

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
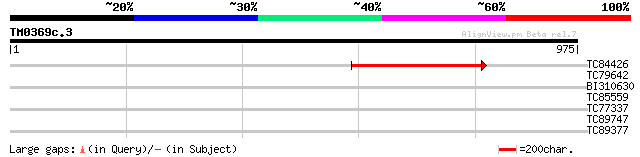
Query= TM0369c.3
(975 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC84426 361 e-100
TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like pr... 33 0.41
BI310630 similar to GP|18087548|gb AT5g55600/MDF20_4 {Arabidopsi... 33 0.41
TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC ... 33 0.70
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 31 2.0
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 29 7.7
TC89377 similar to GP|10177940|dbj|BAB11299. gene_id:MQD19.5~unk... 29 7.7
>TC84426
Length = 707
Score = 361 bits (927), Expect = e-100
Identities = 178/233 (76%), Positives = 197/233 (84%)
Frame = +1
Query: 588 GFCATEKNLLKWTVYLEKQKRPNGSTETIVKQSSVIREPKVEHIADTKKTGVRKPHPAVQ 647
GFCATEK LLKWT+YLEKQKR NGSTE+I KQSSVI P VE+IAD K+TG + HPA+Q
Sbjct: 1 GFCATEKYLLKWTIYLEKQKRLNGSTESIGKQSSVIGGPNVEYIADRKRTGDGQSHPALQ 180
Query: 648 NGGGDQRTSLGISKDDDKKISFGRGFPIFKMRKPFSEVVAGSAAVDSGFPPLQQRKLPAS 707
NG D RTSL I++ DDK ISFGRGFPIFKMRKPFSEVVAGSAAVDSGFPPLQQRKLP
Sbjct: 181 NGDEDLRTSLDINRTDDKNISFGRGFPIFKMRKPFSEVVAGSAAVDSGFPPLQQRKLPTL 360
Query: 708 GSEKGMKLSKPSNQNGERVNAAVDYEISQKSQDILLTEGSLDGSGNNSCRDDDPFLRIGS 767
GSEKG+K S+PSNQN ERVNA ++++ISQKSQD+ TEG L G+GNN RD DPFLRIGS
Sbjct: 361 GSEKGVKQSRPSNQNAERVNATINHQISQKSQDMSFTEGPLHGNGNNGSRDGDPFLRIGS 540
Query: 768 NVVPVYLSGGERSKPHSSLKHVIVYIGFEHECPRGHRFLLNARHLNELGSSYS 820
NV+PVYL G R+KPHSSLKH VY+GFEHECPRGHRFLLNA HL ELGS YS
Sbjct: 541 NVLPVYLDDGTRNKPHSSLKHETVYVGFEHECPRGHRFLLNADHLTELGSLYS 699
>TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like protein
(clone PKCBP) - potato, partial (42%)
Length = 2321
Score = 33.5 bits (75), Expect = 0.41
Identities = 26/97 (26%), Positives = 45/97 (45%)
Frame = -3
Query: 490 DVETSNSNIGASPPSHKPHSSGYFFLHACACGRSRQLRPDPFDFESADASCFSDCDKLLP 549
D+ NS + S + ++S + L +C+ RSR+L F F S D+ C LL
Sbjct: 738 DLAAINSCLYPSIFLSRSYTSDFTALRSCSSFRSRELLSSEFLFSSTDSFCSKIAFSLLK 559
Query: 550 SVKLPEIEVAGPVQSSAWSLLRIGSAKYYESSKGLLQ 586
+ + E + P+Q + S L S + S++ L+
Sbjct: 558 VLIVSSAESSSPLQYFS-SYLSFCSTWLFVSTRSFLE 451
>BI310630 similar to GP|18087548|gb AT5g55600/MDF20_4 {Arabidopsis thaliana},
partial (19%)
Length = 730
Score = 33.5 bits (75), Expect = 0.41
Identities = 23/57 (40%), Positives = 32/57 (55%), Gaps = 5/57 (8%)
Frame = -3
Query: 54 PLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGL-----GSYASLFP 105
P PS H+S ++ PTS+ N+ S G G V R G +S +SGL G ++S FP
Sbjct: 722 PHPSLHHAS-TAVPTSTINSCSSRGGAGLMVDRPGCLISSLSGLANLKAGIHSSRFP 555
>TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC 6.4.1.2) -
potato chloroplast, partial (66%)
Length = 5011
Score = 32.7 bits (73), Expect = 0.70
Identities = 27/125 (21%), Positives = 53/125 (41%)
Frame = -1
Query: 37 QAAKHAMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSG 96
+ + + PF++ + P SS SS+P S ++S S S+ S +
Sbjct: 3700 EGSSSSSKPFSEGSSSSSKPFSEGSSSSSKPFSEGSSSK-------LFSEGSSSSSKLFS 3542
Query: 97 LGSYASLFPGQCIPVTLFVFVDDFSNLSNSSTNGEESSDVSSLSQSNSMSSAAKANLPAK 156
GS + LF + S+ S+ + SS+++S+ + +MSS+ +P
Sbjct: 3541 EGSSSKLF-------------SEGSSSSSKPFSEGSSSELNSMGSTGNMSSSIGFQVPGS 3401
Query: 157 VSGSV 161
+ S+
Sbjct: 3400 IKSSI 3386
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 31.2 bits (69), Expect = 2.0
Identities = 31/109 (28%), Positives = 49/109 (44%), Gaps = 13/109 (11%)
Frame = -2
Query: 57 SRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYAS-------------L 103
S S SSLSS T S+++SSP + S + S+ S S S +S
Sbjct: 733 SSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSAPPAPPSGA 554
Query: 104 FPGQCIPVTLFVFVDDFSNLSNSSTNGEESSDVSSLSQSNSMSSAAKAN 152
F + + L + S+ ++SS + S SS S S+S+ S+ A+
Sbjct: 553 FFELLLLLELLPYPPPSSDDNSSSPSPNAPSPSSSSSSSSSLPSSPSAS 407
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 29.3 bits (64), Expect = 7.7
Identities = 20/76 (26%), Positives = 29/76 (37%), Gaps = 5/76 (6%)
Frame = -2
Query: 428 PTSQHEDHLDKALHAFHSMVKGPSVQ-----VFARKLEEECTSIWKSGRQLCDAISLTGK 482
P H +K L +GPS +F + + C + QL A TGK
Sbjct: 765 PHRYHAQKREKPLSHREQQREGPSPSQFQQIIFLQACQTSCLLLHSLQHQLSLAKHTTGK 586
Query: 483 PCMHQRHDVETSNSNI 498
PC+ H + + NI
Sbjct: 585 PCVAPPHQQSSHSVNI 538
>TC89377 similar to GP|10177940|dbj|BAB11299. gene_id:MQD19.5~unknown
protein {Arabidopsis thaliana}, partial (66%)
Length = 1480
Score = 29.3 bits (64), Expect = 7.7
Identities = 17/50 (34%), Positives = 27/50 (54%)
Frame = -3
Query: 120 FSNLSNSSTNGEESSDVSSLSQSNSMSSAAKANLPAKVSGSVVMLARPAS 169
FS+L T E+ + LS ++S +SA+ LPA+ S + RP+S
Sbjct: 788 FSSLDMPDTAFPEACSLVLLSVNSSSASASSLELPARKKSSSLRSRRPSS 639
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,695,769
Number of Sequences: 36976
Number of extensions: 435242
Number of successful extensions: 2122
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2046
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2100
length of query: 975
length of database: 9,014,727
effective HSP length: 105
effective length of query: 870
effective length of database: 5,132,247
effective search space: 4465054890
effective search space used: 4465054890
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0369c.3


