

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
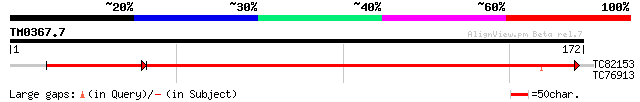
Query= TM0367.7
(172 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC82153 similar to SP|O81263|KITH_ORYSA Thymidine kinase (EC 2.7... 201 9e-61
TC76913 similar to SP|O04450|TCPE_ARATH T-complex protein 1 eps... 27 3.1
>TC82153 similar to SP|O81263|KITH_ORYSA Thymidine kinase (EC 2.7.1.21).
[Rice] {Oryza sativa}, partial (68%)
Length = 731
Score = 201 bits (512), Expect(2) = 9e-61
Identities = 98/131 (74%), Positives = 112/131 (84%), Gaps = 1/131 (0%)
Frame = +3
Query: 42 GVDAYEQLDVIGIDEAQFFDDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPL 101
G +AY++LDVIGIDEAQFF+DLYDFC +AAD DGK VVVAGLDG+Y+RR+FGSVL IIP+
Sbjct: 93 GHEAYQKLDVIGIDEAQFFEDLYDFCCKAADEDGKIVVVAGLDGDYMRRSFGSVLHIIPI 272
Query: 102 ADSVTKLTARCEICGKHAFFTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETT-RL 160
AD+VTKLTARCE+CGK AFFTLRKT E Q ELIGG D+YMPVCR HYVNGQV +E +
Sbjct: 273 ADTVTKLTARCELCGKRAFFTLRKTGEKQTELIGGADLYMPVCRHHYVNGQVVVEAAMKS 452
Query: 161 VLESQKVECGS 171
V SQKV CGS
Sbjct: 453 VFGSQKVGCGS 485
Score = 48.5 bits (114), Expect(2) = 9e-61
Identities = 19/30 (63%), Positives = 23/30 (76%)
Frame = +2
Query: 12 RYGLDSIVTHDGTKLPCWALANLSSFKQKF 41
RY +DS+VTHDG K PCWAL +L FK K+
Sbjct: 2 RYAVDSVVTHDGIKFPCWALPDLMLFKDKY 91
>TC76913 similar to SP|O04450|TCPE_ARATH T-complex protein 1 epsilon
subunit (TCP-1-epsilon) (CCT-epsilon). [Mouse-ear
cress], complete
Length = 1965
Score = 27.3 bits (59), Expect = 3.1
Identities = 12/43 (27%), Positives = 22/43 (50%)
Frame = +1
Query: 38 KQKFGVDAYEQLDVIGIDEAQFFDDLYDFCREAADHDGKTVVV 80
K K +D E+ + E ++FDD+ C++ G T+V+
Sbjct: 883 KHKVDIDTVEKFQTLRKQEQKYFDDMVQQCKDV----GATLVI 999
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,778,773
Number of Sequences: 36976
Number of extensions: 56480
Number of successful extensions: 211
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 211
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 211
length of query: 172
length of database: 9,014,727
effective HSP length: 89
effective length of query: 83
effective length of database: 5,723,863
effective search space: 475080629
effective search space used: 475080629
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0367.7


