

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
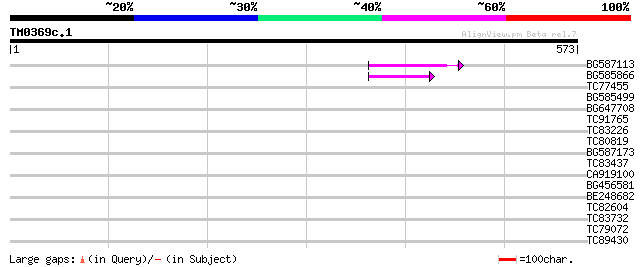
Query= TM0369c.1
(573 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 51 1e-06
BG585866 50 3e-06
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 36 0.035
BG585499 35 0.060
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 35 0.060
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 33 0.23
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 33 0.23
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 33 0.30
BG587173 weakly similar to GP|4512630|gb| Contains reverse trans... 32 0.51
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 31 1.5
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 30 1.9
BG456581 30 3.3
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 29 4.3
TC82604 similar to GP|11994348|dbj|BAB02307. gene_id:MSJ11.16~un... 29 5.6
TC83732 similar to GP|11762164|gb|AAG40360.1 AT4g22310 {Arabidop... 28 9.6
TC79072 28 9.6
TC89430 similar to SP|P46573|APKB_ARATH Protein kinase APK1B (EC... 28 9.6
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 50.8 bits (120), Expect = 1e-06
Identities = 30/98 (30%), Positives = 44/98 (44%), Gaps = 2/98 (2%)
Frame = -3
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS- 421
+WK + KV FLW ++LPT H+ K +C RCG E V H L +CP +
Sbjct: 288 VWKYNTSPKVRHFLWRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYAR 109
Query: 422 -VWVRSRFAKALPGLSAGALMTRSSMVL**DNLHEILA 458
+W S G+ +L + NLH +L+
Sbjct: 108 LIWATSPIHAPPYGIMYDSLYS---------NLHRVLS 22
>BG585866
Length = 828
Score = 49.7 bits (117), Expect = 3e-06
Identities = 26/69 (37%), Positives = 35/69 (50%), Gaps = 2/69 (2%)
Frame = +3
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS- 421
IW+ EK FFLWLA +A+PT H ++V C RCG E H +R C S
Sbjct: 462 IWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCRFSK 641
Query: 422 -VWVRSRFA 429
+W + F+
Sbjct: 642 IIWHKIGFS 668
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 36.2 bits (82), Expect = 0.035
Identities = 19/70 (27%), Positives = 31/70 (44%), Gaps = 3/70 (4%)
Frame = -2
Query: 352 LRHPKFAVMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFD---ACFRCGASV 408
L + + + + +WK+ A KV+ F W D +PT + L++ + C CG
Sbjct: 355 LSYEEEVIFRELWKSKAPAKVLAFSWTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQD 176
Query: 409 EDVEHALRRC 418
E V H C
Sbjct: 175 ETVVHLFLHC 146
>BG585499
Length = 792
Score = 35.4 bits (80), Expect = 0.060
Identities = 21/67 (31%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Frame = +3
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC-P 419
K++W + F+WL + TN +R C CG + E V H L C P
Sbjct: 228 KMLWGWRGPHRTQTFMWLVAHGCILTNYRRSRWGTRVLATCPCCGNADETVLHVLCDCRP 407
Query: 420 GS-VWVR 425
S VW+R
Sbjct: 408 ASQVWIR 428
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 35.4 bits (80), Expect = 0.060
Identities = 24/69 (34%), Positives = 36/69 (51%), Gaps = 4/69 (5%)
Frame = +1
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRM-FCSNVVDQRKQ-- 66
I+HL D LLF + + I Q L + ASG VN +KS + + NV +Q K+
Sbjct: 145 ITHLLFADDSLLFARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMI 324
Query: 67 -QELSIIVG 74
Q+++I G
Sbjct: 325 CQQIAIKTG 351
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 33.5 bits (75), Expect = 0.23
Identities = 21/72 (29%), Positives = 36/72 (49%)
Frame = +3
Query: 31 QLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQELSIIVGITRASYLGKYLDLPQL 90
Q++ L + E SG ++L+KS ++CS V + ++ I+G+ KYL LP +
Sbjct: 333 QIMKNILILYEEDSGKAISLRKS*IYCSRNVPDILKTSITYILGVQFMLGTCKYLGLPSM 512
Query: 91 KARVTKEHFAPI 102
R F+ I
Sbjct: 513 IGRDRTTTFSSI 548
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 33.5 bits (75), Expect = 0.23
Identities = 19/70 (27%), Positives = 26/70 (37%), Gaps = 2/70 (2%)
Frame = +3
Query: 359 VMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC 418
+ K IW + LW D+LP + + C RC + E + H C
Sbjct: 117 IWKKIWSLHTIPRHKVLLWRILNDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSC 296
Query: 419 PGS--VWVRS 426
P S VW S
Sbjct: 297 PLSKRVWFGS 326
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 33.1 bits (74), Expect = 0.30
Identities = 25/84 (29%), Positives = 34/84 (39%), Gaps = 4/84 (4%)
Frame = +2
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYH-SHLVKFDACFRCG-ASVEDVEHALRRCP- 419
+W KV F+W R+ LPT H L+ +A CG E H C
Sbjct: 626 VWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNV 805
Query: 420 -GSVWVRSRFAKALPGLSAGALMT 442
S+W R +P +S+G L T
Sbjct: 806 FCSLWSLVRNWLGIPSMSSGELRT 877
>BG587173 weakly similar to GP|4512630|gb| Contains reverse transcriptase
domain (rvt) PF|00078. {Arabidopsis thaliana}, partial
(5%)
Length = 292
Score = 32.3 bits (72), Expect = 0.51
Identities = 18/59 (30%), Positives = 28/59 (46%), Gaps = 2/59 (3%)
Frame = +1
Query: 362 LIWKATAAEKVMFFLWLAGRDALPTNMK--RYHSHLVKFDACFRCGASVEDVEHALRRC 418
++W + K F +W+A D LPT + + + +VK C CG E +H RC
Sbjct: 97 VVWFSGHIPKHAFNMWVANYDRLPTRCRFAAWDTSIVK--TCLLCGHYDESRDHLFLRC 267
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 30.8 bits (68), Expect = 1.5
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 3/76 (3%)
Frame = +2
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
+SHL + LL + ++ + L F SG+KVN KS + C N+ +
Sbjct: 542 VSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVNFHKSGLVCVNIAPSWLSEAA 721
Query: 70 SII---VGITRASYLG 82
S++ VG YLG
Sbjct: 722 SVLSWKVGKVPFLYLG 769
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 30.4 bits (67), Expect = 1.9
Identities = 18/67 (26%), Positives = 28/67 (40%), Gaps = 4/67 (5%)
Frame = -2
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDA--CFRCGASVEDVEHALRRC 418
++IW KV F W RD LPT + ++ +A C ++E +H C
Sbjct: 590 EMIWHRQVPLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLSC 411
Query: 419 P--GSVW 423
S+W
Sbjct: 410 SYFASLW 390
>BG456581
Length = 683
Score = 29.6 bits (65), Expect = 3.3
Identities = 22/84 (26%), Positives = 29/84 (34%), Gaps = 4/84 (4%)
Frame = +2
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGSV 422
IW + F W D LPT+ C C VE +H RC V
Sbjct: 80 IWNSCIPPSHSFICWRLAHDRLPTDDNLSSRGCALVSMCSFCLEQVETSDHLFLRCKFVV 259
Query: 423 ----WVRSRFAKALPGLSAGALMT 442
W+ S+ L S AL++
Sbjct: 260 TLWSWLCSQLRVGLDFSSFKALLS 331
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 29.3 bits (64), Expect = 4.3
Identities = 19/58 (32%), Positives = 27/58 (45%)
Frame = +3
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
+ + G +SHL D L + L + L+ F ASG+KVN KS + NV
Sbjct: 138 VKRGGTRVSHLQYADDTLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINV 311
>TC82604 similar to GP|11994348|dbj|BAB02307. gene_id:MSJ11.16~unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 713
Score = 28.9 bits (63), Expect = 5.6
Identities = 15/47 (31%), Positives = 24/47 (50%), Gaps = 3/47 (6%)
Frame = +2
Query: 471 MLERHGCWKFLH---ENSRKSSCISRNCHVVLFFSNFNAILCVF*RK 514
+ RHGC + H S + S +S C FF++++ L +F* K
Sbjct: 122 LCNRHGCLRHSHHPCSPSSEISILSSPCFFFFFFNSYSTFLFIF**K 262
>TC83732 similar to GP|11762164|gb|AAG40360.1 AT4g22310 {Arabidopsis
thaliana}, partial (35%)
Length = 716
Score = 28.1 bits (61), Expect = 9.6
Identities = 16/65 (24%), Positives = 30/65 (45%)
Frame = +1
Query: 456 ILAFNPCKFSLFLAQMLERHGCWKFLHENSRKSSCISRNCHVVLFFSNFNAILCVF*RKC 515
IL +PC+ +F+ + C N+ SSC +++L++ F ++ C F
Sbjct: 514 ILFIHPCEKCVFVDSLKGTDMCK*MNLRNAEYSSCKQLLTYILLYWGLFRSMRC*FLLFS 693
Query: 516 AKLRW 520
+L W
Sbjct: 694 TQLSW 708
>TC79072
Length = 2401
Score = 28.1 bits (61), Expect = 9.6
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 20/84 (23%)
Frame = +2
Query: 483 ENSRKS----SCISRNCHVVLFFSNFNAILCV-----------F*RKCAKLRWGRPTQEP 527
+ SRKS S ++ NC V+ FN +LC+ F* C L + R P
Sbjct: 2150 DGSRKSPSTTS*LTTNCAQVVLVLAFNNLLCLYK*YISLHSTYF*NICVCLEFQR*NTTP 2329
Query: 528 TLIAASYE-----EN*VSLHIDGS 546
L+ Y+ EN*++L+I S
Sbjct: 2330 ILLDEKYKSICILEN*INLNIYSS 2401
>TC89430 similar to SP|P46573|APKB_ARATH Protein kinase APK1B (EC 2.7.1.-).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (71%)
Length = 1663
Score = 28.1 bits (61), Expect = 9.6
Identities = 19/53 (35%), Positives = 23/53 (42%)
Frame = +2
Query: 459 FNPCKFSLFLAQMLERHGCWKFLHENSRKSSCISRNCHVVLFFSNFNAILCVF 511
F P +SL L L C FLH KS + +L SN+NA L F
Sbjct: 743 FQPLSWSLRLKVALGAAKCLAFLHSAETKSIYQNLKTSNILLDSNYNAKLFNF 901
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.339 0.147 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,124,081
Number of Sequences: 36976
Number of extensions: 338189
Number of successful extensions: 2495
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1831
Number of HSP's successfully gapped in prelim test: 87
Number of HSP's that attempted gapping in prelim test: 638
Number of HSP's gapped (non-prelim): 1965
length of query: 573
length of database: 9,014,727
effective HSP length: 101
effective length of query: 472
effective length of database: 5,280,151
effective search space: 2492231272
effective search space used: 2492231272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0369c.1


