

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
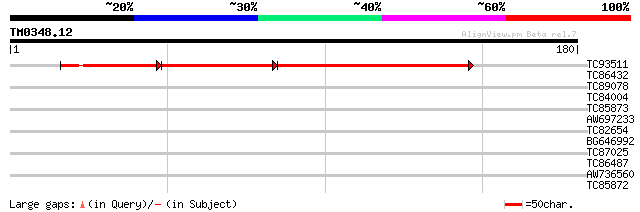
Query= TM0348.12
(180 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93511 similar to GP|21554137|gb|AAM63217.1 unknown {Arabidopsi... 74 3e-30
TC86432 similar to PIR|T05120|T05120 hypothetical protein F7H19.... 32 0.14
TC89078 similar to GP|14335172|gb|AAK59866.1 AT4g27390/M4I22_200... 29 0.88
TC84004 type IIB calcium ATPase [Medicago truncatula] 28 2.6
TC85873 homologue to SP|P52427|PSA4_SPIOL Proteasome subunit alp... 27 3.4
AW697233 similar to PIR|T46230|T462 NAC2-like protein - Arabidop... 27 4.4
TC82654 similar to PIR|T06603|T06603 hypothetical protein F16J13... 27 4.4
BG646992 similar to PIR|T06133|T061 hypothetical protein F23E12.... 27 5.7
TC87025 similar to PIR|S27757|S27757 embryonic abundant protein ... 27 5.7
TC86487 similar to PIR|C84473|C84473 probable protein kinase [im... 26 7.5
AW736560 similar to GP|20260382|gb| unknown protein {Arabidopsis... 26 7.5
TC85872 homologue to SP|P52427|PSA4_SPIOL Proteasome subunit alp... 26 9.8
>TC93511 similar to GP|21554137|gb|AAM63217.1 unknown {Arabidopsis
thaliana}, partial (24%)
Length = 650
Score = 74.3 bits (181), Expect(3) = 3e-30
Identities = 35/62 (56%), Positives = 43/62 (68%)
Frame = +1
Query: 86 AWALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVI 145
AW LF AFGS A +GP+ Y++ +AYQNAF GLS+GS L +VN I+F LGFVI
Sbjct: 463 AWTLFFAFGSSAMVGPMFYSISLAYQNAFNLGLSYGSQASELSPLFMVNTILFTALGFVI 642
Query: 146 GY 147
GY
Sbjct: 643 GY 648
Score = 65.9 bits (159), Expect(3) = 3e-30
Identities = 30/37 (81%), Positives = 33/37 (89%)
Frame = +2
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*Q 85
W GS+S V YPRCSIRRQSTYADAEGR+I LWFLH*+
Sbjct: 350 WYGSKSVVKYPRCSIRRQSTYADAEGRKICLWFLH*R 460
Score = 28.1 bits (61), Expect(3) = 3e-30
Identities = 19/32 (59%), Positives = 23/32 (71%)
Frame = +3
Query: 17 LKLFVCNRHEMLIVVVSV**LMTCLIEWS*SI 48
LKLF+ + M I S+**LMTCLIE S*S+
Sbjct: 255 LKLFIL-KQGMHITEESL**LMTCLIE*S*SL 347
>TC86432 similar to PIR|T05120|T05120 hypothetical protein F7H19.70 -
Arabidopsis thaliana, partial (72%)
Length = 1235
Score = 32.0 bits (71), Expect = 0.14
Identities = 35/128 (27%), Positives = 51/128 (39%)
Frame = +2
Query: 51 GSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAY 110
GS V PRCS+R + Y+D + ++ L+ A +A G L L+
Sbjct: 554 GSEIVVEGPRCSLRSKKVYSDLSVDYLRMFLLN--VPATVVALGLFFFLDDLT------- 706
Query: 111 QNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRG 170
G +++ P FS I + L + L A +K D V L+G
Sbjct: 707 ----GFEITYLLELPE-PFSFIFTWFAAVPLIVYLALSLTRAIIK--------DFVILKG 847
Query: 171 ACPNCGEE 178
CPNCG E
Sbjct: 848 PCPNCGTE 871
>TC89078 similar to GP|14335172|gb|AAK59866.1 AT4g27390/M4I22_200
{Arabidopsis thaliana}, partial (49%)
Length = 1211
Score = 29.3 bits (64), Expect = 0.88
Identities = 22/68 (32%), Positives = 33/68 (48%), Gaps = 2/68 (2%)
Frame = +2
Query: 113 AFGSGLSHGSHNPGLGFSI--IVNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRG 170
A +GL++ S LG+ I IV+ IF VL ++G + LW ++G
Sbjct: 467 ALAAGLTYLSMTGQLGWIIDAIVSIWIFAVLVPIVG---------IGAFLWWAGRDIMKG 619
Query: 171 ACPNCGEE 178
CPNCG +
Sbjct: 620 TCPNCGND 643
>TC84004 type IIB calcium ATPase [Medicago truncatula]
Length = 3655
Score = 27.7 bits (60), Expect = 2.6
Identities = 12/37 (32%), Positives = 21/37 (56%)
Frame = +1
Query: 135 NIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGA 171
N++ +++ FV SAP+ +Q LW N ++ GA
Sbjct: 2698 NVVALIINFVSACITGSAPLTAVQLLWVNLIMDTLGA 2808
>TC85873 homologue to SP|P52427|PSA4_SPIOL Proteasome subunit alpha type 4
(EC 3.4.25.1) (20S proteasome alpha subunit C), partial
(98%)
Length = 1042
Score = 27.3 bits (59), Expect = 3.4
Identities = 19/47 (40%), Positives = 28/47 (59%)
Frame = +3
Query: 10 PSNRTPRLKLFVCNRHEMLIVVVSV**LMTCLIEWS*SIWLGSRSDV 56
PS P+L+L C R +++ V+S L CL+ S*S LGS+ +V
Sbjct: 228 PSCFKPQLQLRKCTRLMIMLHVLS---LGLCLMPTS*STLLGSKHNV 359
>AW697233 similar to PIR|T46230|T462 NAC2-like protein - Arabidopsis
thaliana, partial (4%)
Length = 851
Score = 26.9 bits (58), Expect = 4.4
Identities = 16/50 (32%), Positives = 23/50 (46%), Gaps = 6/50 (12%)
Frame = +3
Query: 112 NAFGSGLSHGSHNPGLGF------SIIVNNIIFIVLGFVIGYPLASAPVK 155
N SG+ HG H PG+G S I N + L + GY A+ ++
Sbjct: 348 NRTDSGVRHGDHGPGVGGTVSSL*SFIHNR*TYTSLFYSDGYTYANLAIR 497
>TC82654 similar to PIR|T06603|T06603 hypothetical protein F16J13.30 -
Arabidopsis thaliana, partial (75%)
Length = 1207
Score = 26.9 bits (58), Expect = 4.4
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 164 DLVALRGACPNCGEE 178
D + L+G CPNCG E
Sbjct: 806 DFLILKGPCPNCGTE 850
>BG646992 similar to PIR|T06133|T061 hypothetical protein F23E12.200 -
Arabidopsis thaliana, partial (11%)
Length = 761
Score = 26.6 bits (57), Expect = 5.7
Identities = 15/50 (30%), Positives = 25/50 (50%), Gaps = 9/50 (18%)
Frame = -3
Query: 126 GLGFSIIVNNIIFIVLGFVIGYPL-----ASAPVKVIQGL----WRNDLV 166
GLG +II+NN +F + + + L + +I G+ WRN L+
Sbjct: 324 GLGLTIIINNFLFYNMNLIFNFRLIFFIFKMTMIMMIIGMIRIRWRNKLL 175
>TC87025 similar to PIR|S27757|S27757 embryonic abundant protein 59K -
soybean, partial (56%)
Length = 1353
Score = 26.6 bits (57), Expect = 5.7
Identities = 10/31 (32%), Positives = 18/31 (57%)
Frame = -3
Query: 118 LSHGSHNPGLGFSIIVNNIIFIVLGFVIGYP 148
L+H H F + +I+F++ FV+G+P
Sbjct: 598 LTHLIHGSIFSFLCLFRSIVFVLPSFVLGFP 506
>TC86487 similar to PIR|C84473|C84473 probable protein kinase [imported] -
Arabidopsis thaliana, partial (75%)
Length = 2229
Score = 26.2 bits (56), Expect = 7.5
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 1/70 (1%)
Frame = -2
Query: 65 RQSTYADAEGRRIYLWFLH*QA-WALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSH 123
R T D G + LW H* A A FLA S C L + + + +F +GL+H +
Sbjct: 1484 RVLTTVDIVGLFLGLWLKH**ANAAAFLAPISEYCPSSLGSIILESLRASFNTGLAHSTR 1305
Query: 124 NPGLGFSIIV 133
LG +V
Sbjct: 1304 FCSLGGRTLV 1275
>AW736560 similar to GP|20260382|gb| unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 200
Score = 26.2 bits (56), Expect = 7.5
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = +3
Query: 112 NAFGSGLSHGSHNPGLG 128
N++GS SHGS+N G+G
Sbjct: 108 NSYGSVGSHGSYNDGIG 158
>TC85872 homologue to SP|P52427|PSA4_SPIOL Proteasome subunit alpha type 4
(EC 3.4.25.1) (20S proteasome alpha subunit C), complete
Length = 1139
Score = 25.8 bits (55), Expect = 9.8
Identities = 19/47 (40%), Positives = 27/47 (57%)
Frame = +3
Query: 10 PSNRTPRLKLFVCNRHEMLIVVVSV**LMTCLIEWS*SIWLGSRSDV 56
PS P+L+L C R L++++ V L C + S*SI LGS+ V
Sbjct: 285 PSFCKPQLQLRKCTR---LMIMLHVLLLGLCPMPTS*SILLGSKHSV 416
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,584,343
Number of Sequences: 36976
Number of extensions: 98912
Number of successful extensions: 588
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 582
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 587
length of query: 180
length of database: 9,014,727
effective HSP length: 90
effective length of query: 90
effective length of database: 5,686,887
effective search space: 511819830
effective search space used: 511819830
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0348.12


