

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
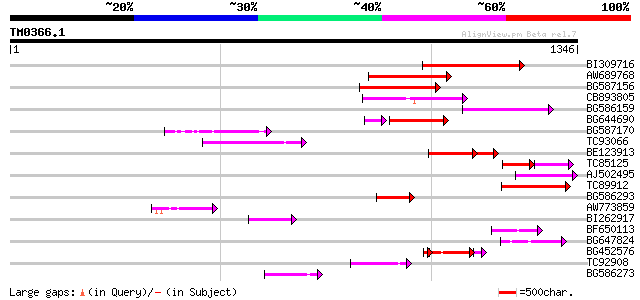
Query= TM0366.1
(1346 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 270 4e-72
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 195 1e-49
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 182 1e-45
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 179 8e-45
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 169 9e-42
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 125 4e-37
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 139 7e-33
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 138 1e-32
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 94 3e-28
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 67 6e-25
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 113 6e-25
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 110 3e-24
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 104 2e-22
AW773859 99 8e-21
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 93 8e-19
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 92 1e-18
BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F... 91 4e-18
BG452576 weakly similar to GP|12005223|gb|A reverse transcriptas... 71 1e-16
TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol po... 82 1e-15
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 80 4e-15
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 270 bits (689), Expect = 4e-72
Identities = 132/241 (54%), Positives = 173/241 (71%)
Frame = +2
Query: 981 QVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDV 1040
+VC+L KS+YGLKQASR+WY KLS L + G+ Q++SD SLF KF+ SSFT LLVYVDD+
Sbjct: 17 KVCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDI 196
Query: 1041 ILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETG 1100
+L GN ++E Q VK L F IKDLG L+YFLGLEVA S GI L QRKY L+LL+++G
Sbjct: 197 VLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSG 376
Query: 1101 TLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMS 1160
L K TP D S++L Y+D YRRL+G+L+YLTTTRPDIS A QQLSQ++S
Sbjct: 377 NLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVS 556
Query: 1161 NPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYI 1220
P H++AA RVL+YLK +P GL + S + + ++D+DWA CP TR+S++GY ++
Sbjct: 557 KPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVFL 736
Query: 1221 G 1221
G
Sbjct: 737 G 739
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 195 bits (495), Expect = 1e-49
Identities = 96/196 (48%), Positives = 131/196 (65%)
Frame = +1
Query: 853 DLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKG 912
D W +AM+ E KAL N TW LV LP K IG KWVYRVK N DG++ ++KARLVAKG
Sbjct: 88 DPRWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKG 267
Query: 913 YNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPA 972
++Q G DY +TFSPV K T+R+IL +A + W + Q+D++NAFL+G L E+VYM+ P
Sbjct: 268 FSQTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQ 447
Query: 973 GVPSVKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTG 1032
G + + VCKL KSLYGLKQA R WYE L++ GF ++ D SL + Q +
Sbjct: 448 GFEAANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIY 627
Query: 1033 LLVYVDDVILFGNTVT 1048
L +YVDD+++ G++ +
Sbjct: 628 LXIYVDDILITGSSAS 675
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 182 bits (461), Expect = 1e-45
Identities = 86/194 (44%), Positives = 130/194 (66%), Gaps = 1/194 (0%)
Frame = -1
Query: 830 AHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKW 889
AH A+ +N+ ++ P ++ EA +D W+E++ AE A+ KN TW +LP+G K + ++W
Sbjct: 600 AHCAFMVNLNENHIPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKKAVSSRW 421
Query: 890 VYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLH 949
++ +K DG++ R K RLVA+G+ G DY +TF+PVAKL T+R++L+LA + W L
Sbjct: 420 IFTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVNLGWGLW 241
Query: 950 QLDVDNAFLHGNLDEDVYMTIPAGVPS-VKPNQVCKLLKSLYGLKQASRKWYEKLSAHLE 1008
Q+DV NAFL G L+++VYM P G+ VK V +L K++YGLKQ+ R WY KLS L
Sbjct: 240 QMDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNKLSTTLN 61
Query: 1009 TLGFKQTASDHSLF 1022
GF+++ DH+LF
Sbjct: 60 GRGFRKSELDHTLF 19
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 179 bits (453), Expect = 8e-45
Identities = 96/258 (37%), Positives = 150/258 (57%), Gaps = 8/258 (3%)
Frame = +3
Query: 838 IAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNV 897
+ +P+TF EA K WR +M E++A E+N TW+L DL G K IG KW+++ K N
Sbjct: 24 LTMTSDPTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNE 203
Query: 898 DGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAF 957
+G + +YKARLVAKGY+Q G+DY + F+PVA+ T+R+++ALAA Q+ D
Sbjct: 204 NGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAA-------QIKRDGVC 362
Query: 958 L------HGNLDEDVYMTIPAGVPSVKPNQVC-KLLKSLYGLKQASRKWYEKLSAHLETL 1010
+ H ++ + + + + ++ ++LYGLKQA R WY ++ A+
Sbjct: 363 IS*M*KAHSCMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKE 542
Query: 1011 GFKQTASDHSLFVKF-QGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVL 1069
GF++ +H+LFVK +G + +YVDD+I GN F+ K S+ + F + DLG +
Sbjct: 543 GFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKM 722
Query: 1070 KYFLGLEVAHSTSGISLC 1087
YFLG+EV + GI +C
Sbjct: 723 HYFLGVEVTQNEKGIYIC 776
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 169 bits (427), Expect = 9e-42
Identities = 80/217 (36%), Positives = 130/217 (59%)
Frame = +1
Query: 1075 LEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRL 1134
+EV + GI +CQRKY DLL+ G S P+ P +L +++ D Y+++
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 1135 VGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTIN 1194
VG L+YL TRPD+ + +S++M+ P + H A +RVLRYL + LG+++ RN +
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 1195 IQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCEL 1254
++ Y+D+D+AG D R+S SGY F + VSW +KKQ V+ S+ +AE+ A A+ C+
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 1255 QWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFH 1291
W+ +L+ L T + + ++CDN S + ++ NPV H
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 125 bits (315), Expect(2) = 4e-37
Identities = 63/141 (44%), Positives = 96/141 (67%), Gaps = 1/141 (0%)
Frame = -2
Query: 901 LARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHG 960
+ R K++LV +GYNQ EG+DY + FSPVA++ +R+++A AA + L+Q+DV +AF++G
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 961 NLDEDVYMTIPAGVPSVK-PNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDH 1019
+L E+V++ P G + PN V +L K+LYGLKQA R WYE+LS L GFK+ D+
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 1020 SLFVKFQGSSFTGLLVYVDDV 1040
+LF+ + + VYVDD+
Sbjct: 64 TLFLLKRE*ELLIIQVYVDDI 2
Score = 48.9 bits (115), Expect(2) = 4e-37
Identities = 22/52 (42%), Positives = 29/52 (55%)
Frame = -3
Query: 843 EPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVK 894
EP EA +D W +MQ E+ E++ W LV PEG IG +WV+R K
Sbjct: 588 EPKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRWVFRNK 433
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 139 bits (350), Expect = 7e-33
Identities = 91/258 (35%), Positives = 141/258 (54%), Gaps = 3/258 (1%)
Frame = -3
Query: 367 VQTQRRIGLGSLQPDELYHLVVSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGHL 426
+++ + IG G + D LY L S + +S+S +N ALWH RLGH
Sbjct: 710 IESSQLIGEGVTKGD-LYMLEKLDPVSN-YKCSFTSSSSLNK-------DALWHARLGH- 561
Query: 427 SHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVHMDIWGPLAQ 486
H R L L + + +K+ C+ C L K + +F + + + CFDL++ D+W
Sbjct: 560 PHGRALNL-MLPGVVFENKN--CEACILGKHCKNVFPRTSTVYENCFDLIYTDLW-TAPS 393
Query: 487 ASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKAIRSDNGPE 546
S NHKYF+T +D+ S++ W+ L+ +K V KNF V + +K +RSDNG E
Sbjct: 392 LSRDNHKYFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGE 213
Query: 547 FLLPAFYS---AQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSV 603
+ AF S GI+HQ SC TPQQNG +RK++H++ +AR+L+FQ+ +V
Sbjct: 212 YTSYAFKSHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQA---------NV 60
Query: 604 LHAVFLMNRIPSKLLKNK 621
A +L+N IP+K+L+ K
Sbjct: 59 STACYLINWIPTKVLRIK 6
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 138 bits (348), Expect = 1e-32
Identities = 90/256 (35%), Positives = 130/256 (50%), Gaps = 7/256 (2%)
Frame = +1
Query: 457 QKRKMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGE 516
+K+ F+ + + + D +H D+WGP S +Y +T++DD+ R VWV L K E
Sbjct: 49 RKKVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNE 228
Query: 517 VQQQVKNFITLVKTQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGR 573
K + LV+TQ G+ VK + +DN EF F + GI K+ PQQNG
Sbjct: 229 TFPTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGV 408
Query: 574 VERKHQHILNIARALLFQSKL--PKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREA 631
ER + +L AR +L + L + +W + A L+NR P L K P ++
Sbjct: 409 AERMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGNL 588
Query: 632 VDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVL--MDLITQEIFISR 689
VD +++FG C A L N+ KL PRA +CIFL Y KGY L D +Q++ +SR
Sbjct: 589 VDYSNLRIFG--CPAYALVND-GKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLILSR 759
Query: 690 DVIFHEHVLPYKGSNS 705
DV F+E L G S
Sbjct: 760 DVTFNEDALLSSGKQS 807
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 93.6 bits (231), Expect(2) = 3e-28
Identities = 49/118 (41%), Positives = 74/118 (62%), Gaps = 1/118 (0%)
Frame = +1
Query: 994 QASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLL-VYVDDVILFGNTVTEFQL 1052
Q+ R W+++ + ++ G+ Q +DH++F+K + +L VYVDD+ L G+ +
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1053 VKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATP 1110
+K+ L + F IKDLG LKYFLG+EVA G S+ QRKY LDLL+ET +G K + P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Score = 51.6 bits (122), Expect(2) = 3e-28
Identities = 25/58 (43%), Positives = 35/58 (60%)
Frame = +2
Query: 1102 LGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYM 1159
L KP TP+D +++L D Y+RLVG+L+YL+ TRPDIS +SQ+M
Sbjct: 329 LDVKPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 67.0 bits (162), Expect(2) = 6e-25
Identities = 37/92 (40%), Positives = 52/92 (56%)
Frame = +3
Query: 1246 ALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVR 1305
+L A E W+ L+++L V +CD+QSALHIA NP FH RTKH+ I H VR
Sbjct: 228 SLPQACKEAIWMQRLMEELGHKQEQITV-YCDSQSALHIARNPAFHSRTKHIGIQYHFVR 404
Query: 1306 EKLQAGILKLLPIPTTLQVADVFTKALQPRVF 1337
E ++ G + + I T +AD TK++ F
Sbjct: 405 EVVEEGSVDMQKIHTNDNLADAMTKSINTDKF 500
Score = 67.0 bits (162), Expect(2) = 6e-25
Identities = 32/76 (42%), Positives = 50/76 (65%)
Frame = +1
Query: 1171 QRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAK 1230
+R++RY+K + G+ + F S + ++GY D+D+AG D R+S +GY F + VSW +K
Sbjct: 4 KRIMRYIKGTSGVAVCFG-GSELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLSK 180
Query: 1231 KQTTVSRSSNEAEYRA 1246
QT V+ S+ EAEY A
Sbjct: 181 LQTVVALSTTEAEYMA 228
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 113 bits (282), Expect = 6e-25
Identities = 56/146 (38%), Positives = 85/146 (57%)
Frame = +2
Query: 1201 ADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYL 1260
+DWAG +TR+S SGY F++G +SW +KKQ V+ S+ EAEY A + WL +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1261 LQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPT 1320
L+ + ++CDN+SA+ ++ NPVFH R+KH+DI H +RE + + + PT
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1321 TLQVADVFTKALQPRVFQGFATKLAM 1346
++AD+FTK L+ F L M
Sbjct: 362 EEKIADIFTKPLKIESFYKLKKMLGM 439
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 110 bits (276), Expect = 3e-24
Identities = 60/165 (36%), Positives = 101/165 (60%), Gaps = 2/165 (1%)
Frame = +1
Query: 1168 KAAQRVLRYLKASPGLGLLFPRNSTIN--IQGYSDADWAGCPDTRRSISGYCFYIGRSLV 1225
+A + VL+YL S L + + + ++GY DAD+AG DTR+S+SG+ F + + +
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 1226 SWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIA 1285
SWKA +Q+ V+ S+ +AEY A + WL ++ +L +T + CD+QSA+H+A
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKI-HCDSQSAIHLA 357
Query: 1286 ANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTK 1330
+ V+HERTKH+DI H +R+ +++ + + + + ADVFTK
Sbjct: 358 NHQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 104 bits (260), Expect = 2e-22
Identities = 48/90 (53%), Positives = 69/90 (76%)
Frame = +2
Query: 872 TWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKL 931
T KLV P GVKPIG +W+Y++KRN DGTL +YKARLVAKGY + +G+D+ + F+PV ++
Sbjct: 56 TLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVVRI 235
Query: 932 TTVRVILALAASQNWHLHQLDVDNAFLHGN 961
T+ ++LALAA+ +H +DV AFL+G+
Sbjct: 236 ETI*LLLALAATNGC*IHHIDVKIAFLNGH 325
>AW773859
Length = 538
Score = 99.4 bits (246), Expect = 8e-21
Identities = 59/172 (34%), Positives = 97/172 (56%), Gaps = 14/172 (8%)
Frame = -3
Query: 336 QKMVVLL*MVVL-------L*IMW*RLLLAK-------EMLQDMEVQTQRRIGLGSLQPD 381
Q++V +L ++VL *I W LL+ + +++ E ++ + IG G L +
Sbjct: 506 QRVVGVLSLMVLPRSNMNTW*ICWLLLLVPQVRWFHTLQVILIQEKRSLKMIGSGELF-E 330
Query: 382 ELYHLVVSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGHLSHDRLLALHTVQNSI 441
LY L + S + SS + +P ALWHFRLGHLS+ +LL+LH+ I
Sbjct: 329 GLYFLTTQDTPSATIA---SSQVQPQPQPTFLPQEALWHFRLGHLSNRKLLSLHSNFPFI 159
Query: 442 SVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVHMDIWGPLAQASVHNHK 493
++ ++ VCD+CH ++ K+ F +S ++A KC++L H DIWGP + S+HN +
Sbjct: 158 TIDQNSVCDICHYSRHKKLPFQLSTNRASKCYELFHFDIWGPFSTQSIHNQR 3
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 92.8 bits (229), Expect = 8e-19
Identities = 46/113 (40%), Positives = 68/113 (59%)
Frame = +1
Query: 567 TPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYEL 626
TPQQNG ER ++ +L RA+L + + K W +V A +++NR PS ++ K+P E+
Sbjct: 85 TPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLKTPMEM 264
Query: 627 LYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMD 679
+ VD + VFG + + R+KLDP++RKCIFLGY +KGY L D
Sbjct: 265 WKGKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYXLWD 423
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 92.4 bits (228), Expect = 1e-18
Identities = 50/124 (40%), Positives = 73/124 (58%), Gaps = 3/124 (2%)
Frame = +1
Query: 1145 RPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFP---RNSTINIQGYSDA 1201
RPDI ++ +S++M +P H AA R+LRY++ + GLLFP ++ + YSD+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 1202 DWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLL 1261
DW G RRS SGY F + +SW KKQ + SS EAEY A +AT + WL ++
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 1262 QDLK 1265
++LK
Sbjct: 472 KELK 483
>BG647824 weakly similar to PIR|G96722|G96 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (5%)
Length = 721
Score = 90.5 bits (223), Expect = 4e-18
Identities = 61/158 (38%), Positives = 86/158 (53%), Gaps = 1/158 (0%)
Frame = -3
Query: 1166 HFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLV 1225
H +AA ++L YLK SP + FP S I I+ + D+D F +
Sbjct: 590 HPQAAIQIL-YLKISPSQ*IFFP--S*IQIKAFCDSD*IDQAA*TLENQSVIFASS*ATH 420
Query: 1226 SWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSAL-HI 1284
S+ + + N YR++ CE++WL YLL DLK T +L+CDNQSA HI
Sbjct: 419 SYAGNLK---KKRYNFKIYRSI*STICEIKWLTYLLNDLKFTFIKPAMLYCDNQSAARHI 249
Query: 1285 AANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTL 1322
AAN F ERTKH+++DCH+VR KLQ + +L I ++L
Sbjct: 248 AANSSFLERTKHIELDCHIVRVKLQLKLFHILHILSSL 135
>BG452576 weakly similar to GP|12005223|gb|A reverse transcriptase-like protein
{Spiranthes hongkongensis}, partial (25%)
Length = 676
Score = 70.9 bits (172), Expect(3) = 1e-16
Identities = 47/106 (44%), Positives = 67/106 (62%)
Frame = +2
Query: 997 RKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDS 1056
RK Y KL + L +LG+KQ+ +DHS+F G FT LVY+DD++ + +E QLVK
Sbjct: 236 RK*Y*KLLSTLISLGYKQSPNDHSIFSF--GRRFTIFLVYLDDIVFARDDHSETQLVKSH 409
Query: 1057 LHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTL 1102
L + F I LG L Y +G+++A S S I Q KY ++LL+E+G L
Sbjct: 410 LDKNFIIIGLGTLHY-VGVKIA*SES*IIDDQCKYTIELLEESGQL 544
Score = 27.7 bits (60), Expect(3) = 1e-16
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = +3
Query: 982 VCKLLKSLYGLKQASRKWYEK 1002
VCK S+YGLK A R+ Y K
Sbjct: 165 VCKFHNSIYGLKHAHRQ*YSK 227
Score = 26.9 bits (58), Expect(3) = 1e-16
Identities = 14/34 (41%), Positives = 17/34 (49%), Gaps = 4/34 (11%)
Frame = +3
Query: 1102 LGSKPVATPLDPSIRLS----QEQGKPYDDPAAY 1131
LG KP +TP DPS++L E PY Y
Sbjct: 546 LGMKPSSTPRDPSLKLKLYNIYETSSPYHVKLGY 647
>TC92908 similar to GP|10140704|gb|AAG13538.1 putative gag-pol polyprotein
{Oryza sativa}, partial (1%)
Length = 638
Score = 82.4 bits (202), Expect = 1e-15
Identities = 51/144 (35%), Positives = 76/144 (52%)
Frame = +2
Query: 810 IRSSTAHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEK 869
+ SS+ + HY S HRA I + EP + +A + W AM+ EI+ LE+
Sbjct: 179 LNSSSKSYHIFHYVGYSFSAKHRASLAAITSNIEPKNYVQAAQ*QEWLAAMEQEIQVLEE 358
Query: 870 NGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVA 929
N T L L EG K + + VY++ +G + +YKA+LVAK + QVEG D+ D
Sbjct: 359 NNTSTLEPLREGKKWVDCRPVYKIIHKANGEIEKYKAQLVAKDFVQVEGEDF*D-LCLSN 535
Query: 930 KLTTVRVILALAASQNWHLHQLDV 953
K R +L +AA++ LH +DV
Sbjct: 536 KDDNCRCLLTIAAAKG*QLHLMDV 607
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 80.5 bits (197), Expect = 4e-15
Identities = 48/136 (35%), Positives = 72/136 (52%)
Frame = -2
Query: 606 AVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIF 665
A +L+NRIP+++LK+++P+E+L + L M+VFG LC+ R+KL+ R+RK +F
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTYMRVFGCLCYVLVPGELRNKLEARSRKAMF 525
Query: 666 LGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQAESTADKLN 725
+GY KGY D + + +SRDV F E Y+ N DL+ A L
Sbjct: 524 IGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEEKNQE-----DLRDLTSDKAGVLR 360
Query: 726 VASEIQSAKGDASVST 741
V E K + ST
Sbjct: 359 VILEGLGIKMNQDQST 312
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.144 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,419,576
Number of Sequences: 36976
Number of extensions: 571766
Number of successful extensions: 4371
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 2457
Number of HSP's successfully gapped in prelim test: 200
Number of HSP's that attempted gapping in prelim test: 1834
Number of HSP's gapped (non-prelim): 2783
length of query: 1346
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1238
effective length of database: 5,021,319
effective search space: 6216392922
effective search space used: 6216392922
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0366.1


