

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0342b.6
(427 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
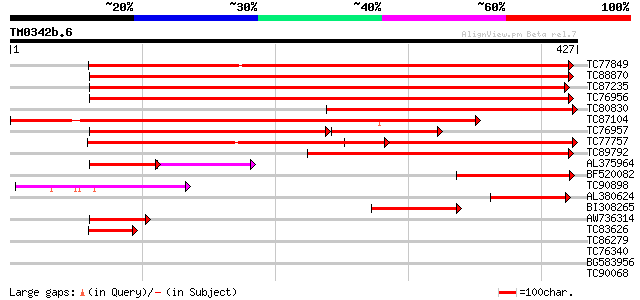
Score E
Sequences producing significant alignments: (bits) Value
TC77849 similar to PIR|A96610|A96610 probable pectinacetylestera... 440 e-124
TC88870 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3... 411 e-115
TC87235 weakly similar to PIR|S68805|S68805 pectin acetylesteras... 402 e-112
TC76956 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3... 394 e-110
TC80830 weakly similar to GP|6478931|gb|AAF14036.1| putative pec... 345 1e-95
TC87104 weakly similar to GP|10176854|dbj|BAB10060. pectinacetyl... 319 1e-87
TC76957 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3... 208 3e-75
TC77757 weakly similar to PIR|A96610|A96610 probable pectinacety... 258 4e-69
TC89792 weakly similar to GP|16226325|gb|AAL16135.1 AT3g05910/F2... 203 1e-52
AL375964 similar to GP|10176999|dbj pectin acetylesterase {Arabi... 75 1e-25
BF520082 similar to PIR|T01197|T011 pectin acetylesterase homolo... 112 4e-25
TC90898 similar to PIR|A96610|A96610 probable pectinacetylestera... 100 1e-21
AL380624 similar to PIR|T02194|T021 probable pectinacetylesteras... 81 9e-16
BI308265 similar to PIR|S68805|S688 pectin acetylesterase (EC 3.... 80 2e-15
AW736314 weakly similar to PIR|S68805|S688 pectin acetylesterase... 50 2e-06
TC83626 similar to PIR|T01197|T01197 pectin acetylesterase homol... 47 2e-05
TC86279 homologue to PIR|T07093|T07093 acetyl-CoA carboxylase (E... 32 0.36
TC76340 homologue to SP|Q9SM09|VATA_CITUN Vacuolar ATP synthase ... 32 0.36
BG583956 homologue to GP|10177159|dbj gb|AAF63824.1~gene_id:K21P... 32 0.36
TC90068 similar to GP|20197988|gb|AAD22321.2 expressed protein {... 32 0.61
>TC77849 similar to PIR|A96610|A96610 probable pectinacetylesterase
precursor T8L23.6 [imported] - Arabidopsis thaliana,
partial (86%)
Length = 1589
Score = 440 bits (1131), Expect = e-124
Identities = 195/365 (53%), Positives = 262/365 (71%)
Frame = +3
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
+IP T + A+ +GA+CLDG+ PG+HF G GSG+ SWL+ LEGGGWCN+I SC RK T
Sbjct: 246 MIPLTLIHGAVSKGAVCLDGTLPGYHFHPGSGSGANSWLIQLEGGGWCNTIRSCVFRKTT 425
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSSKYME +PF+GILS+ QNPDFFNWN+V +RYCDGASF+G ++E L FRG
Sbjct: 426 RRGSSKYMEKQLPFTGILSNKAEQNPDFFNWNRVKVRYCDGASFSGDSQNEAAQ-LQFRG 602
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
Q IW A M+EL+S GM A QALLSGCSAGGLA+++HCD+F+ + PK T VKCL+DAGFF
Sbjct: 603 QKIWLAAMEELMSRGMKNANQALLSGCSAGGLASILHCDEFQSLFPKSTKVKCLSDAGFF 782
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
LD D+ G +T+R+ + VV LQ V K+L K CL ++P+ C FP ++ +++TP+FL++
Sbjct: 783 LDATDVSGGHTLRNLFGGVVNLQEVQKNLPKSCLNHLDPTSCFFPQNLIDHVQTPLFLLN 962
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQ + L P +DP G W +CK N NCN++ I IL FR+ +L + F Q
Sbjct: 963 AAYDAWQFQESLAPHSADPHGSWNNCKSNHANCNSSQIQILQNFRNQMLNDIKGFSTTSQ 1142
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
G+F+NSCF HCQ+ R +TW++ +SP + N IA ++ +WFFDR+V+ IDC YPC+ TC
Sbjct: 1143SGLFINSCFAHCQSERQDTWFADDSPLLNNMPIAVAIGNWFFDRQVVKAIDCAYPCDNTC 1322
Query: 420 RNLDF 424
NL F
Sbjct: 1323HNLVF 1337
>TC88870 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (93%)
Length = 1409
Score = 411 bits (1056), Expect = e-115
Identities = 177/365 (48%), Positives = 259/365 (70%), Gaps = 1/365 (0%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+P T L +A+ +GA+CLDGS P +H KGFG+G SWL+ EGGGWCN++++C RK
Sbjct: 315 VPITLLKSAVSKGAVCLDGSPPAYHLDKGFGTGINSWLVQFEGGGWCNNVTTCLSRKTYR 494
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKG-SGLFFRG 179
LGSSK M + FSGIL++ R NPDF+NWN++ +RYCDG+SF G E+ + L FRG
Sbjct: 495 LGSSKQMANQIAFSGILNNRRQFNPDFYNWNRIKVRYCDGSSFTGDVEAVNPVTKLHFRG 674
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
I+ A+M++LL+ GM AK A++SGCSAGGLA+++HCD FR +LP+ VKCL++AG+F
Sbjct: 675 ARIFNAVMEDLLAKGMKNAKNAIISGCSAGGLASILHCDRFRALLPRGAKVKCLSNAGYF 854
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
++ +D+ G + + ++ VV G AK+L + C +++ P C FP ++ I TP+FLV+
Sbjct: 855 INARDVSGAHHVEQYFTQVVTTHGSAKNLPRSCTSRLSPRLCFFPQYVISQIATPIFLVN 1034
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+NIL P +DP GHW SCKL+I+NC++N +D++ FR+ L+AL
Sbjct: 1035AAYDSWQIKNILAPGAADPRGHWHSCKLDINNCSSNQLDLMQGFRTQFLRALTAVGNSTS 1214
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
GMF++SC+ HCQT TW+S +SP + +IA++VADWF++R++ + IDC YPCNP+C
Sbjct: 1215KGMFIDSCYAHCQTEMQGTWFSSDSPSLAKTSIAKAVADWFYERKLFHKIDCSYPCNPSC 1394
Query: 420 RNLDF 424
+N F
Sbjct: 1395KNRVF 1409
>TC87235 weakly similar to PIR|S68805|S68805 pectin acetylesterase (EC
3.1.1.-) precursor - mung bean, partial (89%)
Length = 1447
Score = 402 bits (1033), Expect = e-112
Identities = 171/362 (47%), Positives = 253/362 (69%), Gaps = 1/362 (0%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+P T L +A+ +GA+CLDGS P +HF +G G+ +W++H+EGGGWC++++ C R+ +
Sbjct: 123 VPLTRLESAVSKGAVCLDGSPPAYHFDQGHDEGANNWIVHIEGGGWCHNVTYCLYRRDSR 302
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
LGSS ME FSG LS ++ NPDF+NWN+V +RYCDG+SF G E + + L++RG
Sbjct: 303 LGSSHEMEEQTYFSGYLSDNQQYNPDFYNWNRVKVRYCDGSSFTGDVEEVDPTTKLYYRG 482
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
I+ A+M+ELL+ GM A+ A+LSGCSAGGL T++HCD FR + P ET VKC++DAG+F
Sbjct: 483 ARIFSAVMEELLAKGMDHAENAILSGCSAGGLTTILHCDGFRALFPNETRVKCVSDAGYF 662
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
++ DI G++ + +Y+ VV G KSL C + + P C FP + +I+TP+F+V+
Sbjct: 663 VNVNDISGDHYIEDYYSQVVATHGSEKSLPSSCTSMLSPGLCFFPQYMASSIQTPIFIVN 842
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+NIL P +DP G W+SCK N++NC+ ++I+ +R+ L+AL+
Sbjct: 843 AAYDSWQIKNILAPGDADPDGQWRSCKTNLNNCSPEQLNIMQDYRTQFLEALSPISNSPS 1022
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
GMF++SC++HCQT ETW+ +SP + NKT+A++V DWF++R IDC YPCNPTC
Sbjct: 1023NGMFIDSCYVHCQTEPQETWFKSDSPMVGNKTVAKAVGDWFYERSPSREIDCTYPCNPTC 1202
Query: 420 RN 421
+N
Sbjct: 1203QN 1208
>TC76956 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (93%)
Length = 1573
Score = 394 bits (1011), Expect = e-110
Identities = 172/365 (47%), Positives = 247/365 (67%), Gaps = 1/365 (0%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+P T + +A+ +GA+CLDGS P +HF KGF +G +W++H EGGGWCN+ ++C R T
Sbjct: 204 VPITFVQSAVAKGAVCLDGSPPAYHFDKGFEAGIDNWIVHFEGGGWCNNATTCLDRIDTR 383
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
LGSSK M+ + FSG SS + NPDF+NWN++ +RYCDG+SF G E+ + + L +RG
Sbjct: 384 LGSSKKMDKTLSFSGFFSSGKKFNPDFYNWNRIKVRYCDGSSFTGDVEAVDPKTNLHYRG 563
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
I+ A++++LL+ GM AK A+LSGCSAGGL +++ CD FR +LP VKC++DAG+F
Sbjct: 564 GRIFVAVIEDLLAKGMKNAKNAILSGCSAGGLTSILQCDRFRTLLPAAAKVKCVSDAGYF 743
Query: 240 LDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVH 299
++ K + G + + FY+ VVQ G AK+L C +++ P C FP + IKTP+F V+
Sbjct: 744 INVKAVSGASHIEQFYSQVVQTHGSAKNLPSSCTSRLSPGLCFFPQNVAAQIKTPIFFVN 923
Query: 300 SAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQ 359
+AYD WQI+NIL P +DP G W++CKL+I +C+AN + + FR+ LKA++
Sbjct: 924 AAYDSWQIKNILAPGVADPHGTWRNCKLDIKSCSANQLSTMQGFRTEFLKAISVVSNSPS 1103
Query: 360 IGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTC 419
GMF++ C+ HCQT ETW +SP + TIA++V DW++DR IDCPYPCNPTC
Sbjct: 1104KGMFIDGCYSHCQTGMQETWMRTDSPVLAKTTIAKAVGDWYYDRSTFQQIDCPYPCNPTC 1283
Query: 420 RNLDF 424
N F
Sbjct: 1284HNRVF 1298
>TC80830 weakly similar to GP|6478931|gb|AAF14036.1| putative
pectinacetylesterase {Arabidopsis thaliana}, partial
(43%)
Length = 1045
Score = 345 bits (886), Expect = 1e-95
Identities = 157/189 (83%), Positives = 174/189 (91%)
Frame = +1
Query: 239 FLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLV 298
FLDEKDI GN+TM+SFY+DVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLV
Sbjct: 7 FLDEKDIAGNSTMKSFYHDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLV 186
Query: 299 HSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRK 358
H AYDFWQI NILVPEGSDP WKSC+LNI +C+AN+I IL+ FRSSLLKA+NEFQQRK
Sbjct: 187 HPAYDFWQIHNILVPEGSDPHRRWKSCRLNIQSCDANMISILDSFRSSLLKAVNEFQQRK 366
Query: 359 QIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPT 418
IGMF++SCFIHCQTW GETW+SP SPKI +KTIAESVADWFFDR+V+ LIDCP+PCNPT
Sbjct: 367 DIGMFIDSCFIHCQTWMGETWHSPRSPKINHKTIAESVADWFFDRQVVKLIDCPFPCNPT 546
Query: 419 CRNLDFTRV 427
C N+DFTRV
Sbjct: 547 CHNMDFTRV 573
>TC87104 weakly similar to GP|10176854|dbj|BAB10060. pectinacetylesterase
{Arabidopsis thaliana}, partial (73%)
Length = 1499
Score = 319 bits (817), Expect = 1e-87
Identities = 164/358 (45%), Positives = 220/358 (60%), Gaps = 4/358 (1%)
Frame = +2
Query: 1 MSSSPSFSITNLRLRTLVVFFRKLTTKDYAIAAFAFFLLLSLAFFSQLHHHHHSHSLSNL 60
+SS SF + L + + AFA F L+++ + HS L
Sbjct: 401 LSSQISFVLRELESSGKTMIHNSFSVTMNFAVAFALFYLITVESWRV-----HSQEPKKL 565
Query: 61 -IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
+ T + NA GA CLDGS P +H +GFG+G +WLL EGGGWCN + SC R T
Sbjct: 566 YVNMTLVNNARETGAFCLDGSLPAYHLDRGFGAGEDNWLLQFEGGGWCNDLKSCLERAKT 745
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKG-SGLFFR 178
GS+ YM F+GILS++ + NPDF+NWN+V +RYCDGASF G+ G + L+F+
Sbjct: 746 RRGSTNYMTKYETFNGILSNNATVNPDFYNWNRVKLRYCDGASFTGNKVFNNGTTKLYFK 925
Query: 179 GQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGF 238
GQ IWEA++ +LL G+ KA++ALLSGCSAGGLAT HCD+F + LP +VKCL+DAGF
Sbjct: 926 GQKIWEALIADLLPKGLGKARKALLSGCSAGGLATFHHCDNFTKYLPTNASVKCLSDAGF 1105
Query: 239 FLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKME--PSKCLFPSEILKNIKTPVF 296
FLD +D+ N+TMR F+ VV+LQG ++L+K C + M P C FP +LK I TP F
Sbjct: 1106FLDGRDVSLNHTMRYFFKSVVRLQGSVQNLNKNCTSAMPSYPDLCFFPQYVLKYISTPYF 1285
Query: 297 LVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEF 354
+++SAYD +Q NILVP +DP GHW CK + C I+ L FR S++ AL F
Sbjct: 1286ILNSAYDVFQFHNILVPPSTDPRGHWIHCKKDPAACTPTEINTLQGFRLSMIAALKPF 1459
>TC76957 similar to PIR|S68805|S68805 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (68%)
Length = 858
Score = 208 bits (530), Expect(2) = 3e-75
Identities = 91/182 (50%), Positives = 133/182 (73%), Gaps = 1/182 (0%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTA 120
+PFT + +A+ +GA+CLDGS P +HF KGF +G +W++H EGGGWCN+ ++C R T
Sbjct: 24 VPFTFVQSAVAKGAVCLDGSPPAYHFDKGFEAGIDNWIVHFEGGGWCNNATTCLDRIDTR 203
Query: 121 LGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRG 179
LGSSK M+ + FSG SS + NPDF+NWN++ +RYCDG+SF G E+ + + L +RG
Sbjct: 204 LGSSKKMDKTLSFSGFFSSGKKFNPDFYNWNRIKVRYCDGSSFTGDVEAVDPKTNLHYRG 383
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFF 239
I+ A++++LL+ GM AK A+LSGCSAGGL +++ CD FR +LP VKC++DAG+F
Sbjct: 384 GRIFVAVIEDLLAKGMKNAKNAILSGCSAGGLTSILQCDRFRTLLPAAAKVKCVSDAGYF 563
Query: 240 LD 241
++
Sbjct: 564 IN 569
Score = 91.7 bits (226), Expect(2) = 3e-75
Identities = 38/84 (45%), Positives = 54/84 (64%)
Frame = +2
Query: 243 KDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAY 302
K + G + + FY+ VVQ G AK+L C +++ P C FP + IKTP+F V++AY
Sbjct: 605 KAVSGASHIEQFYSQVVQTHGSAKNLPSSCTSRLSPGLCFFPQNVAAQIKTPIFFVNAAY 784
Query: 303 DFWQIRNILVPEGSDPSGHWKSCK 326
D WQI+NIL P +DP G W++CK
Sbjct: 785 DSWQIKNILAPGVADPHGTWRNCK 856
>TC77757 weakly similar to PIR|A96610|A96610 probable pectinacetylesterase
precursor T8L23.6 [imported] - Arabidopsis thaliana,
partial (86%)
Length = 1536
Score = 258 bits (658), Expect = 4e-69
Identities = 122/228 (53%), Positives = 162/228 (70%)
Frame = +2
Query: 59 NLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKF 118
+++ T + A +GA+CLDG+ PG+H +GF SG+ SWL+HLEGGGWC+++ +C RK
Sbjct: 332 HMVDLTLIQGADSKGAVCLDGTLPGYHLDRGFESGANSWLIHLEGGGWCDTVRNCVYRKK 511
Query: 119 TALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFR 178
T GSSKYME +PF+GILS+ +NPDFFNWN+V +RYCDGASFAG E+E +GL FR
Sbjct: 512 THRGSSKYMENQIPFTGILSNKPEENPDFFNWNRVKLRYCDGASFAGDSENE-AAGLQFR 688
Query: 179 GQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGF 238
GQ IW A ++ELLS GM A+QALLSGCSAGGLA+++HCD+F+ +LPK + VKC +DAGF
Sbjct: 689 GQKIWLAAIEELLSQGMQNAEQALLSGCSAGGLASIIHCDEFQSLLPKSSKVKCFSDAGF 868
Query: 239 FLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSE 286
FLD DI G T+R + L G + K+ + P+ L SE
Sbjct: 869 FLDAIDISGGRTLREYVWRCSSLTGSPEKSAKKLSQQNGPNFVLLSSE 1012
Score = 201 bits (512), Expect = 3e-52
Identities = 86/175 (49%), Positives = 122/175 (69%)
Frame = +3
Query: 253 SFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILV 312
+ + VV LQ V K+L K CL KM+P+ C FP ++++++TP+FL+++AYD WQIR L
Sbjct: 912 NMFGGVVHLQEVQKNLPKSCLNKMDPTSCFFPQNVVEHVETPLFLLNAAYDVWQIRASLA 1091
Query: 313 PEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQ 372
P +DP G W CK N NCN++ I +L FR+ +L + +F + Q G+F+NSCF HCQ
Sbjct: 1092 PSTADPLGSWNDCKSNNANCNSSQIQLLQDFRNQMLDDVKDFSRSSQTGLFINSCFAHCQ 1271
Query: 373 TWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDFTRV 427
+ R ETW++ +SP I +K IA +V DW+FDREV+ IDCPYPC+ +C NL F RV
Sbjct: 1272 SERQETWFADDSPLIDDKPIAVAVGDWYFDREVVKAIDCPYPCDNSCHNLVFRRV 1436
>TC89792 weakly similar to GP|16226325|gb|AAL16135.1 AT3g05910/F2O10_3
{Arabidopsis thaliana}, partial (45%)
Length = 1012
Score = 203 bits (516), Expect = 1e-52
Identities = 88/200 (44%), Positives = 127/200 (63%)
Frame = +3
Query: 225 PKETTVKCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFP 284
P+ T VKC +DAG FLD D+ G ++R+ + VV LQG KSL + C + P C FP
Sbjct: 9 PRTTRVKCFSDAGLFLDSVDVSGRRSLRNLFGSVVTLQGAHKSLPRSCTNHLNPILCFFP 188
Query: 285 SEILKNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFR 344
++ +++TP+FL+++AYD WQI+ L P +D +W C+ N C++ I L FR
Sbjct: 189 QHLIASVRTPLFLLNAAYDTWQIQASLAPPSADYHWNWYDCRKNYARCSSPQIQYLQGFR 368
Query: 345 SSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDRE 404
+ +L+ F + +Q G+F+NSCF HCQ+ R +TW++ SP I NK IA+SV +WFFDR
Sbjct: 369 NQMLRVTRRFSRSRQNGLFINSCFAHCQSERQDTWHARGSPHIGNKGIADSVGNWFFDRV 548
Query: 405 VINLIDCPYPCNPTCRNLDF 424
+ I CPYPC+ TC NL F
Sbjct: 549 GVQAIGCPYPCDKTCHNLVF 608
>AL375964 similar to GP|10176999|dbj pectin acetylesterase {Arabidopsis
thaliana}, partial (24%)
Length = 515
Score = 74.7 bits (182), Expect(2) = 1e-25
Identities = 25/53 (47%), Positives = 41/53 (77%)
Frame = +3
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSC 113
+P T L +A+ +GA+CLDGS P +HF +G G+ +W++H+EGGGWC++++ C
Sbjct: 108 VPLTRLESAVSKGAVCLDGSPPAYHFDQGHDEGANNWIVHIEGGGWCHNVTYC 266
Score = 60.1 bits (144), Expect(2) = 1e-25
Identities = 29/75 (38%), Positives = 43/75 (56%), Gaps = 1/75 (1%)
Frame = +1
Query: 112 SCSRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-E 170
S S L + + + SG LS ++ NPDF+NWN+V +RYCDG+SF G E +
Sbjct: 289 SXSXXSXXXLAHTGHEVXXLTLSGYLSDNQQYNPDFYNWNRVKVRYCDGSSFTGDAEEVD 468
Query: 171 KGSGLFFRGQIIWEA 185
+ L++RG I+ A
Sbjct: 469 PTTKLYYRGARIFSA 513
>BF520082 similar to PIR|T01197|T011 pectin acetylesterase homolog F21E10.11
- Arabidopsis thaliana, partial (20%)
Length = 522
Score = 112 bits (279), Expect = 4e-25
Identities = 46/89 (51%), Positives = 65/89 (72%)
Frame = -3
Query: 337 IDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESV 396
I L FR+ +L ++ +F + + G+F+NSCF HCQT R +TW+S NSP I+NK IA +V
Sbjct: 520 IKFLQGFRTHMLNSIKDFSRSNKNGLFINSCFAHCQTERQDTWFSDNSPVIRNKVIALAV 341
Query: 397 ADWFFDREVINLIDCPYPCNPTCRNLDFT 425
DW+FDRE + +IDCPYPC+ TC +L F+
Sbjct: 340 GDWYFDREGVKVIDCPYPCDKTCHHLVFS 254
>TC90898 similar to PIR|A96610|A96610 probable pectinacetylesterase
precursor T8L23.6 [imported] - Arabidopsis thaliana,
partial (18%)
Length = 579
Score = 100 bits (249), Expect = 1e-21
Identities = 58/159 (36%), Positives = 86/159 (53%), Gaps = 27/159 (16%)
Frame = +1
Query: 5 PSFSITNLRLRTLVV-FFRKLTTKDYA-----IAAFAFFLLLSLAFFSQL----HHH--- 51
P F N + T V FF K KD + F ++++L F + + ++H
Sbjct: 103 PFFPTINQKFCTFVFCFFHKTKHKDLEETITMVNLFWVCIVIALVFTNWVDANAYYHINE 282
Query: 52 --------HHSHSLSNLIP------FTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSW 97
H + S S+L+ T + +A +GA+CLDG+ P +HF G+GSG+ SW
Sbjct: 283 TELSILEAHEASSFSSLVAQPHMVGITLIQSAAAKGAVCLDGTLPAYHFDHGYGSGANSW 462
Query: 98 LLHLEGGGWCNSISSCSRRKFTALGSSKYMETVVPFSGI 136
L++LEGGGWCN+ +C RK T GSSK+ME +PF+GI
Sbjct: 463 LVNLEGGGWCNNRXTCVYRKTTRRGSSKFMEKAIPFTGI 579
>AL380624 similar to PIR|T02194|T021 probable pectinacetylesterase At2g46930
- Arabidopsis thaliana, partial (14%)
Length = 460
Score = 80.9 bits (198), Expect = 9e-16
Identities = 34/60 (56%), Positives = 42/60 (69%)
Frame = +2
Query: 363 FVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNL 422
F+NSCF HCQ+ +TW +SP+I N TIAE+V DW+F R ID PYPC+ TCRNL
Sbjct: 2 FINSCFAHCQSESQDTWSGADSPRIINTTIAEAVGDWYFCRNKSKAIDWPYPCDTTCRNL 181
>BI308265 similar to PIR|S68805|S688 pectin acetylesterase (EC 3.1.1.-)
precursor - mung bean, partial (21%)
Length = 678
Score = 79.7 bits (195), Expect = 2e-15
Identities = 32/68 (47%), Positives = 46/68 (67%)
Frame = +3
Query: 273 LAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNC 332
L +P C FP + IKTP+F V++AYD WQI+NIL P +DP G W++CKL+I +C
Sbjct: 378 LQDSQPGLCFFPQNVAAQIKTPIFFVNAAYDSWQIKNILAPGVADPHGTWRNCKLDIKSC 557
Query: 333 NANLIDIL 340
+AN + +
Sbjct: 558 SANQLSTM 581
>AW736314 weakly similar to PIR|S68805|S688 pectin acetylesterase (EC
3.1.1.-) precursor - mung bean, partial (13%)
Length = 203
Score = 49.7 bits (117), Expect = 2e-06
Identities = 20/46 (43%), Positives = 30/46 (64%)
Frame = +2
Query: 61 IPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGW 106
+P T +A GA+ LDG P +H +GF +G+ +W++H EGGGW
Sbjct: 65 VPITFDQSASA*GAV*LDGRPPAYHCDQGFQAGNDNWIVHFEGGGW 202
>TC83626 similar to PIR|T01197|T01197 pectin acetylesterase homolog
F21E10.11 - Arabidopsis thaliana, partial (9%)
Length = 562
Score = 46.6 bits (109), Expect = 2e-05
Identities = 18/37 (48%), Positives = 26/37 (69%)
Frame = +2
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRS 96
++ T + A +GA+CLDGS P +HF +G+GSGS S
Sbjct: 428 MVGLTLINGAAAKGAVCLDGSLPAYHFHRGYGSGSNS 538
>TC86279 homologue to PIR|T07093|T07093 acetyl-CoA carboxylase (EC 6.4.1.2)
biotin carboxylase chain precursor - soybean, partial
(86%)
Length = 2189
Score = 32.3 bits (72), Expect = 0.36
Identities = 19/83 (22%), Positives = 37/83 (43%)
Frame = -1
Query: 324 SCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPN 383
+C H+ NL+ + F S L K L + R N CF+HC + ++PN
Sbjct: 854 TCGCLDHHRETNLVCQSDSFFSGLQKPLTSWNSR-------NICFLHCFSCSCFVTHNPN 696
Query: 384 SPKIQNKTIAESVADWFFDREVI 406
+ ++++ I ++ F ++
Sbjct: 695 TVRVRSNKIDSMISTHFNKNSIL 627
>TC76340 homologue to SP|Q9SM09|VATA_CITUN Vacuolar ATP synthase catalytic
subunit A (EC 3.6.3.14) (V-ATPase A subunit), complete
Length = 2417
Score = 32.3 bits (72), Expect = 0.36
Identities = 15/34 (44%), Positives = 20/34 (58%)
Frame = +3
Query: 29 YAIAAFAFFLLLSLAFFSQLHHHHHSHSLSNLIP 62
+ I F FF FF+ +HHHHH S+SN +P
Sbjct: 18 FLIQIFFFFFF----FFNFIHHHHHL*SVSNKMP 107
>BG583956 homologue to GP|10177159|dbj
gb|AAF63824.1~gene_id:K21P3.18~similar to unknown
protein {Arabidopsis thaliana}, partial (43%)
Length = 818
Score = 32.3 bits (72), Expect = 0.36
Identities = 16/28 (57%), Positives = 19/28 (67%)
Frame = +1
Query: 34 FAFFLLLSLAFFSQLHHHHHSHSLSNLI 61
F+F L SL+ S LHHHHH HSL L+
Sbjct: 19 FSFSLYNSLSTLS-LHHHHHHHSLDLLL 99
>TC90068 similar to GP|20197988|gb|AAD22321.2 expressed protein {Arabidopsis
thaliana}, partial (88%)
Length = 1352
Score = 31.6 bits (70), Expect = 0.61
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +2
Query: 19 VFFRKLTTKDYAIAAFAFFLLLSLAFFSQLHHHHHSHSLSN 59
VF K T + + +LLL L F Q+HH HH ++L++
Sbjct: 503 VFMFKATHQPFIQQHQLIYLLLPLLMFCQVHHPHHLNTLTS 625
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.138 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,206,869
Number of Sequences: 36976
Number of extensions: 323752
Number of successful extensions: 5041
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 3799
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4454
length of query: 427
length of database: 9,014,727
effective HSP length: 99
effective length of query: 328
effective length of database: 5,354,103
effective search space: 1756145784
effective search space used: 1756145784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0342b.6


