

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369b.1
(266 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
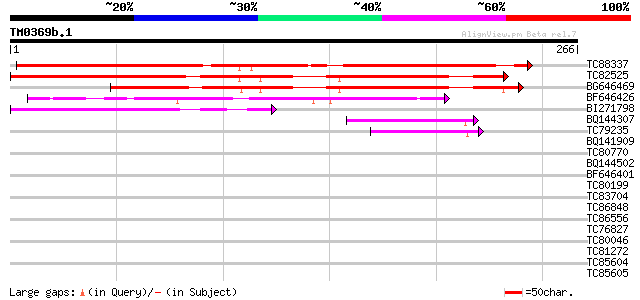
TC88337 similar to GP|10177240|dbj|BAB10614. emb|CAB89309.1~gene... 300 4e-82
TC82525 similar to GP|10177240|dbj|BAB10614. emb|CAB89309.1~gene... 289 9e-79
BG646469 similar to GP|10177240|db emb|CAB89309.1~gene_id:MRN17.... 229 8e-61
BF646426 similar to GP|10177240|dbj emb|CAB89309.1~gene_id:MRN17... 91 4e-19
BI271798 weakly similar to GP|10177240|dbj emb|CAB89309.1~gene_i... 74 6e-14
BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 43 1e-04
TC79235 weakly similar to GP|7297741|gb|AAF52992.1| CG17108 gene... 41 5e-04
BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox ... 40 0.001
TC80770 similar to GP|4680336|gb|AAD27627.1| hypothetical protei... 39 0.002
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 39 0.002
BF646401 similar to PIR|A02951|KRMS keratin type II cytoskeleta... 38 0.004
TC80199 weakly similar to GP|21593112|gb|AAM65061.1 unknown {Ara... 36 0.018
TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19... 36 0.018
TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {A... 36 0.018
TC86556 similar to PIR|T07598|T07598 proline-rich protein GPP1 -... 36 0.018
TC76827 similar to GP|17154773|gb|AAL35979.1 extensin-like prote... 35 0.023
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 35 0.023
TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 ... 35 0.023
TC85604 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago t... 35 0.023
TC85605 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago t... 35 0.023
>TC88337 similar to GP|10177240|dbj|BAB10614.
emb|CAB89309.1~gene_id:MRN17.16~similar to unknown
protein {Arabidopsis thaliana}, partial (50%)
Length = 787
Score = 300 bits (768), Expect = 4e-82
Identities = 163/245 (66%), Positives = 189/245 (76%), Gaps = 3/245 (1%)
Frame = +2
Query: 4 QRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTI 63
QR+ LL+ AY++QGKSM+E++Q+L+YTT+ELEQTR VQEELRKRD+QL NLK+L N I
Sbjct: 110 QRSPLLSWAYYYQGKSMDELKQTLMYTTMELEQTRVTVQEELRKRDEQLHNLKELFNKVI 289
Query: 64 RERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDE--PRRGID-SNNGLSSS 120
RERDEAQEKCQRL LEK LFQ Q+QQ Q DP SG+SSIEDE +RGI+ SNNG+S S
Sbjct: 290 RERDEAQEKCQRLFLEKLLFQQQKQQNQ---DPLSGISSIEDEQIQKRGIESSNNGISLS 460
Query: 121 DCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTL 180
DCEESIVSSP QP + + S+A+ +KPLPEKGKLLQAVMKAGPLLQTL
Sbjct: 461 DCEESIVSSPQQTRTEQPMM-MMIDSIAK-------DKPLPEKGKLLQAVMKAGPLLQTL 616
Query: 181 LLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCG 240
LLAGPLPQWRHPPPPLESFEIPPVTI SPP QPPQ QLL QESF N S CG
Sbjct: 617 LLAGPLPQWRHPPPPLESFEIPPVTIHSPPPQPPQHQHQLLPQESFVN--------SNCG 772
Query: 241 RVSRK 245
+++ +
Sbjct: 773 KINSR 787
>TC82525 similar to GP|10177240|dbj|BAB10614.
emb|CAB89309.1~gene_id:MRN17.16~similar to unknown
protein {Arabidopsis thaliana}, partial (50%)
Length = 710
Score = 289 bits (739), Expect = 9e-79
Identities = 161/239 (67%), Positives = 181/239 (75%), Gaps = 5/239 (2%)
Frame = +1
Query: 1 MDYQRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLN 60
MD Q SLLN AYF QGKSME++RQSL+YTTLELEQTR QEEL K+D+QL+NLKDL+N
Sbjct: 85 MDNQSNSLLNWAYFCQGKSMEDLRQSLLYTTLELEQTRVTAQEELTKKDEQLMNLKDLIN 264
Query: 61 NTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDE--PRRGIDSNNG-- 116
N IRERDEAQEKCQRLLLEK +F QQQ DP SGVSSIEDE R+GIDSNNG
Sbjct: 265 NIIRERDEAQEKCQRLLLEKLVF------HQQQNDPLSGVSSIEDEQVTRKGIDSNNGFS 426
Query: 117 LSSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIE-LTPEKPLPEKGKLLQAVMKAGP 175
LSSSDCEESIVSSP+ D QSMIE LTP +PLPEKGKLLQAVMKAGP
Sbjct: 427 LSSSDCEESIVSSPIID---------------QSMIEVLTPNRPLPEKGKLLQAVMKAGP 561
Query: 176 LLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTS 234
LL+TLLLAGPLPQW+HPPPPLESFEIPPV+I P++LHQ+S F+ + T+
Sbjct: 562 LLKTLLLAGPLPQWKHPPPPLESFEIPPVSI-----------PRILHQDSIFSSNIDTT 705
>BG646469 similar to GP|10177240|db emb|CAB89309.1~gene_id:MRN17.16~similar
to unknown protein {Arabidopsis thaliana}, partial (40%)
Length = 663
Score = 229 bits (584), Expect = 8e-61
Identities = 129/200 (64%), Positives = 149/200 (74%), Gaps = 6/200 (3%)
Frame = +2
Query: 48 RDDQLLNLKDLLNNTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEP 107
+D+QL+NLKDL+NN IRERDEAQEKCQRLLLEK +F QQQ DP SGVSSIEDE
Sbjct: 146 KDEQLMNLKDLINNIIRERDEAQEKCQRLLLEKLVFH------QQQNDPLSGVSSIEDEQ 307
Query: 108 --RRGIDSNNG--LSSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIE-LTPEKPLPE 162
R+GIDSNNG LSSSDCEESIVSSP+ D QSMIE LTP +PLPE
Sbjct: 308 VTRKGIDSNNGFSLSSSDCEESIVSSPIID---------------QSMIEVLTPNRPLPE 442
Query: 163 KGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
KGKLLQAVMKAGPLL+TLLLAGPLPQW+HPPPPLESFEIPPV+I P++LH
Sbjct: 443 KGKLLQAVMKAGPLLKTLLLAGPLPQWKHPPPPLESFEIPPVSI-----------PRILH 589
Query: 223 QESFFNGS-SSTSTHSQCGR 241
Q+S F+ + +T+ +S CG+
Sbjct: 590 QDSIFSSNIDTTNANSHCGK 649
>BF646426 similar to GP|10177240|dbj emb|CAB89309.1~gene_id:MRN17.16~similar
to unknown protein {Arabidopsis thaliana}, partial (14%)
Length = 651
Score = 91.3 bits (225), Expect = 4e-19
Identities = 73/222 (32%), Positives = 104/222 (45%), Gaps = 24/222 (10%)
Frame = +3
Query: 9 LNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTIRERDE 68
L+ + +Q + E+RQ L+ TTLELE ++ +L NL L ERDE
Sbjct: 24 LDSVFIYQERE-HELRQKLIDTTLELEA--------MKNMKTELFNL---LKTAYLERDE 167
Query: 69 AQEKCQRLL--------LEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSS 120
A+ + Q+L+ +KF+ QQ+ N ++ ++G
Sbjct: 168 AKNQLQKLMNNPSPIHFQDKFISGFQQENPLMFHSSNPSITESSS-------LSHGSPPV 326
Query: 121 DCEESIVSSPVFDNMPQPQFP---------LPLPSVAQ-------SMIELTPEKPLPEKG 164
D VSSP F N L +PS ++ + EK LP+KG
Sbjct: 327 DSFFETVSSPEFSNTNMAYLNHHQNQHFNYLRVPSEKPVSDHGTVAIDSIAKEKVLPQKG 506
Query: 165 KLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTI 206
KLLQAV+ AGPLLQT+LLAGPLP W + PPPL ++PP+ I
Sbjct: 507 KLLQAVIDAGPLLQTILLAGPLPTWSN-PPPLNDIQVPPLNI 629
>BI271798 weakly similar to GP|10177240|dbj
emb|CAB89309.1~gene_id:MRN17.16~similar to unknown
protein {Arabidopsis thaliana}, partial (20%)
Length = 390
Score = 73.9 bits (180), Expect = 6e-14
Identities = 45/125 (36%), Positives = 72/125 (57%)
Frame = +3
Query: 1 MDYQRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLN 60
M+Y+ + L Y +E+++ SL+YTTLELE T + +EE+ +++ +L++ DLL+
Sbjct: 42 MEYECSPLTWEFYNHDEGLLEDLKHSLLYTTLELEATIVSAKEEITRKECELIHANDLLS 221
Query: 61 NTIRERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSS 120
I+ERDEA+ KCQ L+LEK Q+ Q D SG + + + +SS
Sbjct: 222 KVIKERDEARAKCQNLMLEK---------QEFQIDNKSGSEN---------EYSTQNASS 347
Query: 121 DCEES 125
DCEE+
Sbjct: 348 DCEEN 362
>BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (24%)
Length = 1059
Score = 42.7 bits (99), Expect = 1e-04
Identities = 27/66 (40%), Positives = 31/66 (46%), Gaps = 4/66 (6%)
Frame = -2
Query: 159 PLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQ----PP 214
P PE Q + P+ L P P R PPPPL S +IPP P+P LQ PP
Sbjct: 533 PSPEIICAAQFHISPPPVPPHPLPISPPPPLRPPPPPLPSPDIPPTPTPAPRLQLTPSPP 354
Query: 215 QQPPQL 220
PP L
Sbjct: 353 HAPPPL 336
Score = 32.3 bits (72), Expect = 0.19
Identities = 21/58 (36%), Positives = 25/58 (42%), Gaps = 2/58 (3%)
Frame = -1
Query: 182 LAGPLPQWR--HPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHS 237
L P+P PPPP F PP PSP PP L + N S TST++
Sbjct: 516 LRSPIPYLSTPRPPPPSPHFSPPPPPAPSP-------PPPLSRHPAHPNPRSPTSTYA 364
Score = 30.0 bits (66), Expect = 0.96
Identities = 16/36 (44%), Positives = 18/36 (49%), Gaps = 2/36 (5%)
Frame = -3
Query: 185 PLPQWRHPPPPLESFEI--PPVTIPSPPLQPPQQPP 218
PLP R P PPL F + PP T P + P PP
Sbjct: 418 PLPTSRPPQPPLPDFNLRPPPPTHLPP*IHLPTPPP 311
Score = 29.6 bits (65), Expect = 1.3
Identities = 35/130 (26%), Positives = 50/130 (37%), Gaps = 36/130 (27%)
Frame = -3
Query: 125 SIVSSPVFDNMPQPQFPLPLPSVAQSMIELT--PEKPLPEKGKLL----------QAVMK 172
S +S ++P+P+ PL LPS+ S + T + P + KL+ ++
Sbjct: 775 SRISHIASHSLPRPRAPLSLPSLHISPVPHTTSSNRTPPARTKLMVTSPRPHTPPNPIVT 596
Query: 173 AGPLLQTLLLA---GPLPQWRHP----------------PPPLESFEIPPVTIPSPPLQP 213
+ P L + A GP HP PP L F PP + P PP P
Sbjct: 595 SLPPLSLIK*ADVHGPWSFCYHPPKSFAQPNSISLHPPSPPTLSPFLPPPPSGPLPPPSP 416
Query: 214 -----PQQPP 218
P QPP
Sbjct: 415 LPTSRPPQPP 386
>TC79235 weakly similar to GP|7297741|gb|AAF52992.1| CG17108 gene product
{Drosophila melanogaster}, partial (21%)
Length = 648
Score = 40.8 bits (94), Expect = 5e-04
Identities = 21/54 (38%), Positives = 26/54 (47%), Gaps = 1/54 (1%)
Frame = -2
Query: 170 VMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQP-PQQPPQLLH 222
+M GP L P+P HPPPP + F PP PP P P PP ++H
Sbjct: 488 IMGRGPPLGPGPPPPPMPHSHHPPPPHDPFAPPPPVFHHPPPPPNPYAPPPVVH 327
Score = 30.4 bits (67), Expect = 0.74
Identities = 18/52 (34%), Positives = 21/52 (39%), Gaps = 16/52 (30%)
Frame = -2
Query: 183 AGPLPQWRHPPPPLESFEIPPVT-----------IPSPPLQ-----PPQQPP 218
A P P + HPPPP + PPV P PP PP +PP
Sbjct: 398 APPPPVFHHPPPPPNPYAPPPVVHHHHTPLHAHPAPPPPQPGHPGPPPPRPP 243
Score = 28.1 bits (61), Expect = 3.7
Identities = 14/33 (42%), Positives = 16/33 (48%)
Frame = -2
Query: 190 RHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
RH PPP PP+ PPL P PP + H
Sbjct: 518 RHNPPPPG----PPIMGRGPPLGPGPPPPPMPH 432
>BQ141909 weakly similar to GP|14253159|emb VMP3 protein {Volvox carteri f.
nagariensis}, partial (13%)
Length = 1223
Score = 40.0 bits (92), Expect = 0.001
Identities = 37/132 (28%), Positives = 54/132 (40%), Gaps = 2/132 (1%)
Frame = -3
Query: 89 QQQQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCEESIVSSPVFDNMPQPQFP--LPLPS 146
++ +DP+ + S+ ++S N S+S C S+ S P P P P LP+P
Sbjct: 546 RRSYSSDPSLSLRSLPPSHILCVNSCNDDSTSVCSNSLFSLP----FPPPLSPRLLPVPF 379
Query: 147 VAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTI 206
+ S+ L P P + + P LL PLP PPPL P+ +
Sbjct: 378 LP-SIFPLPPPSVPPPLSRFPLPFVPPPPFFPLPLLPPPLP-----PPPLL-----PLPL 232
Query: 207 PSPPLQPPQQPP 218
PP PP P
Sbjct: 231 SDPPSAPPPPSP 196
Score = 33.5 bits (75), Expect = 0.087
Identities = 24/66 (36%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Frame = -2
Query: 156 PEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPP- 214
P P+P L A +++ P L P + PPPP S PP PS PL+PP
Sbjct: 394 PPCPIPSLYFSLAAPLRSSPSLTLPPSFCSSPPFLPPPPPPPS-PPPPPPPPSSPLRPPL 218
Query: 215 QQPPQL 220
+ PP L
Sbjct: 217 RSPPPL 200
Score = 30.8 bits (68), Expect = 0.57
Identities = 28/83 (33%), Positives = 33/83 (39%), Gaps = 4/83 (4%)
Frame = -1
Query: 140 FPLPLP---SVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLL-LAGPLPQWRHPPPP 195
FPL P S++ S P +P P + A P L L L+ P P P PP
Sbjct: 419 FPLHFPPASSLSHSFPLFFPCRPPP-----FLPLSHASPFLLFLPPLSSPSPSSPLPSPP 255
Query: 196 LESFEIPPVTIPSPPLQPPQQPP 218
S P T P P PP PP
Sbjct: 254 PPSSLFPSPTPPPLPPPPPPPPP 186
Score = 29.6 bits (65), Expect = 1.3
Identities = 30/94 (31%), Positives = 39/94 (40%), Gaps = 4/94 (4%)
Frame = -2
Query: 129 SPVFDNMPQPQFPLPLPSVAQSM---IELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
SP + P P P+PS+ S+ + +P LP + + P L P
Sbjct: 427 SPFSPSTFPPPPPCPIPSLYFSLAAPLRSSPSLTLPP------SFCSSPPFLPP---PPP 275
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQP-PQQPP 218
P PPPP S PP+ P PPL P P PP
Sbjct: 274 PPSPPPPPPPPSSPLRPPLRSP-PPLPPLPPLPP 176
>TC80770 similar to GP|4680336|gb|AAD27627.1| hypothetical protein {Oryza
sativa subsp. indica} [Oryza sativa (indica
cultivar-group)], partial (7%)
Length = 507
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/94 (34%), Positives = 36/94 (38%), Gaps = 8/94 (8%)
Frame = -2
Query: 136 PQPQFPLPLPS---VAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWR-- 190
PQP P P PS + M+ PLP +L P PQ
Sbjct: 263 PQPPHPSPQPSPHPLLFFMLS*FSHPPLPT------------------ILPHPPPQPAPQ 138
Query: 191 ---HPPPPLESFEIPPVTIPSPPLQPPQQPPQLL 221
HPPP F I P P PP QPP QPP L+
Sbjct: 137 PPPHPPPQPPLFTIFPQPPPHPPPQPPPQPPLLI 36
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 38.9 bits (89), Expect = 0.002
Identities = 29/90 (32%), Positives = 36/90 (39%), Gaps = 1/90 (1%)
Frame = -2
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
P D P P P PLP ++ +P P P G A + P P P
Sbjct: 286 PPLDRAPPPP-PAPLPLASRLRPRASPPPPPPPPGPAASAPPRPPP--------PPPPPP 134
Query: 190 RHPPPPLESFEIPPVT-IPSPPLQPPQQPP 218
+ PPPP PP T P PP PP++ P
Sbjct: 133 QRPPPP-----PPPHTHQPEPPTSPPRRQP 59
Score = 32.0 bits (71), Expect = 0.25
Identities = 21/66 (31%), Positives = 26/66 (38%), Gaps = 3/66 (4%)
Frame = -1
Query: 156 PEKPLPEKGKLLQAVMKAG---PLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQ 212
P P PE ++ A G P L L+ P+ PPPP+ PP P
Sbjct: 674 PPPPPPEWERVPLAKKIGGQPRPALVPLVRVATYPRPPRPPPPVPPPPSPPRAPPQSEAA 495
Query: 213 PPQQPP 218
PP PP
Sbjct: 494 PPPPPP 477
Score = 27.7 bits (60), Expect = 4.8
Identities = 15/34 (44%), Positives = 15/34 (44%), Gaps = 3/34 (8%)
Frame = -3
Query: 189 WRHPP---PPLESFEIPPVTIPSPPLQPPQQPPQ 219
W PP P S IPP T P P P PPQ
Sbjct: 621 WAAPPGARPSRPSRYIPPPTSPPPASPPAPVPPQ 520
Score = 27.3 bits (59), Expect = 6.3
Identities = 13/37 (35%), Positives = 16/37 (43%)
Frame = -2
Query: 182 LAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
L P P+ PPP PP P+PP P + P
Sbjct: 514 LHNPRPRPPPRPPPAAPARAPPSGPPAPPTPPATRAP 404
Score = 26.9 bits (58), Expect = 8.2
Identities = 14/33 (42%), Positives = 16/33 (48%)
Frame = -3
Query: 190 RHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
RH PPP PP+ P PP PP+ P H
Sbjct: 207 RHRPPPPGPPPPPPLG-PRPPPHPPRSGPPRPH 112
>BF646401 similar to PIR|A02951|KRMS keratin type II cytoskeletal - mouse
(fragment), partial (5%)
Length = 557
Score = 38.1 bits (87), Expect = 0.004
Identities = 34/123 (27%), Positives = 50/123 (40%), Gaps = 21/123 (17%)
Frame = +2
Query: 120 SDCEESIVSSPVFDNMPQPQ-FPLPLPSVAQ----SMIELTPEKPLPEKGK--------L 166
S C SI SP P P+ F L ++Q + + L +P PE +
Sbjct: 5 SSCSRSIFLSPQLSLHPLPEPFSLTRDLLSQPRAAAKLSLPSMRPPPEPPPQNNQTVVTI 184
Query: 167 LQAVMKAGPLLQTL-LLAGPLPQWRHP------PPPLESFEIPPVTIPSPPL-QPPQQPP 218
++ + Q L L+ P P W P PPP+ S + + P P L PP +PP
Sbjct: 185 FSLILNPKTMFQALPLMVAPPPSWLAPTRPPLEPPPISSTTVKFIVPPDPDLCCPPPKPP 364
Query: 219 QLL 221
L+
Sbjct: 365 WLI 373
>TC80199 weakly similar to GP|21593112|gb|AAM65061.1 unknown {Arabidopsis
thaliana}, partial (41%)
Length = 934
Score = 35.8 bits (81), Expect = 0.018
Identities = 33/123 (26%), Positives = 55/123 (43%), Gaps = 5/123 (4%)
Frame = +2
Query: 119 SSDCEESIVSSPVFDNM-PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAG--P 175
SS+ + S P+ M PQ + PL + ++ ++ +T E KL++ G P
Sbjct: 23 SSNPYTILTSIPIKTIMDPQSKIPLRILNLILLLVCVTLPINAMESRKLVEKCTPCGDTP 202
Query: 176 LLQT--LLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSST 233
+ ++ P P +P PP S PP+ SPP P++PP + SSS+
Sbjct: 203 IYSPPPIIYNSPPPPIVYPSPPPPSPSPPPIVYYSPPPPSPKKPPSANCPPPPSSPSSSS 382
Query: 234 STH 236
S +
Sbjct: 383 SPY 391
>TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19.70 -
Arabidopsis thaliana, partial (15%)
Length = 716
Score = 35.8 bits (81), Expect = 0.018
Identities = 29/127 (22%), Positives = 44/127 (33%)
Frame = +1
Query: 88 QQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCEESIVSSPVFDNMPQPQFPLPLPSV 147
+ + +P +E P +N+ S+ + + QP P+ LP
Sbjct: 178 RDEPMYVEPQEVRMKLEPPPPVAFANNDSTVPPVATTSLPEPLPYQHKEQPPIPVTLP-- 351
Query: 148 AQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIP 207
P PL + + + P L L P HPPPP S ++PP
Sbjct: 352 --------PPPPLSKLPPATREQLPPPPPLSKL----PPAAREHPPPPPPSAKLPPAARE 495
Query: 208 SPPLQPP 214
PL PP
Sbjct: 496 RLPLPPP 516
>TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {Arabidopsis
thaliana}, partial (92%)
Length = 1250
Score = 35.8 bits (81), Expect = 0.018
Identities = 29/83 (34%), Positives = 34/83 (40%), Gaps = 1/83 (1%)
Frame = -2
Query: 138 PQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPL-LQTLLLAGPLPQWRHPPPPL 196
P P PLP P PLP PL L + L+ PLP PPPL
Sbjct: 268 PFMPPPLPPPRPPPRPPLPPPPLPP------------PLALNGVPLSPPLPPKPPLPPPL 125
Query: 197 ESFEIPPVTIPSPPLQPPQQPPQ 219
PP++ P PPL PP P+
Sbjct: 124 -----PPLSPPLPPLNPPPPRPR 71
Score = 30.4 bits (67), Expect = 0.74
Identities = 15/28 (53%), Positives = 15/28 (53%)
Frame = -2
Query: 193 PPPLESFEIPPVTIPSPPLQPPQQPPQL 220
PPPL PP P PPL PP PP L
Sbjct: 259 PPPLP----PPRPPPRPPLPPPPLPPPL 188
Score = 28.1 bits (61), Expect = 3.7
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = -2
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAV 170
P+P P PLP ++ + L P P P G + V
Sbjct: 151 PKPPLPPPLPPLSPPLPPLNPPPPRPRTGGAIAVV 47
Score = 27.7 bits (60), Expect = 4.8
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -2
Query: 196 LESFEIPPVTIPSPPLQPPQQPPQL 220
L F PP+ P PP +PP PP L
Sbjct: 274 LPPFMPPPLPPPRPPPRPPLPPPPL 200
>TC86556 similar to PIR|T07598|T07598 proline-rich protein GPP1 - potato,
partial (24%)
Length = 1250
Score = 35.8 bits (81), Expect = 0.018
Identities = 27/93 (29%), Positives = 41/93 (44%), Gaps = 3/93 (3%)
Frame = +3
Query: 129 SPVFDNMPQPQFPLPLPSVAQSMIELTP--EKPLPEKGKLLQAVMKAGPLLQTLLLAGPL 186
+PV++ P P+P + + TP EKPLP + + + L P+
Sbjct: 30 TPVYEK----PLPPPVPVYHKPLPPPTPVYEKPLPPPVPVYHKPLPPPTPVYHKPLPPPV 197
Query: 187 PQWRH-PPPPLESFEIPPVTIPSPPLQPPQQPP 218
P + PPP +E +PP + PP P QPP
Sbjct: 198 PVKKPCPPPKVEHPVLPPAPVYKPPPVPKPQPP 296
Score = 34.7 bits (78), Expect = 0.039
Identities = 31/92 (33%), Positives = 39/92 (41%), Gaps = 6/92 (6%)
Frame = +3
Query: 136 PQP-----QFPLPLPSVAQSMIELTPE-KPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PQP + P P P V ++ P KP P V KA P + P P+
Sbjct: 285 PQPPPVPVKKPCPPPKVEHPVLPPVPVYKPPP--------VPKALPPPVPIKKPCP-PKI 437
Query: 190 RHPPPPLESFEIPPVTIPSPPLQPPQQPPQLL 221
HP P PPV IP PP+ P +PP +L
Sbjct: 438 EHPVLPPVPIYKPPVVIPKPPVVPIYKPPVIL 533
>TC76827 similar to GP|17154773|gb|AAL35979.1 extensin-like protein {Cucumis
sativus}, partial (60%)
Length = 1093
Score = 35.4 bits (80), Expect = 0.023
Identities = 30/106 (28%), Positives = 46/106 (43%), Gaps = 7/106 (6%)
Frame = +3
Query: 141 PLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPP--PPLES 198
P+ P + + + + P P+P V+ P+ TL + LP PP P +
Sbjct: 228 PIVKPPIVKPPVTIPPTLPIPP-------VLPHLPIPPTLPIPPVLPHLPIPPTLPIPPT 386
Query: 199 FEIPPV-----TIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQC 239
IPP T+P PP+ P P +L+ S +G S+ S S C
Sbjct: 387 LPIPPTLPIPPTLPIPPVLPHLPVPPVLNPPS--SGGSTPSPTSPC 518
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 35.4 bits (80), Expect = 0.023
Identities = 23/48 (47%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Frame = +1
Query: 177 LQTLLLAGPLPQWRHPPPPLESFEIPPVTIPS-PPLQPPQQPPQLLHQ 223
LQT L PLPQ PP + PP P PP +PPQ+PPQ L Q
Sbjct: 250 LQTRSL--PLPQRPPQRPPQRPPQKPPQRPPQKPPQRPPQRPPQKLPQ 387
Score = 29.6 bits (65), Expect = 1.3
Identities = 16/39 (41%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Frame = +1
Query: 187 PQWRHPPPPLESFEIPPVTIPS-PPLQPPQQPPQLLHQE 224
PQ PP + PP +P PP + PQ+PPQ L Q+
Sbjct: 322 PQRPPQKPPQRPPQRPPQKLPQRPPQKLPQRPPQKLPQK 438
Score = 27.7 bits (60), Expect = 4.8
Identities = 13/29 (44%), Positives = 18/29 (61%), Gaps = 2/29 (6%)
Frame = +1
Query: 193 PPPLESFEIPPVTIPS--PPLQPPQQPPQ 219
P PL++ +P P PP +PPQ+PPQ
Sbjct: 241 PIPLQTRSLPLPQRPPQRPPQRPPQKPPQ 327
>TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 -
Arabidopsis thaliana, partial (3%)
Length = 757
Score = 35.4 bits (80), Expect = 0.023
Identities = 38/118 (32%), Positives = 50/118 (42%), Gaps = 9/118 (7%)
Frame = +1
Query: 138 PQFPLP-LPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPL------PQWR 190
P LP LPS ++ L P+P A+M PL L+L PL P
Sbjct: 214 PALVLPVLPSPVLALPVLPSPVPVP-LALPSPALMPVAPLSPALVLPAPLSPALVLPVLP 390
Query: 191 HPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSS--TSTHSQCGRVSRKR 246
PPP + IP P+PP PP P L + +SS +S+ S+ G SR R
Sbjct: 391 SPPPRPAALLIPD---PAPPTSPPPAPVSLSPASAPLPSASSPASSSFSKGGCKSRGR 555
>TC85604 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago truncatula},
partial (47%)
Length = 790
Score = 35.4 bits (80), Expect = 0.023
Identities = 29/105 (27%), Positives = 49/105 (46%), Gaps = 7/105 (6%)
Frame = +1
Query: 123 EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPE-KPLPEKGKLLQAVMKAGPLLQTLL 181
E+ V P + P + P+ P V + ++E P KP EK + + ++ PL + +
Sbjct: 10 EKPPVHKPPVEKPPVHKSPVEKPPVHKPLVEKLPVYKPSVEKPPVYKPPVEKPPLHKPPV 189
Query: 182 LAGPL--PQWRHPP---PPLESFEIPPVTIPSPPL-QPPQQPPQL 220
P+ P PP PP+E + +I PP+ +PP + P L
Sbjct: 190 EKPPVHKPPVEKPPVHKPPVEKLPVYKPSIEKPPVYKPPVEKPPL 324
Score = 27.3 bits (59), Expect = 6.3
Identities = 25/100 (25%), Positives = 39/100 (39%), Gaps = 8/100 (8%)
Frame = +1
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTP-EKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
V P P + P+ PS+ + + P EKP K + + M P+ + + P
Sbjct: 217 VEKPPVHKPPVEKLPVYKPSIEKPPVYKPPVEKPPLHKPPVEKPPMHKPPVEKPPVHKPP 396
Query: 186 LPQWRHPPPPLES-------FEIPPVTIPSPPLQPPQQPP 218
+ + PP+E E PPV P P +PP
Sbjct: 397 VEKLPVYKPPVEKPPVYKPHVEKPPVNKPPVEKPPVHKPP 516
Score = 27.3 bits (59), Expect = 6.3
Identities = 25/99 (25%), Positives = 41/99 (41%), Gaps = 3/99 (3%)
Frame = +1
Query: 123 EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTP-EKPLPEKGKLLQAVMKAGPLLQTLL 181
E+ V P + P + P+ P + + +E P KP EK + + ++ P+ + +
Sbjct: 250 EKLPVYKPSIEKPPVYKPPVEKPPLHKPPVEKPPMHKPPVEKPPVHKPPVEKLPVYKPPV 429
Query: 182 LAGPL--PQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P+ P PP E PPV P P +PP
Sbjct: 430 EKPPVYKPHVEKPPVNKPPVEKPPVHKPPVEKPPVHKPP 546
>TC85605 similar to GP|437310|gb|AAA62850.1|| nodulin {Medicago truncatula},
partial (62%)
Length = 1077
Score = 35.4 bits (80), Expect = 0.023
Identities = 29/105 (27%), Positives = 49/105 (46%), Gaps = 7/105 (6%)
Frame = +3
Query: 123 EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPE-KPLPEKGKLLQAVMKAGPLLQTLL 181
E+ V P + P + P+ P V + ++E P KP EK + + ++ PL + +
Sbjct: 366 EKPPVHKPPVEKPPVHKSPVEKPPVHKPLVEKLPVYKPSVEKPPVYKPPVEKPPLHKPPV 545
Query: 182 LAGPL--PQWRHPP---PPLESFEIPPVTIPSPPL-QPPQQPPQL 220
P+ P PP PP+E + +I PP+ +PP + P L
Sbjct: 546 EKPPVHKPPVEKPPVHKPPVEKLPVYKPSIEKPPVYKPPVEKPPL 680
Score = 29.6 bits (65), Expect = 1.3
Identities = 24/90 (26%), Positives = 37/90 (40%), Gaps = 3/90 (3%)
Frame = +3
Query: 132 FDNMPQPQFPLPLPSVAQSMIELTP-EKPLPEKGKLLQAVMKAGPLLQTLLLAGPL--PQ 188
++ P+ + P+ P + IE P KPL EK + ++ P+ + + P P
Sbjct: 123 YEKPPEYKPPVQNPQFYKPHIEKPPVHKPLDEKPPVHHPPIEKPPIYKPPVEKPPAYKPP 302
Query: 189 WRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
H P E PPV P P +PP
Sbjct: 303 VEHHPVYRPHVEKPPVNKPPVEKPPVHKPP 392
Score = 27.3 bits (59), Expect = 6.3
Identities = 25/100 (25%), Positives = 39/100 (39%), Gaps = 8/100 (8%)
Frame = +3
Query: 127 VSSPVFDNMPQPQFPLPLPSVAQSMIELTP-EKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
V P P + P+ PS+ + + P EKP K + + M P+ + + P
Sbjct: 573 VEKPPVHKPPVEKLPVYKPSIEKPPVYKPPVEKPPLHKPPVEKPPMHKPPVEKPPVHKPP 752
Query: 186 LPQWRHPPPPLES-------FEIPPVTIPSPPLQPPQQPP 218
+ + PP+E E PPV P P +PP
Sbjct: 753 VEKLPVYKPPVEKPPVYKPHVEKPPVNKPPVEKPPVHKPP 872
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,324,394
Number of Sequences: 36976
Number of extensions: 193989
Number of successful extensions: 5078
Number of sequences better than 10.0: 363
Number of HSP's better than 10.0 without gapping: 2232
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3684
length of query: 266
length of database: 9,014,727
effective HSP length: 94
effective length of query: 172
effective length of database: 5,538,983
effective search space: 952705076
effective search space used: 952705076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0369b.1


