

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0349b.2
(400 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
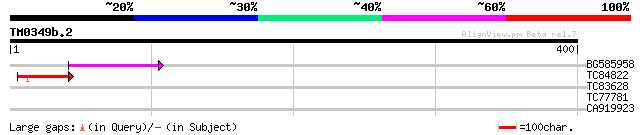
Sequences producing significant alignments: (bits) Value
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 66 3e-11
TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsi... 48 8e-06
TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_18... 38 0.008
TC77781 weakly similar to GP|23237921|dbj|BAC16494. contains EST... 31 0.97
CA919923 weakly similar to GP|23237921|dbj contains ESTs D41667(... 30 1.3
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 65.9 bits (159), Expect = 3e-11
Identities = 35/68 (51%), Positives = 41/68 (59%), Gaps = 1/68 (1%)
Frame = +2
Query: 42 SAPRG-RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDN 100
S PR R+++ R+ +RL +DYF P Y D FRRRYRM KHVFLRIV L D
Sbjct: 461 SRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVFLRIVEALGQHDE 640
Query: 101 YFTQRVDA 108
YF VDA
Sbjct: 641 YFQLTVDA 664
>TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsis
thaliana}, partial (47%)
Length = 636
Score = 47.8 bits (112), Expect = 8e-06
Identities = 25/42 (59%), Positives = 29/42 (68%), Gaps = 2/42 (4%)
Frame = +1
Query: 6 MDPPEF--DLEAYLEKSEREDTYILNRFRDRRNLILEDSAPR 45
MDP +F D+EAY KSE E+ YILNRFR+RR I E PR
Sbjct: 40 MDPSKFRFDIEAYKRKSEIEERYILNRFRERRKQIEEGYTPR 165
>TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_180
{Arabidopsis thaliana}, partial (23%)
Length = 773
Score = 37.7 bits (86), Expect = 0.008
Identities = 27/106 (25%), Positives = 45/106 (41%)
Frame = +2
Query: 68 NEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRML 127
N + D FRR +RM K F I +L SS + + ++ I + + L
Sbjct: 431 NREDFPDDEFRRCFRMSKQTFNMICNELDSS----VTKKNTTLRDAIPVRQRVAVCIYRL 598
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQE 173
A G V + +G +T + + C I + Q++LR P +E
Sbjct: 599 ATGEPLRLVSKKFGLGISTCHKLVLEVCAAIKSVLMQKFLRWPDEE 736
>TC77781 weakly similar to GP|23237921|dbj|BAC16494. contains ESTs
D41667(S4320) AU033165(S4320)~mucin-like protein,
partial (22%)
Length = 1150
Score = 30.8 bits (68), Expect = 0.97
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +3
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKGI 158
ANGVA ++Y+ + ET C+R + KGI
Sbjct: 741 ANGVALSKDEDYVVVCETWKFRCVRHWLKGI 833
>CA919923 weakly similar to GP|23237921|dbj contains ESTs D41667(S4320)
AU033165(S4320)~mucin-like protein, partial (20%)
Length = 774
Score = 30.4 bits (67), Expect = 1.3
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -2
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKGI 158
ANGVA ++Y+ + ET CL+ + KGI
Sbjct: 668 ANGVALSKDEDYLVVCETWKFRCLKHWLKGI 576
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,702,429
Number of Sequences: 36976
Number of extensions: 207226
Number of successful extensions: 1383
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1325
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1383
length of query: 400
length of database: 9,014,727
effective HSP length: 98
effective length of query: 302
effective length of database: 5,391,079
effective search space: 1628105858
effective search space used: 1628105858
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0349b.2


