

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.6
(143 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
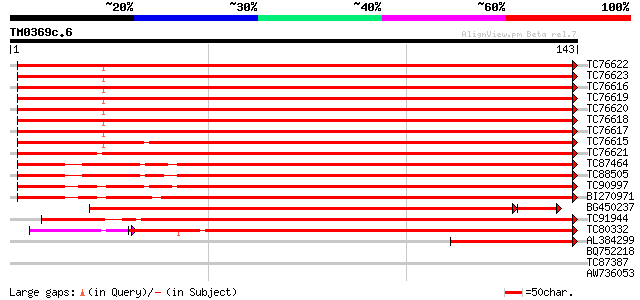
Score E
Sequences producing significant alignments: (bits) Value
TC76622 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 241 7e-65
TC76623 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 241 7e-65
TC76616 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 241 7e-65
TC76619 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 241 7e-65
TC76620 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 239 2e-64
TC76618 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 239 2e-64
TC76617 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 238 8e-64
TC76615 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 235 5e-63
TC76621 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 234 7e-63
TC87464 homologue to GP|7635498|emb|CAB88668.1 histone H2B {Cice... 226 2e-60
TC88505 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland co... 224 7e-60
TC90997 homologue to PIR|T06390|T06390 histone H2B-3 - tomato (f... 223 2e-59
BI270971 homologue to PIR|T06390|T06 histone H2B-3 - tomato (fra... 222 5e-59
BG450237 homologue to SP|O22582|H2B_ Histone H2B. [Upland cotton... 179 4e-46
TC91944 similar to PIR|S11937|S11937 histone H2B - Emericella ni... 168 8e-43
TC80332 similar to PIR|S11937|S11937 histone H2B - Emericella ni... 157 3e-40
AL384299 PIR|T30054|HSK histone H2B [validated] - Caenorhabditis... 62 8e-11
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 36 0.006
TC87387 homologue to PIR|T47775|T47775 hypothetical protein F24I... 34 0.017
AW736053 homologue to GP|10177235|dbj histone H2B like protein {... 33 0.030
>TC76622 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 904
Score = 241 bits (615), Expect = 7e-65
Identities = 126/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAGDKKKKRSKKNVETYKI
Sbjct: 83 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGDKKKKRSKKNVETYKI 262
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 263 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 442
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 443 LVLPGELAKHAVSEGTKAVTKFTSS 517
>TC76623 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 647
Score = 241 bits (615), Expect = 7e-65
Identities = 126/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAGDKKKKRSKKNVETYKI
Sbjct: 81 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGDKKKKRSKKNVETYKI 260
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 261 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 440
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 441 LVLPGELAKHAVSEGTKAVTKFTSS 515
>TC76616 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 698
Score = 241 bits (615), Expect = 7e-65
Identities = 126/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAGDKKKKRSKKNVETYKI
Sbjct: 93 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGDKKKKRSKKNVETYKI 272
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 273 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 452
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 453 LVLPGELAKHAVSEGTKAVTKFTSS 527
>TC76619 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 718
Score = 241 bits (615), Expect = 7e-65
Identities = 126/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAGDKKKKRSKKNVETYKI
Sbjct: 84 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGDKKKKRSKKNVETYKI 263
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 264 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 443
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 444 LVLPGELAKHAVSEGTKAVTKFTSS 518
>TC76620 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 718
Score = 239 bits (611), Expect = 2e-64
Identities = 125/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAG+KKKKRSKKNVETYKI
Sbjct: 44 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGEKKKKRSKKNVETYKI 223
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 224 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 403
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 404 LVLPGELAKHAVSEGTKAVTKFTSS 478
>TC76618 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 755
Score = 239 bits (611), Expect = 2e-64
Identities = 125/145 (86%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAG+KKKKRSKKNVETYKI
Sbjct: 71 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGEKKKKRSKKNVETYKI 250
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 251 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 430
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 431 LVLPGELAKHAVSEGTKAVTKFTSS 505
>TC76617 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 760
Score = 238 bits (606), Expect = 8e-64
Identities = 124/145 (85%), Positives = 132/145 (90%), Gaps = 4/145 (2%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P + G AAG+KKKKRSKK+VETYKI
Sbjct: 102 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLPKEGGSAAGEKKKKRSKKSVETYKI 281
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 282 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 461
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 462 LVLPGELAKHAVSEGTKAVTKFTSS 536
>TC76615 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, complete
Length = 685
Score = 235 bits (599), Expect = 5e-63
Identities = 125/145 (86%), Positives = 131/145 (90%), Gaps = 4/145 (2%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAA----AEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKI 58
KG+KKPAEKKPA ++ A AEKKP+A KK+P KEGG+A KKKRSKKNVETYKI
Sbjct: 84 KGEKKPAEKKPAEEKKSTVAEKAPAEKKPKAGKKLP-KEGGSAAGDKKKRSKKNVETYKI 260
Query: 59 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 118
YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR
Sbjct: 261 YIFKVLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVR 440
Query: 119 LVLPGELAKHAVSEGTKAVTKFTSS 143
LVLPGELAKHAVSEGTKAVTKFTSS
Sbjct: 441 LVLPGELAKHAVSEGTKAVTKFTSS 515
>TC76621 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, partial (89%)
Length = 589
Score = 234 bits (598), Expect = 7e-63
Identities = 121/141 (85%), Positives = 131/141 (92%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK 62
KG+KKPAE+K + EKA A EKKP+A KK+P + G A+G+KKKKR+KK+VETYKIYIFK
Sbjct: 65 KGEKKPAEEKKSTVAEKAPA-EKKPKAGKKLPKESGSASGEKKKKRNKKSVETYKIYIFK 241
Query: 63 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 122
VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP
Sbjct: 242 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 421
Query: 123 GELAKHAVSEGTKAVTKFTSS 143
GELAKHAVSEGTKAVTKFTSS
Sbjct: 422 GELAKHAVSEGTKAVTKFTSS 484
>TC87464 homologue to GP|7635498|emb|CAB88668.1 histone H2B {Cicer
arietinum}, complete
Length = 677
Score = 226 bits (576), Expect = 2e-60
Identities = 122/141 (86%), Positives = 130/141 (91%)
Frame = +3
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK 62
K +KKPAEKKPA + A AEKKP+AEKKI +KEG + DKKKKR+KK+VETYKIYIFK
Sbjct: 69 KAEKKPAEKKPA----EKAPAEKKPKAEKKI-SKEGSS--DKKKKRTKKSVETYKIYIFK 227
Query: 63 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 122
VLKQVHPDIG+SSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP
Sbjct: 228 VLKQVHPDIGVSSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 407
Query: 123 GELAKHAVSEGTKAVTKFTSS 143
GELAKHAVSEGTKAVTKFTSS
Sbjct: 408 GELAKHAVSEGTKAVTKFTSS 470
>TC88505 homologue to SP|O22582|H2B_GOSHI Histone H2B. [Upland cotton]
{Gossypium hirsutum}, partial (92%)
Length = 690
Score = 224 bits (572), Expect = 7e-60
Identities = 122/141 (86%), Positives = 129/141 (90%)
Frame = +1
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK 62
K +KKPAEKKPA + + AEKKP+AEKKI +KEGG DKKKKR KK+VETYKIYIFK
Sbjct: 85 KAEKKPAEKKPA----EKSPAEKKPKAEKKI-SKEGG---DKKKKRVKKSVETYKIYIFK 240
Query: 63 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 122
VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQE+SRLARYNKKPTITSREIQTAVRLVLP
Sbjct: 241 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQEASRLARYNKKPTITSREIQTAVRLVLP 420
Query: 123 GELAKHAVSEGTKAVTKFTSS 143
GELAKHAVSEGTKAVTKFTSS
Sbjct: 421 GELAKHAVSEGTKAVTKFTSS 483
>TC90997 homologue to PIR|T06390|T06390 histone H2B-3 - tomato (fragment),
partial (94%)
Length = 665
Score = 223 bits (569), Expect = 2e-59
Identities = 124/141 (87%), Positives = 130/141 (91%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK 62
KG+KKPAEKKPA EKA A +K +AEKKIP KE +A DKKKKR+KK+VETYKIYIFK
Sbjct: 59 KGEKKPAEKKPA---EKAPA--EKTKAEKKIP-KEASSA-DKKKKRNKKSVETYKIYIFK 217
Query: 63 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 122
VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP
Sbjct: 218 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 397
Query: 123 GELAKHAVSEGTKAVTKFTSS 143
GELAKHAVSEGTKAVTKFTSS
Sbjct: 398 GELAKHAVSEGTKAVTKFTSS 460
>BI270971 homologue to PIR|T06390|T06 histone H2B-3 - tomato (fragment),
partial (94%)
Length = 441
Score = 222 bits (565), Expect = 5e-59
Identities = 121/141 (85%), Positives = 129/141 (90%)
Frame = +2
Query: 3 KGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFK 62
KG+KKPAEKKPA EKA A +K +AEKKIP + ++ DKKKKR+KK+VETYKIYIFK
Sbjct: 26 KGEKKPAEKKPA---EKAPA--EKTKAEKKIPKE--ASSTDKKKKRNKKSVETYKIYIFK 184
Query: 63 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 122
VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP
Sbjct: 185 VLKQVHPDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLP 364
Query: 123 GELAKHAVSEGTKAVTKFTSS 143
GELAKHAVSEGTKAVTKFTSS
Sbjct: 365 GELAKHAVSEGTKAVTKFTSS 427
>BG450237 homologue to SP|O22582|H2B_ Histone H2B. [Upland cotton] {Gossypium
hirsutum}, partial (80%)
Length = 682
Score = 179 bits (454), Expect(2) = 4e-46
Identities = 93/108 (86%), Positives = 96/108 (88%)
Frame = -2
Query: 21 AAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVLKQVHPDIGISSKAMGI 80
A AEKKP+A KK+P + G AAGD K KRSKKNVETYKIYIFKVLKQVHPDIG SSKAMGI
Sbjct: 486 APAEKKPKAGKKLPKEGGSAAGDXKXKRSKKNVETYKIYIFKVLKQVHPDIGXSSKAMGI 307
Query: 81 MNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGELAKH 128
MNSFINDIF KLAQESSRLARYNKK TITSREIQT VRLVL GELAKH
Sbjct: 306 MNSFINDIFXKLAQESSRLARYNKKXTITSREIQTXVRLVLXGELAKH 163
Score = 21.2 bits (43), Expect(2) = 4e-46
Identities = 9/11 (81%), Positives = 11/11 (99%)
Frame = -3
Query: 129 AVSEGTKAVTK 139
AVSEGTKAV++
Sbjct: 161 AVSEGTKAVSQ 129
>TC91944 similar to PIR|S11937|S11937 histone H2B - Emericella nidulans,
partial (79%)
Length = 772
Score = 168 bits (425), Expect = 8e-43
Identities = 90/135 (66%), Positives = 108/135 (79%)
Frame = +3
Query: 9 AEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVETYKIYIFKVLKQVH 68
AEKKPA + A A AEKK +K+ A+GDK KK+ K ETY YI+KVLKQVH
Sbjct: 105 AEKKPATGGKAPAKAP----AEKK-DSKKPVASGDKTKKKRKVRKETYSSYIYKVLKQVH 269
Query: 69 PDIGISSKAMGIMNSFINDIFEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGELAKH 128
PD GIS+KAM I+NSF+NDIFE+++ E+S+LA YNK+ TI+SREIQTAVRL+LPGELAKH
Sbjct: 270 PDTGISNKAMSILNSFVNDIFERISGEASKLAAYNKRSTISSREIQTAVRLILPGELAKH 449
Query: 129 AVSEGTKAVTKFTSS 143
AVSEGTKAVTK+TSS
Sbjct: 450 AVSEGTKAVTKYTSS 494
>TC80332 similar to PIR|S11937|S11937 histone H2B - Emericella nidulans,
partial (92%)
Length = 562
Score = 157 bits (398), Expect(2) = 3e-40
Identities = 82/115 (71%), Positives = 97/115 (84%), Gaps = 2/115 (1%)
Frame = +3
Query: 31 KKIPAKEGGAA--GDKKKKRSKKNVETYKIYIFKVLKQVHPDIGISSKAMGIMNSFINDI 88
K +P K+ A+ GDK KK+ K ETY YI+KVLKQVHPD GIS+KAM IMNSF+NDI
Sbjct: 96 KHLPKKKKAASAPGDKVKKK-KARKETYSSYIYKVLKQVHPDTGISNKAMSIMNSFVNDI 272
Query: 89 FEKLAQESSRLARYNKKPTITSREIQTAVRLVLPGELAKHAVSEGTKAVTKFTSS 143
FE++A E+S+LA YNK+ TI+SREIQTAVRL+LPGELAKHAVSEGTKAVTK+TSS
Sbjct: 273 FERIAAEASKLAAYNKRSTISSREIQTAVRLILPGELAKHAVSEGTKAVTKYTSS 437
Score = 23.1 bits (48), Expect(2) = 3e-40
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +2
Query: 6 KKPAEKKPAAAEEKAAAAEKKPRAEKK 32
K AEKKPA KA AA K P +K+
Sbjct: 41 KAVAEKKPATTGGKAPAA-KAPSKKKE 118
>AL384299 PIR|T30054|HSK histone H2B [validated] - Caenorhabditis elegans,
partial (26%)
Length = 411
Score = 62.0 bits (149), Expect = 8e-11
Identities = 30/32 (93%), Positives = 32/32 (99%)
Frame = +1
Query: 112 EIQTAVRLVLPGELAKHAVSEGTKAVTKFTSS 143
EIQTAVRL+LPGELAKHAVSEGTKAVTK+TSS
Sbjct: 1 EIQTAVRLILPGELAKHAVSEGTKAVTKYTSS 96
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 35.8 bits (81), Expect = 0.006
Identities = 21/49 (42%), Positives = 24/49 (48%)
Frame = -2
Query: 6 KKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVE 54
KK AEKK A E+AA KK R AKE +KK K K+ E
Sbjct: 600 KKEAEKKKKKAREEAAKKAKKAREAAYKKAKEEKKEAEKKAKEEKRQAE 454
Score = 33.9 bits (76), Expect = 0.023
Identities = 23/54 (42%), Positives = 33/54 (60%)
Frame = -2
Query: 2 AKGDKKPAEKKPAAAEEKAAAAEKKPRAEKKIPAKEGGAAGDKKKKRSKKNVET 55
AK +KK AEKK A+E+ AEKK + EKK K+ A KK++ +K+ E+
Sbjct: 513 AKEEKKEAEKK---AKEEKRQAEKKIKEEKKEAEKK---AKKDKKEQDRKDAES 370
>TC87387 homologue to PIR|T47775|T47775 hypothetical protein F24I3.230 -
Arabidopsis thaliana, partial (14%)
Length = 1074
Score = 34.3 bits (77), Expect = 0.017
Identities = 25/59 (42%), Positives = 30/59 (50%), Gaps = 9/59 (15%)
Frame = +3
Query: 5 DKKPAEKKPAAAEEKAAAA--EKKPRAEKKIPAKEGGAAG-------DKKKKRSKKNVE 54
DKK +KK +E AAA E+K EKK KE G G DKKKK+ K+ E
Sbjct: 381 DKKEKKKKKKKDKENGAAASDEEKVEKEKKKKHKEKGEDGSPEVEKSDKKKKKHKETSE 557
>AW736053 homologue to GP|10177235|dbj histone H2B like protein {Arabidopsis
thaliana}, partial (11%)
Length = 152
Score = 33.5 bits (75), Expect = 0.030
Identities = 16/17 (94%), Positives = 16/17 (94%)
Frame = -3
Query: 108 ITSREIQTAVRLVLPGE 124
ITSR IQTAVRLVLPGE
Sbjct: 51 ITSRNIQTAVRLVLPGE 1
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.308 0.125 0.326
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,795,099
Number of Sequences: 36976
Number of extensions: 27895
Number of successful extensions: 380
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 268
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 310
length of query: 143
length of database: 9,014,727
effective HSP length: 87
effective length of query: 56
effective length of database: 5,797,815
effective search space: 324677640
effective search space used: 324677640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0369c.6


