

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.9
(434 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
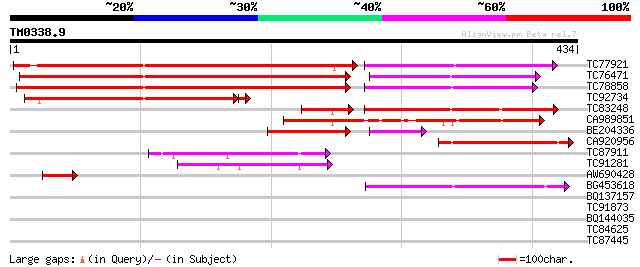
Score E
Sequences producing significant alignments: (bits) Value
TC77921 similar to GP|21553837|gb|AAM62930.1 phytochelatin synth... 411 e-136
TC76471 similar to PIR|T51392|T51392 probable phytochelatin synt... 376 e-115
TC78858 similar to PIR|T51392|T51392 probable phytochelatin synt... 370 e-115
TC92734 similar to GP|21553837|gb|AAM62930.1 phytochelatin synth... 266 1e-72
TC83248 similar to GP|13477083|dbj|BAB02996. phytochelatin synth... 143 9e-48
CA989851 similar to GP|20453072|gb AT5g60920/MSL3_40 {Arabidopsi... 151 4e-37
BE204336 similar to PIR|T51392|T51 probable phytochelatin synthe... 95 6e-22
CA920956 similar to GP|20453072|gb| AT5g60920/MSL3_40 {Arabidops... 99 3e-21
TC87911 similar to PIR|E71427|E71427 hypothetical protein - Arab... 77 1e-14
TC91281 similar to PIR|E71427|E71427 hypothetical protein - Arab... 68 8e-12
AW690428 similar to GP|21553837|gb| phytochelatin synthetase-lik... 57 2e-08
BG453618 similar to PIR|T51392|T51 probable phytochelatin synthe... 47 2e-05
BQ137157 32 0.37
TC91873 weakly similar to GP|13877655|gb|AAK43905.1 Unknown prot... 31 0.82
BQ144035 similar to GP|21732311|emb hypothetical protein {Homo s... 29 4.0
TC84625 similar to GP|3065951|gb|AAC14346.1| Notch3 {Homo sapien... 28 5.3
TC87445 similar to GP|13937224|gb|AAK50104.1 At1g74910/F9E10_24 ... 28 9.0
>TC77921 similar to GP|21553837|gb|AAM62930.1 phytochelatin synthetase-like
protein {Arabidopsis thaliana}, partial (82%)
Length = 1727
Score = 411 bits (1056), Expect(2) = e-136
Identities = 182/267 (68%), Positives = 223/267 (83%), Gaps = 4/267 (1%)
Frame = +1
Query: 4 LLLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRH 63
+LLF + TL +S+EAYDPLDPNGNITI+WD++ W DGY+A VT+ NFQQYRH
Sbjct: 178 VLLFLLISCSTLF----TSTEAYDPLDPNGNITIKWDVMGWNPDGYIAIVTMYNFQQYRH 345
Query: 64 IQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYS 123
IQEPGW+LGWTWAK E+IW M+G QTTEQGDCSKFK + PHCCKKDP VDL+PG PY+
Sbjct: 346 IQEPGWTLGWTWAKKEVIWNMMGSQTTEQGDCSKFKAGI-PHCCKKDPTVVDLLPGTPYN 522
Query: 124 EQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPA 183
+Q+ANCCKGG+L S QDP+NAV+SF++ VG AGTT K+V+LP+NFT ++PGPGYTCGPA
Sbjct: 523 QQIANCCKGGILNSWAQDPSNAVSSFQISVGSAGTTNKTVKLPRNFTLRAPGPGYTCGPA 702
Query: 184 RIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCAC 243
+IV+PT+F+ DKRRVTQA+M+WNVTCTYSQFLA K PTCCVSLS+FYNDTIV CPTC C
Sbjct: 703 KIVRPTQFITTDKRRVTQALMTWNVTCTYSQFLAQKTPTCCVSLSSFYNDTIVDCPTCTC 882
Query: 244 GCQS----GNCVDPNSPNLTSVISNPG 266
GCQ+ +CV+P+SP+L+SV+S G
Sbjct: 883 GCQNKTQPDSCVNPDSPHLSSVVSASG 963
Score = 91.7 bits (226), Expect(2) = e-136
Identities = 59/148 (39%), Positives = 86/148 (57%)
Frame = +3
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
M+ + + SSLACE L VL +G ++ FQLQ+EL +E + S A Q+
Sbjct: 990 MYKSHVSDSSSLACET*L*RVLEDQGHNHKFQLQNELLTMEPCGAASQF**SYSAFQLQV 1169
Query: 332 RVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSED 391
+VI SL G+ +*Y A+G+ +LQ*F+ SW +W C +R K +NF+ *R+G S +D
Sbjct: 1170 QVIKSLRGL-K*YCYAMGSKILQ*FSFISWITW*CAIRNLI*KRQINFHI**RMGLSKKD 1346
Query: 392 LFQW*QLCDATT*CLSKVTECRFAAKGF 419
LFQW* LC A++ C+ V + F F
Sbjct: 1347 LFQW**LCYASSRCIPLVAKRNFKISFF 1430
>TC76471 similar to PIR|T51392|T51392 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (94%)
Length = 1772
Score = 376 bits (965), Expect(2) = e-115
Identities = 168/255 (65%), Positives = 206/255 (79%), Gaps = 1/255 (0%)
Frame = +1
Query: 8 RFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEP 67
R ++ +++L S + AYDPLDPNGNITI+WD++SWT DGYVAAVT++NFQ YRHI P
Sbjct: 148 RLLISAICVIVLFSYAAAYDPLDPNGNITIKWDVVSWTADGYVAAVTMSNFQMYRHIMNP 327
Query: 68 GWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVA 127
GW+LGW+WAK E+IW M+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++Q +
Sbjct: 328 GWTLGWSWAKKEVIWSMVGSQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQQFS 504
Query: 128 NCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVK 187
NCCKGGV+ + QDP++AV++F+V VG+AGT+ K+V+LPKNFT +PGPGYTCGPA+IV
Sbjct: 505 NCCKGGVVAAWGQDPSSAVSAFQVSVGQAGTSNKTVKLPKNFTLLAPGPGYTCGPAKIVP 684
Query: 188 PTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS 247
T FL DKRR TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYN TI CP+CACGCQ+
Sbjct: 685 STTFLTTDKRRKTQALMTWNVTCTYSQFLARKNPSCCVSLSSFYNSTITPCPSCACGCQN 864
Query: 248 -GNCVDPNSPNLTSV 261
NCV S L V
Sbjct: 865 KKNCVKGRSKFLDMV 909
Score = 59.3 bits (142), Expect(2) = e-115
Identities = 47/131 (35%), Positives = 67/131 (50%)
Frame = +3
Query: 276 FMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQLRVIN 335
++P+ +LACE L +L S+ Y FQL ELF L+ T K Q + L+
Sbjct: 963 YVPH*GTLACED*L*GLLESQDCCYKFQL*IELFSLDSCCTTSKS*QCHSSFQL*LQAFA 1142
Query: 336 SLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSEDLFQW 395
L +++*+ L N +L * SW W C++R A K N + R+G S E L QW
Sbjct: 1143 CL-SIHK*HWYVLWNEIL**SVDGSWTKWKCSIRSASSKEQENIHTQPRMGIS*ESLLQW 1319
Query: 396 *QLCDATT*CL 406
*++ DA T* L
Sbjct: 1320 *RMHDAFT*YL 1352
>TC78858 similar to PIR|T51392|T51392 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (94%)
Length = 1602
Score = 370 bits (950), Expect(2) = e-115
Identities = 164/257 (63%), Positives = 205/257 (78%), Gaps = 1/257 (0%)
Frame = +2
Query: 6 LFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQ 65
L + V+ +L+L S + AYDPLDPNGNITI+WD++SWT DGYVA VT++NFQ +RHI
Sbjct: 98 LMKLVISSFCVLVLVSYAVAYDPLDPNGNITIKWDVMSWTPDGYVATVTIHNFQMFRHIM 277
Query: 66 EPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQ 125
PGW+LGWTWAK E+IW + G QTTEQGDCSKFKG + PHCCKK P VDL+PG PY++Q
Sbjct: 278 NPGWTLGWTWAKKEVIWSVTGAQTTEQGDCSKFKGNI-PHCCKKIPTVVDLLPGVPYNQQ 454
Query: 126 VANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARI 185
+NCCKGGV+ + QDPA A+++F+V VG+AGT+ ++V+LPKNFT +PGPGYTCGPA++
Sbjct: 455 FSNCCKGGVVAAWGQDPAEAISAFQVSVGQAGTSNRTVKLPKNFTLLAPGPGYTCGPAKV 634
Query: 186 VKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC 245
V T FL D+RR TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYN+TI CP CACGC
Sbjct: 635 VPSTNFLTPDRRRKTQALMTWNVTCTYSQFLARKNPSCCVSLSSFYNETITPCPHCACGC 814
Query: 246 QS-GNCVDPNSPNLTSV 261
Q+ NCV S L+ V
Sbjct: 815 QNKKNCVKSESKILSMV 865
Score = 62.4 bits (150), Expect(2) = e-115
Identities = 49/133 (36%), Positives = 70/133 (51%)
Frame = +1
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
MH +PN SLACE KL +LAS+ FQLQDEL LE N T K Q + QL
Sbjct: 907 MHSSHVPN*GSLACEDKL*GLLASQNRSNKFQLQDELLSLESCNSTSKS*QCHTSFQLQL 1086
Query: 332 RVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSED 391
+ SL +++*+ + L N +LQ* + SW ++R + + R+G SS+
Sbjct: 1087 QATCSLC-IHK*HWHVLWNEILQ*STNGSWTKRKYSIRGIASEKQRYLHIQARMGISSQG 1263
Query: 392 LFQW*QLCDATT* 404
L W* + ++T*
Sbjct: 1264 LL*W**MHASST* 1302
>TC92734 similar to GP|21553837|gb|AAM62930.1 phytochelatin synthetase-like
protein {Arabidopsis thaliana}, partial (36%)
Length = 651
Score = 266 bits (681), Expect(2) = 1e-72
Identities = 118/166 (71%), Positives = 138/166 (83%), Gaps = 2/166 (1%)
Frame = +1
Query: 12 PCTLLLLLPS--SSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
PC L L L S S++AYDPLDPNGNITI+WDI+ WT DGYVA VT+NNFQQYRHI PGW
Sbjct: 127 PCILFLFLLSFTSTDAYDPLDPNGNITIKWDIIQWTPDGYVATVTMNNFQQYRHIASPGW 306
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
SLGWTWAK E+IW M+G QTTEQGDCSKFKG V PHCCKK+P VDL+PG PY++Q+ANC
Sbjct: 307 SLGWTWAKKEVIWSMVGSQTTEQGDCSKFKGNV-PHCCKKNPTVVDLLPGTPYNQQIANC 483
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPG 175
CKGGVL+S QDP AV +F++ VGRAGTT K+V++PKNFT +PG
Sbjct: 484 CKGGVLSSWAQDPTMAVGAFQISVGRAGTTNKTVKVPKNFTLNAPG 621
Score = 25.0 bits (53), Expect(2) = 1e-72
Identities = 8/9 (88%), Positives = 9/9 (99%)
Frame = +3
Query: 176 PGYTCGPAR 184
PGYTCGPA+
Sbjct: 624 PGYTCGPAK 650
>TC83248 similar to GP|13477083|dbj|BAB02996. phytochelatin synthetase-like
protein {Arabidopsis thaliana}, partial (39%)
Length = 667
Score = 143 bits (360), Expect(2) = 9e-48
Identities = 90/149 (60%), Positives = 101/149 (67%)
Frame = +2
Query: 272 MH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQL 331
M+* ++ PSSLA A LQ VLA +G Y* QLQDEL LEL T KL QSD FQL
Sbjct: 173 MY*SYVSYPSSLAH*A*LQGVLARKGYCY*LQLQDELLRLELGCSTSKL*QSDSTFQFQL 352
Query: 332 RVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSED 391
+ NSL V++*YSNALG+*+LQ*F HASW W CT+R F KG F F *RLGFS +D
Sbjct: 353 KTNNSL-RVDK*YSNALGS*ILQ*FTHASWSYW*CTIRATFPKG-FKF*FR*RLGFSPKD 526
Query: 392 LFQW*QLCDATT*CLSKVTECRFAAKGFP 420
LFQ QLCDATT*CLS V C F K FP
Sbjct: 527 LFQRRQLCDATT*CLSMVA*CWFQTKRFP 613
Score = 65.5 bits (158), Expect(2) = 9e-48
Identities = 27/45 (60%), Positives = 37/45 (82%), Gaps = 5/45 (11%)
Frame = +3
Query: 224 CVSLSTFYNDTIVSCPTCACGC-----QSGNCVDPNSPNLTSVIS 263
CV+LS+FYNDTIV CPTC+CGC QSG+C+DP++ +L SV++
Sbjct: 3 CVALSSFYNDTIVPCPTCSCGCQGNSPQSGSCIDPSASHLASVVN 137
>CA989851 similar to GP|20453072|gb AT5g60920/MSL3_40 {Arabidopsis thaliana},
partial (41%)
Length = 799
Score = 151 bits (382), Expect = 4e-37
Identities = 98/213 (46%), Positives = 129/213 (60%), Gaps = 13/213 (6%)
Frame = +3
Query: 210 CTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC-----QSGNCVDPNSPNLTSVISN 264
CTYSQFLA K PTCCV+LS+FYNDTIV CPTC+CGC QSG C+DP++PNL SV ++
Sbjct: 3 CTYSQFLAPKTPTCCVALSSFYNDTIVPCPTCSCGCQGNSAQSGTCIDPSAPNLASVANS 182
Query: 265 PGNNGHYMH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSD 324
N P + S + C ++ + +Y ++++ + ++N ++ SD
Sbjct: 183 FPTNSTM---PLVQCTSHM-CPIQVHWHINLNDKEY-WRVK-----VTITNLNYRMNYSD 332
Query: 325 PAL*FQ------LRVINSL--WGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG* 376
L Q L ++S +N+ + +G+* LQ*F HAS W CT+R F KG*
Sbjct: 333 WNLVVQHPNFYNLTQLSSFNYKSIND-ATMIVGS*TLQ*FTHASRSYW*CTIRANFPKG* 509
Query: 377 VNFYFS*RLGFSSEDLFQW*QLCDATT*CLSKV 409
FYF *RLGFS EDLFQW QLCDATT*CLS V
Sbjct: 510 -EFYFR*RLGFSPEDLFQWRQLCDATT*CLSMV 605
>BE204336 similar to PIR|T51392|T51 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (38%)
Length = 684
Score = 95.1 bits (235), Expect(2) = 6e-22
Identities = 44/65 (67%), Positives = 50/65 (76%), Gaps = 1/65 (1%)
Frame = +1
Query: 198 RVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-GNCVDPNSP 256
R TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYN TI CP+CACGCQ+ NCV S
Sbjct: 232 RKTQALMTWNVTCTYSQFLARKNPSCCVSLSSFYNSTITPCPSCACGCQNKKNCVKGRSK 411
Query: 257 NLTSV 261
L V
Sbjct: 412 FLDMV 426
Score = 26.9 bits (58), Expect(2) = 6e-22
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = +3
Query: 276 FMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPK 319
++P+ +LACE L +L S+ Y FQL ELF L+ T K
Sbjct: 480 YVPH*GTLACED*L*GLLESQDCCYKFQL*IELFSLDSCCTTSK 611
>CA920956 similar to GP|20453072|gb| AT5g60920/MSL3_40 {Arabidopsis
thaliana}, partial (19%)
Length = 618
Score = 99.0 bits (245), Expect = 3e-21
Identities = 59/103 (57%), Positives = 74/103 (71%)
Frame = -1
Query: 329 FQLRVINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFS 388
FQL+VI SLW N+*YSNALG+*VL *F+ + P W C++ KG*+NF+F * LGF
Sbjct: 615 FQLQVIKSLW-FNK*YSNALGS*VL**FS*SCRP*WQCSI*ATLPKG*INFHF**GLGFP 439
Query: 389 SEDLFQW*QLCDATT*CLSKVTECRFAAKGFPLTCFDVNILGN 431
S+DL Q QLCDATT*CLS VT F+A+GF + CF +LG+
Sbjct: 438 SKDLLQRRQLCDATT*CLSMVT*YWFSARGF-IVCFSNGLLGS 313
>TC87911 similar to PIR|E71427|E71427 hypothetical protein - Arabidopsis
thaliana, partial (24%)
Length = 639
Score = 77.4 bits (189), Expect = 1e-14
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 8/147 (5%)
Frame = +1
Query: 107 CKKDPVAVDLIPGAPYSE----QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKS 162
C++ P +DL P +++ ++ +CC+ G + DP+ +++ F++ V +
Sbjct: 52 CQRRPTIIDL-PPTKFNDTDVGKIPSCCRNGTILPKSMDPSKSISRFQMQVYKMPPVLNR 228
Query: 163 VRL--PKNFTFK-SPGPGYTCGPARIVKPTRFLRYDKRRVTQAMM-SWNVTCTYSQFLAA 218
L P+N+ S P Y CGP V PT + + +M SW V C +
Sbjct: 229 SELSPPQNWKISGSLNPDYKCGPLVRVSPTENPDPTGLPLNKTVMASWQVVCNITTAKGT 408
Query: 219 KAPTCCVSLSTFYNDTIVSCPTCACGC 245
+ CCVS S +YN++++ C TCACGC
Sbjct: 409 SSK-CCVSFSAYYNESVIPCKTCACGC 486
>TC91281 similar to PIR|E71427|E71427 hypothetical protein - Arabidopsis
thaliana, partial (36%)
Length = 1107
Score = 67.8 bits (164), Expect = 8e-12
Identities = 38/126 (30%), Positives = 61/126 (48%), Gaps = 7/126 (5%)
Frame = +3
Query: 129 CCKGGVLTSLKQDPANAVASFRVVVGRAGT--TTKSVRLPKNFTFKSP--GPGYTCGPAR 184
CC+ G L D +++ F++ V + S+ P+N+ K P Y CG
Sbjct: 12 CCRNGTLLPPYMDLTMSISRFQIQVYKMPPHLNRSSLTPPQNWEIKGDTINPDYKCGAPI 191
Query: 185 IVKPTRFLRYDKRRVT-QAMMSWNVTCTYSQFLAAKA--PTCCVSLSTFYNDTIVSCPTC 241
+ P++ ++ SW V C S ++K+ P CCVS S +YN++++ C TC
Sbjct: 192 RISPSQSQDLTGLPSNISSIASWQVVCNISTKSSSKSDSPKCCVSFSAYYNESVIPCSTC 371
Query: 242 ACGCQS 247
ACGC S
Sbjct: 372 ACGCPS 389
>AW690428 similar to GP|21553837|gb| phytochelatin synthetase-like protein
{Arabidopsis thaliana}, partial (5%)
Length = 639
Score = 56.6 bits (135), Expect = 2e-08
Identities = 21/27 (77%), Positives = 25/27 (91%)
Frame = +1
Query: 26 YDPLDPNGNITIRWDILSWTGDGYVAA 52
YDPLDPNGNITI+WD++SWT DGYV +
Sbjct: 94 YDPLDPNGNITIKWDVVSWTADGYVVS 174
Score = 37.4 bits (85), Expect = 0.011
Identities = 14/19 (73%), Positives = 16/19 (83%)
Frame = +1
Query: 51 AAVTLNNFQQYRHIQEPGW 69
AAVT++NFQ YRHI PGW
Sbjct: 583 AAVTMSNFQMYRHIMNPGW 639
>BG453618 similar to PIR|T51392|T51 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (35%)
Length = 549
Score = 46.6 bits (109), Expect = 2e-05
Identities = 46/156 (29%), Positives = 80/156 (50%)
Frame = +1
Query: 273 H*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQLR 332
H P + +LA + +LQ +LA + ++ Q+ EL + S TP + P+L*F +
Sbjct: 1 HAPPLSCSCALAYKRQLQGLLACKDCNHQLQI*TELHTMGSSCATP*SQ*CYPSL*F*IH 180
Query: 333 VINSLWGVNE*YSNALGN*VLQ*FAHASWPSWLCTVRPAFWKG*VNFYFS*RLGFSSEDL 392
+SL ++*+ N L +LQ* + SW CT+ A+ +G + Y R+GFS + +
Sbjct: 181 ATSSL*S-HK*HRNVLWFEILQ*SNNGSWV*RECTI*GAYEEGQEHIYTKARMGFS*KGI 357
Query: 393 FQW*QLCDATT*CLSKVTECRFAAKGFPLTCFDVNI 428
W* + +T+* + + +F +K C D N+
Sbjct: 358 L*W**MYASTS*FIPFL--AKFCSKATNEFCNDDNL 459
>BQ137157
Length = 1053
Score = 32.3 bits (72), Expect = 0.37
Identities = 27/95 (28%), Positives = 38/95 (39%), Gaps = 2/95 (2%)
Frame = +3
Query: 126 VANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARI 185
V C+ V+T D A+ R R T+S + + GP P+R+
Sbjct: 447 VTRACRRRVVTQESSDATTESATQRAT-HRPAPKTRSASVSHA---RRKGPARRPAPSRV 614
Query: 186 VKPTRFLRYDKRRVTQAMMSWNVT--CTYSQFLAA 218
TR RYD R S NV CT+S+ +A
Sbjct: 615 RVRTRAHRYDYARAVSCSTSMNVVA*CTHSRLASA 719
>TC91873 weakly similar to GP|13877655|gb|AAK43905.1 Unknown protein
{Arabidopsis thaliana}, partial (5%)
Length = 608
Score = 31.2 bits (69), Expect = 0.82
Identities = 19/62 (30%), Positives = 29/62 (46%)
Frame = +1
Query: 247 SGNCVDPNSPNLTSVISNPGNNGHYMH*PFMPNPSSLACEAKLQAVLASEGDDY*FQLQD 306
S NC+D S +++ ++P G +H P PS AC A + E DD+ + D
Sbjct: 172 SSNCLDTTSAHVSETRTSPLKLGFDLHQP----PSHPACNATTWQAIIGEPDDFDESVSD 339
Query: 307 EL 308
L
Sbjct: 340 LL 345
>BQ144035 similar to GP|21732311|emb hypothetical protein {Homo sapiens},
partial (4%)
Length = 1340
Score = 28.9 bits (63), Expect = 4.0
Identities = 12/36 (33%), Positives = 24/36 (66%), Gaps = 1/36 (2%)
Frame = -2
Query: 209 TCTYSQFLAAKAPTCCVSLSTFYNDTI-VSCPTCAC 243
+C+YS+ APTC +SLS+ ++ ++ ++C +C
Sbjct: 898 SCSYSRHTPHPAPTCTLSLSSLFSVSLALACLAFSC 791
>TC84625 similar to GP|3065951|gb|AAC14346.1| Notch3 {Homo sapiens}, partial
(0%)
Length = 464
Score = 28.5 bits (62), Expect = 5.3
Identities = 15/48 (31%), Positives = 24/48 (49%), Gaps = 3/48 (6%)
Frame = -1
Query: 15 LLLLLPSSSEAYDPLDPNGNIT---IRWDILSWTGDGYVAAVTLNNFQ 59
LLL S++ P+G +T + +LSWTG+G+ TL +
Sbjct: 146 LLLTAGSATNTLTSTQPSGTLT*KRVTSMLLSWTGEGWHCLTTLQQLE 3
>TC87445 similar to GP|13937224|gb|AAK50104.1 At1g74910/F9E10_24 {Arabidopsis
thaliana}, partial (91%)
Length = 1622
Score = 27.7 bits (60), Expect = 9.0
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = -3
Query: 213 SQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCVDPNS 255
++F P CV +ST T+ +CP+ G S NC+ NS
Sbjct: 1332 NRFTKYSDPKLCVVIST----TLDTCPSTNGGFPSNNCMKNNS 1216
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,035,713
Number of Sequences: 36976
Number of extensions: 290087
Number of successful extensions: 2300
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 2252
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2286
length of query: 434
length of database: 9,014,727
effective HSP length: 99
effective length of query: 335
effective length of database: 5,354,103
effective search space: 1793624505
effective search space used: 1793624505
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0338.9


