positives = 103/150 (68%), Gaps = 3/150 (2%)
Frame = +2
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F YDVFISFRG DTR F L+K L KGIRTF+DD+ L+ G+ I+P+L K+IE S +
Sbjct: 95 FIYDVFISFRGIDTRSGFTGHLYKALCDKGIRTFIDDKELQRGDEITPSLLKSIEHSKIA 274
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
IIV S+NYA+S++CLDELV I+ + +LV P+FY VEPSHVRHQ KYGEA+ E
Sbjct: 275 IIVFSENYATSSFCLDELVHIINYFKE-KGRLVLPVFYGVEPSHVRHQNNKYGEALTEFE 451
Query: 134 KKYGINS---DRIRTWRSTLNEVCQLVGHH 160
+ + N DR++ W+ LN+V L G H
Sbjct: 452 EMFQNNKENMDRLQKWKIALNQVGNLSGXH 541
>TC92835 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (13%)
Length = 581
Score = 148 bits (374), Expect = 9e-37
Identities = 75/152 (49%), Positives = 102/152 (66%), Gaps = 3/152 (1%)
Frame = +2
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F Y VF+SFRG DTRH F L+K L KGI+TF+DD L+ G+ I+P+L KAIEES +
Sbjct: 56 FTYQVFLSFRGTDTRHGFTGNLYKALTDKGIKTFIDDNDLQRGDEITPSLLKAIEESRIF 235
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
I V S NYA+S +CLDELV I+ C + +LV P+F+ V+P++VRH G YGEA+A HE
Sbjct: 236 IPVFSINYATSKFCLDELVHIIHCYKT-KGRLVLPVFFGVDPTNVRHHTGPYGEALAGHE 412
Query: 134 KKY---GINSDRIRTWRSTLNEVCQLVGHHIS 162
K++ N +R+ W+ L + L G+H S
Sbjct: 413 KRFQNDKDNMERLHQWKLALTQAANLSGYHSS 508
>TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (12%)
Length = 765
Score = 146 bits (369), Expect = 3e-36
Identities = 74/151 (49%), Positives = 102/151 (67%), Gaps = 3/151 (1%)
Frame = +1
Query: 18 VFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVL 77
VF+SFRG DTR+ F L+K L KGIRTF+DD L+ G+ I+P+L KAIEES + I +
Sbjct: 94 VFLSFRGSDTRNKFTGNLYKALVDKGIRTFIDDNDLERGDEITPSLVKAIEESRIFIPIF 273
Query: 78 SKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYG 137
S NYASS++CLDELV I+ C + LV+P+FY VEP+H+R+Q G YGE + HE+++
Sbjct: 274 SANYASSSFCLDELVHIIHCYKT-KSCLVFPVFYDVEPTHIRNQSGIYGEHLTKHEERFQ 450
Query: 138 INS---DRIRTWRSTLNEVCQLVGHHISTTG 165
N +R+R W+ L + L G+H S G
Sbjct: 451 NNEKNMERLRQWKIALIQAANLSGYHYSPHG 543
>TC92969 weakly similar to GP|20339360|gb|AAM19353.1
TIR-similar-domain-containing protein TSDC {Pisum
sativum}, partial (15%)
Length = 772
Score = 145 bits (365), Expect = 1e-35
Identities = 71/143 (49%), Positives = 105/143 (72%)
Frame = +3
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
+DVF+SFRGEDTR+TF L+ L R I+T++D+E L+ G+ ISP+L KAI+++ + +I
Sbjct: 96 HDVFLSFRGEDTRYTFTSHLYAALTRLQIKTYIDNE-LERGDEISPSLLKAIDDAKLSVI 272
Query: 76 VLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKK 135
+ S+NYASS WCL+ELVKILEC ++ N Q++ PIFYHV P++VR+Q G YG ++A EK+
Sbjct: 273 IFSENYASSRWCLEELVKILEC-KKNNGQILVPIFYHVNPTNVRNQTGSYGLSLAEQEKR 449
Query: 136 YGINSDRIRTWRSTLNEVCQLVG 158
++ +++TWR L E G
Sbjct: 450 RDMH--KVQTWRLALTEAANFSG 512
>TC93448 similar to GP|9858478|gb|AAG01052.1| resistance protein MG23
{Glycine max}, partial (28%)
Length = 673
Score = 143 bits (360), Expect = 4e-35
Identities = 75/140 (53%), Positives = 101/140 (71%)
Frame = +1
Query: 21 SFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVLSKN 80
SFRG DTR+ F L+K L KGI TF+DDE L+ G+ I+P+L +AIEES + IIVLSKN
Sbjct: 139 SFRGLDTRYGFTGNLYKALYDKGIHTFIDDEELQRGHEITPSLLEAIEESRIAIIVLSKN 318
Query: 81 YASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINS 140
YASS++CL ELVKIL+C +G +LV+PIFY V+PS VR Q G YGEA+A +++ N
Sbjct: 319 YASSSFCLHELVKILDCI-KGKGRLVWPIFYDVDPSDVRKQTGSYGEALAMLGERF--ND 489
Query: 141 DRIRTWRSTLNEVCQLVGHH 160
+ ++ W++ L +V L G H
Sbjct: 490 NNLQIWKNALQQVANLSGWH 549
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,431,129
Number of Sequences: 36976
Number of extensions: 71934
Number of successful extensions: 662
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 558
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 561
length of query: 170
length of database: 9,014,727
effective HSP length: 89
effective length of query: 81
effective length of database: 5,723,863
effective search space: 463632903
effective search space used: 463632903
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0370.4


