

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0359.2
(453 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
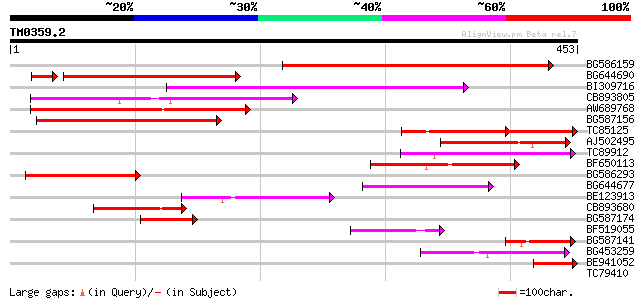
Score E
Sequences producing significant alignments: (bits) Value
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 212 2e-55
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 186 3e-51
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 161 6e-40
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 150 7e-37
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 149 3e-36
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 134 6e-32
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 75 1e-25
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 111 5e-25
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 103 1e-22
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 95 5e-20
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 93 2e-19
BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T... 74 2e-13
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 72 4e-13
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 68 6e-12
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 59 4e-09
BF519055 similar to GP|21592754|gb| unknown {Arabidopsis thalian... 48 9e-06
BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440... 44 1e-04
BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, ... 43 3e-04
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 42 5e-04
TC79410 similar to PIR|H84518|H84518 pathogenesis-related PR-1-l... 37 0.002
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 212 bits (540), Expect = 2e-55
Identities = 103/217 (47%), Positives = 146/217 (66%), Gaps = 1/217 (0%)
Frame = +1
Query: 219 IQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDTSGKVCQKLYRGM 278
++V Q EG YI Q KY +LL++F M +S +++ P+ P C L K++ KV Y+ +
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 279 IGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEYK 338
+G L+YL A+RPD+++ + L +RF + P E H+ AVKR+LRYL GTINLG+MYK+ K
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 339 LSGYCDADYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQM 398
L Y D+DYAGD +RKST G L S VSW+SK+Q + LST +AE+I+AA C+ Q
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 399 LWMKHQLEDYQILES-NIPIYCDNTAAISLSKNPILH 434
+WM+ LE +S +I +YCDN + I LSKNP+LH
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 186 bits (473), Expect(2) = 3e-51
Identities = 88/141 (62%), Positives = 114/141 (80%)
Frame = -2
Query: 44 VVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIILLISFSLNHNIILHQMDVKSAFLNG 103
+ RNK +LV QGY+Q+EGIDY E F+PVAR+EAI +LI+F+ L+QMDVKSAF+NG
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 104 YISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDT 163
+ +EV+V QPPGFED + P+HVF+L K+LYGLKQAPRAWYERLS FLL+N F RGK+D
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 164 TLFCKTYKDDILIVKIYVDDI 184
TLF + ++LI+++YVDDI
Sbjct: 64 TLFLLKRE*ELLIIQVYVDDI 2
Score = 33.5 bits (75), Expect(2) = 3e-51
Identities = 14/21 (66%), Positives = 16/21 (75%)
Frame = -3
Query: 18 LTKKPENVHVIGTKWVFRNKL 38
L +PE VIGT+WVFRNKL
Sbjct: 492 LVPRPEGKTVIGTRWVFRNKL 430
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 161 bits (407), Expect = 6e-40
Identities = 91/241 (37%), Positives = 134/241 (54%)
Frame = +2
Query: 126 VFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVKIYVDDII 185
V +L+KS+YGLKQA R WY +LS L+ +++ D +LF K + +YVDDI+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKFKDSSFTTLLVYVDDIV 199
Query: 186 FGSANQSLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNM 245
+ S + + F++ +G L+YFLG++V ++ +G ++Q KYT ELL+
Sbjct: 200 LAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDSGN 379
Query: 246 LESTVAKTPMHPTCILEKEDTSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSD 305
L TP + L D+ + YR +IG L+YLT +RPDI F+V ++F S
Sbjct: 380 LAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFVSK 559
Query: 306 PRETHLTAVKRILRYLKGTINLGLMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQFLG 365
P++ H A R+L+YLK GL Y TS KLS + D+D+A T RKS G FLG
Sbjct: 560 PQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVFLG 739
Query: 366 S 366
S
Sbjct: 740 S 742
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 150 bits (380), Expect = 7e-37
Identities = 84/220 (38%), Positives = 131/220 (59%), Gaps = 6/220 (2%)
Frame = +3
Query: 17 DLTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEA 76
+LT IG KW+F+ KLNE G++ + K RLVA+GYSQQ G+DYTE FAPVAR +
Sbjct: 132 ELTDLRSGAKTIGLKWIFKTKLNENGEIEKYKARLVAKGYSQQYGVDYTEVFAPVARWDT 311
Query: 77 IILLISFSLN---HNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVF--KLKK 131
I ++I+ + + + M + + + K + ++ V ++K+
Sbjct: 312 IRMVIALAAQIKRDGVCIS*M*KAHSCMEN*MRKFLLINH------RVM*RRVIS*RVKR 473
Query: 132 SLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD-ILIVKIYVDDIIFGSAN 190
+LYGLKQAPRAWY R+ ++ + F + + TLF K + ILI+ +YVDD+IF +
Sbjct: 474 ALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIGND 653
Query: 191 QSLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQTPEGTYI 230
+++ +EF K M+ EF MS +G++ YFLG++V Q +G YI
Sbjct: 654 ENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQNEKGIYI 773
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 149 bits (375), Expect = 3e-36
Identities = 78/176 (44%), Positives = 110/176 (62%)
Frame = +1
Query: 17 DLTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEA 76
DL P + IG KWV+R K N G V + K RLVA+G+SQ G DYTETF+PV +
Sbjct: 151 DLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVAKGFSQTLGCDYTETFSPVIKPVT 330
Query: 77 IILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGL 136
I L+++ ++ + + Q+D+ +AFLNG++ +EVY+ QP GFE K V KL KSLYGL
Sbjct: 331 IRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQPQGFEAANK-SLVCKLNKSLYGL 507
Query: 137 KQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQS 192
KQAPRAWYE L+S ++ F + + D +L + + IYVDDI+ ++ S
Sbjct: 508 KQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACIYLXIYVDDILITGSSAS 675
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 134 bits (338), Expect = 6e-32
Identities = 67/148 (45%), Positives = 96/148 (64%)
Frame = -1
Query: 22 PENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIILLI 81
P+ + ++W+F K G + R K RLVA+G++ G DY ETFAPVA+L I +++
Sbjct: 453 PKGKKAVSSRWIFTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVL 274
Query: 82 SFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPR 141
S ++N L QMDVK+AFL G + EVY++ PPG E K +V +LKK++YGLKQ+PR
Sbjct: 273 SLAVNLGWGLWQMDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPR 94
Query: 142 AWYERLSSFLLENEFVRGKVDTTLFCKT 169
AWY +LS+ L F + ++D TLF T
Sbjct: 93 AWYNKLSTTLNGRGFRKSELDHTLFTLT 10
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 75.5 bits (184), Expect(2) = 1e-25
Identities = 40/87 (45%), Positives = 56/87 (63%)
Frame = +1
Query: 314 VKRILRYLKGTINLGLMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQFLGSNLVSWAS 373
VKRI+RY+KGT + + + SE + GY D+D+AGD +RKST G L VSW S
Sbjct: 1 VKRIMRYIKGTSGVAVCFGG-SELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLS 177
Query: 374 KRQSTIALSTAEAEYISAAICSTQMLW 400
K Q+ +ALST EAEY++A + + L+
Sbjct: 178 KLQTVVALSTTEAEYMAAYLKHARKLF 258
Score = 58.9 bits (141), Expect(2) = 1e-25
Identities = 21/57 (36%), Positives = 39/57 (67%)
Frame = +3
Query: 397 QMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRGYVQK 453
+ +WM+ +E+ + I +YCD+ +A+ +++NP HSR KHI ++YHF+R V++
Sbjct: 249 EAIWMQRLMEELGHKQEQITVYCDSQSALHIARNPAFHSRTKHIGIQYHFVREVVEE 419
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 111 bits (278), Expect = 5e-25
Identities = 54/106 (50%), Positives = 75/106 (69%), Gaps = 2/106 (1%)
Frame = +2
Query: 345 ADYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQ 404
+D+AGD RKST G LG+ +SW+SK+Q +A STAEAEYI++ C+TQ +W++
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 405 LEDYQILESNIP--IYCDNTAAISLSKNPILHSRAKHIEVKYHFIR 448
LE E N P IYCDN +AI+LSKNP+ H R+KHI++++H IR
Sbjct: 182 LE-VMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIR 316
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 103 bits (257), Expect = 1e-22
Identities = 51/142 (35%), Positives = 83/142 (57%), Gaps = 2/142 (1%)
Frame = +1
Query: 313 AVKRILRYLKGTINLGLMYKKTSEYK--LSGYCDADYAGDRTERKSTFGNCQFLGSNLVS 370
A+K +L+YL ++ L Y K ++ + L GY DADYAG+ RKS G L +S
Sbjct: 4 ALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTIS 183
Query: 371 WASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKN 430
W + +QS + LST +AEYI+ +W+K + + I + + I+CD+ +AI L+ +
Sbjct: 184 WKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGITQEYVKIHCDSQSAIHLANH 363
Query: 431 PILHSRAKHIEVKYHFIRGYVQ 452
+ H R KHI+++ HFIR ++
Sbjct: 364 QVYHERTKHIDIRLHFIRDMIE 429
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 95.1 bits (235), Expect = 5e-20
Identities = 49/122 (40%), Positives = 74/122 (60%), Gaps = 3/122 (2%)
Frame = +1
Query: 289 RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMY---KKTSEYKLSGYCDA 345
RPDI +SV + ++F DPR+ HL A RILRY++GT+ GL++ K+ Y+L Y D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 346 DYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQL 405
D+ GD R+ST G +SW +K+Q ALS+ EAEYI+ + Q LW+ +
Sbjct: 301 DWCGD---RRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 406 ED 407
++
Sbjct: 472 KE 477
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 93.2 bits (230), Expect = 2e-19
Identities = 43/92 (46%), Positives = 63/92 (67%)
Frame = +2
Query: 13 RSKLDLTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVA 72
+ L L KKP V IG +W+++ K NE G +++ K RLVA+GY +Q+GID+ E FAPV
Sbjct: 50 KQTLKLVKKPTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVV 229
Query: 73 RLEAIILLISFSLNHNIILHQMDVKSAFLNGY 104
R+E I LL++ + + +H +DVK AFLNG+
Sbjct: 230 RIETI*LLLALAATNGC*IHHIDVKIAFLNGH 325
>BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T15F17.l
{Arabidopsis thaliana}, partial (3%)
Length = 539
Score = 73.6 bits (179), Expect = 2e-13
Identities = 40/104 (38%), Positives = 58/104 (55%)
Frame = -3
Query: 283 LYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEYKLSGY 342
+ LT P+I FS++L +R+ S P H +K I +YLKG I++GL Y K L GY
Sbjct: 531 ILLTLQGPNITFSINLLSRYSSAPTMRH*NGIKHICKYLKGIIDMGLFYSKDCSPDLIGY 352
Query: 343 CDADYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEA 386
+A Y D + +S G G+ ++SW S + STIA S+ A
Sbjct: 351 VNA*YLSDPHKARS*TGYIFTCGNTVISWRSTK*STIATSSNHA 220
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 72.0 bits (175), Expect = 4e-13
Identities = 42/125 (33%), Positives = 67/125 (53%), Gaps = 3/125 (2%)
Frame = +1
Query: 138 QAPRAWYERLSSFLLENEFVRGKVDTTLFCK---TYKDDILIVKIYVDDIIFGSANQSLC 194
Q+PR W++R + + + +++ + D +F K T K ILIV YVDDI +
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIV--YVDDIFLTGDHGK*I 174
Query: 195 KEFSKMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTP 254
K ++ EFE+ +G LKYFLG++V + +G+ I Q KY +LLK+ M+ + P
Sbjct: 175 KRLKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDP 354
Query: 255 MHPTC 259
C
Sbjct: 355 YGCNC 369
Score = 36.2 bits (82), Expect = 0.027
Identities = 20/57 (35%), Positives = 32/57 (56%)
Frame = +2
Query: 246 LESTVAKTPMHPTCILEKEDTSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARF 302
L+ ++TPM T L D V + Y+ ++G L+YL+ +RPDI F V ++F
Sbjct: 329 LDVKPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQF 499
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 68.2 bits (165), Expect = 6e-12
Identities = 35/74 (47%), Positives = 48/74 (64%)
Frame = -2
Query: 68 FAPVARLEAIILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVF 127
F P+ +L I+ L+S N+ L +DVK+AFL G + +++Y+HQP GF E V
Sbjct: 554 FVPIVKLNTIMFLLSIVAIENLYLE*LDVKTAFLRGDLVEDIYMHQPEGFS*E-VGKMVG 378
Query: 128 KLKKSLYGLKQAPR 141
KLKKS+YGLKQ PR
Sbjct: 377 KLKKSMYGLKQGPR 336
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse
transcriptase homolog - rape retrotransposon copia-like
(fragment), partial (84%)
Length = 249
Score = 58.9 bits (141), Expect = 4e-09
Identities = 25/46 (54%), Positives = 34/46 (73%)
Frame = -1
Query: 105 ISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSF 150
+ +++Y+ QP GF K DHV KL+KSLYGLKQ+PR WY+R S+
Sbjct: 246 LEEKIYMTQPEGFLFPGKEDHVCKLRKSLYGLKQSPRQWYKRFDSY 109
>BF519055 similar to GP|21592754|gb| unknown {Arabidopsis thaliana}, partial
(30%)
Length = 675
Score = 47.8 bits (112), Expect = 9e-06
Identities = 23/75 (30%), Positives = 45/75 (59%)
Frame = -3
Query: 273 KLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYK 332
KL ++G+LLY+T + PD+ FS++ ++F P + + +K+++R+ K TI K
Sbjct: 640 KLVHSIVGALLYITVTCPDLSFSINKPSQFMHKPTQINFQQLKKVMRHPKLTI------K 479
Query: 333 KTSEYKLSGYCDADY 347
K + ++ + DAD+
Sbjct: 478 KLFDLQIYAFSDADW 434
>BG587141 similar to PIR|H86461|H86 hypothetical protein AAF32440.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 731
Score = 43.9 bits (102), Expect = 1e-04
Identities = 21/58 (36%), Positives = 35/58 (60%), Gaps = 2/58 (3%)
Frame = +3
Query: 397 QMLWMKHQLED--YQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRGYVQ 452
Q +W++ L + ++ E + I DN + I+L++NP+ H R HI +YHFIR V+
Sbjct: 135 QAMWLQDLLSEVTWEPCEE-VVIRIDNQSVIALTRNPVFHGRGNHIHKRYHFIRECVE 305
>BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (5%)
Length = 657
Score = 42.7 bits (99), Expect = 3e-04
Identities = 33/122 (27%), Positives = 59/122 (48%), Gaps = 3/122 (2%)
Frame = -2
Query: 329 LMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIA--LSTAEA 386
+++K+ + + Y + AG +R ST G FLG N+V +Q+ +A +
Sbjct: 530 IVFKRE*KLSMEVYXNTVCAGWIVDRGSTSGY*MFLGGNMVE*---KQNVVAR*VQRHNF 360
Query: 387 EYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYH 445
E S + +K +L+D I + + ++ +N ++ NP+ H R KHIE+ H
Sbjct: 359 ELCSQGL*RVMDEELKIKLDDLIINYKDPMTLF*NNNFVSRIAHNPVQHYRTKHIEIDQH 180
Query: 446 FI 447
FI
Sbjct: 179 FI 174
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 42.0 bits (97), Expect = 5e-04
Identities = 17/35 (48%), Positives = 23/35 (65%)
Frame = +2
Query: 419 CDNTAAISLSKNPILHSRAKHIEVKYHFIRGYVQK 453
CD +A L+ NP+ HSR KHI + HF+R VQ+
Sbjct: 29 CDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQ 133
>TC79410 similar to PIR|H84518|H84518 pathogenesis-related PR-1-like protein
[imported] - Arabidopsis thaliana, partial (84%)
Length = 1002
Score = 36.6 bits (83), Expect(2) = 0.002
Identities = 18/54 (33%), Positives = 29/54 (53%)
Frame = +3
Query: 32 WVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIILLISFSL 85
+ F K N G + RLVA+G+ Q+ GIDY + F+ V + ++ FS+
Sbjct: 756 YFFCLKRNSDGSTLYYTTRLVAKGFHQRSGIDYKDQFSLVVKQNGVMHFYFFSM 917
Score = 22.3 bits (46), Expect(2) = 0.002
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +2
Query: 101 LNGYISKEVYVHQPPGF 117
LN ++EVY+ QPP F
Sbjct: 911 LNETFNEEVYMSQPPVF 961
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,822,826
Number of Sequences: 36976
Number of extensions: 188068
Number of successful extensions: 927
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 909
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 917
length of query: 453
length of database: 9,014,727
effective HSP length: 99
effective length of query: 354
effective length of database: 5,354,103
effective search space: 1895352462
effective search space used: 1895352462
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0359.2


