

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
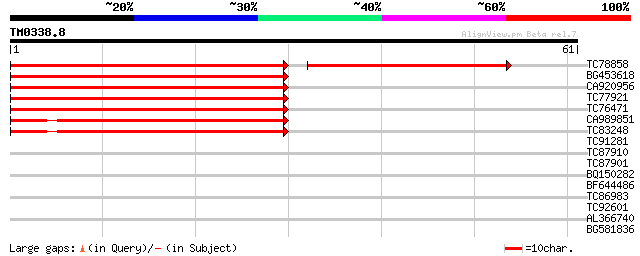
TC78858 similar to PIR|T51392|T51392 probable phytochelatin synt... 65 2e-14
BG453618 similar to PIR|T51392|T51 probable phytochelatin synthe... 67 1e-12
CA920956 similar to GP|20453072|gb| AT5g60920/MSL3_40 {Arabidops... 61 1e-10
TC77921 similar to GP|21553837|gb|AAM62930.1 phytochelatin synth... 61 1e-10
TC76471 similar to PIR|T51392|T51392 probable phytochelatin synt... 57 1e-09
CA989851 similar to GP|20453072|gb AT5g60920/MSL3_40 {Arabidopsi... 55 4e-09
TC83248 similar to GP|13477083|dbj|BAB02996. phytochelatin synth... 54 9e-09
TC91281 similar to PIR|E71427|E71427 hypothetical protein - Arab... 35 0.006
TC87910 similar to PIR|E71427|E71427 hypothetical protein - Arab... 35 0.006
TC87901 similar to PIR|A84751|A84751 probable SF16 protein (Heli... 27 1.6
BQ150282 26 2.7
BF644486 similar to GP|17933295|gb| AT5g67540/K9I9_10 {Arabidops... 25 6.0
TC86983 similar to GP|9971154|dbj|BAB12429.1 mkpA {Dictyostelium... 25 6.0
TC92601 25 7.8
AL366740 weakly similar to GP|4755186|gb unknown protein {Arabid... 25 7.8
BG581836 similar to GP|8778285|gb| F14D16.26 {Arabidopsis thalia... 25 7.8
>TC78858 similar to PIR|T51392|T51392 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (94%)
Length = 1602
Score = 64.7 bits (156), Expect(2) = 2e-14
Identities = 25/30 (83%), Positives = 29/30 (96%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+K+TF+FKQGWAFP KVYFNGDECM+PPP
Sbjct: 1211 KNKDTFTFKQGWAFPHKVYFNGDECMLPPP 1300
Score = 28.9 bits (63), Expect(2) = 2e-14
Identities = 15/22 (68%), Positives = 15/22 (68%)
Frame = +3
Query: 33 THFCQILLLQA*FPSRHSSSHY 54
T F IL LQ * P RHSSSHY
Sbjct: 1308 TLFSLILPLQD*LPLRHSSSHY 1373
>BG453618 similar to PIR|T51392|T51 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (35%)
Length = 549
Score = 67.0 bits (162), Expect = 1e-12
Identities = 26/30 (86%), Positives = 29/30 (96%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDKNTF+ KQGWAFPR+VYFNGDECM+PPP
Sbjct: 302 KDKNTFTLKQGWAFPRRVYFNGDECMLPPP 391
>CA920956 similar to GP|20453072|gb| AT5g60920/MSL3_40 {Arabidopsis
thaliana}, partial (19%)
Length = 618
Score = 60.8 bits (146), Expect = 1e-10
Identities = 22/30 (73%), Positives = 28/30 (93%)
Frame = -2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 482 KDKSTFTFDKGWAFPRRIYFNGDNCVMPPP 393
>TC77921 similar to GP|21553837|gb|AAM62930.1 phytochelatin synthetase-like
protein {Arabidopsis thaliana}, partial (82%)
Length = 1727
Score = 60.8 bits (146), Expect = 1e-10
Identities = 22/30 (73%), Positives = 28/30 (93%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 1294 KDKSTFTFDKGWAFPRRIYFNGDNCVMPPP 1383
>TC76471 similar to PIR|T51392|T51392 probable phytochelatin synthetase -
Arabidopsis thaliana, partial (94%)
Length = 1772
Score = 57.4 bits (137), Expect = 1e-09
Identities = 22/30 (73%), Positives = 26/30 (86%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+K TF+ QGWAFPR+VYFNG+ECMMP P
Sbjct: 1255 KNKKTFTLNQGWAFPRRVYFNGEECMMPSP 1344
>CA989851 similar to GP|20453072|gb AT5g60920/MSL3_40 {Arabidopsis thaliana},
partial (41%)
Length = 799
Score = 55.5 bits (132), Expect = 4e-09
Identities = 22/30 (73%), Positives = 27/30 (89%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+ F+F +GWAFPRK+YFNGD C+MPPP
Sbjct: 502 KDKS-FTFDKGWAFPRKIYFNGDNCVMPPP 588
>TC83248 similar to GP|13477083|dbj|BAB02996. phytochelatin synthetase-like
protein {Arabidopsis thaliana}, partial (39%)
Length = 667
Score = 54.3 bits (129), Expect = 9e-09
Identities = 21/30 (70%), Positives = 26/30 (86%)
Frame = +3
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD N F+F +GWAFPR++YFNGD C+MPPP
Sbjct: 477 KDSN-FNFDKGWAFPRRIYFNGDNCVMPPP 563
>TC91281 similar to PIR|E71427|E71427 hypothetical protein - Arabidopsis
thaliana, partial (36%)
Length = 1107
Score = 35.0 bits (79), Expect = 0.006
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = +3
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTH 34
K+K+ + G FP KV+FNG+EC +P P++
Sbjct: 852 KNKDEINVVGGDGFPSKVFFNGEECSLPSVFPSN 953
>TC87910 similar to PIR|E71427|E71427 hypothetical protein - Arabidopsis
thaliana, partial (25%)
Length = 859
Score = 35.0 bits (79), Expect = 0.006
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +2
Query: 11 GWAFPRKVYFNGDECMMPPPTPT 33
G FP+KV+FNG+EC +P PT
Sbjct: 437 GDGFPKKVFFNGEECSLPSVLPT 505
>TC87901 similar to PIR|A84751|A84751 probable SF16 protein (Helianthus
annuus) [imported] - Arabidopsis thaliana, partial (60%)
Length = 1434
Score = 26.9 bits (58), Expect = 1.6
Identities = 16/52 (30%), Positives = 24/52 (45%), Gaps = 4/52 (7%)
Frame = -3
Query: 13 AFPRKVYFNGDECMMPPPTPTHFCQIL----LLQA*FPSRHSSSHYSSFLFY 60
AFP + +GD +P PTP F +++ L + P + SF FY
Sbjct: 862 AFP---WTSGDPAFLPAPTPASFAKLITT*TLTRVCPPLMRKGISHGSFAFY 716
>BQ150282
Length = 286
Score = 26.2 bits (56), Expect = 2.7
Identities = 14/26 (53%), Positives = 17/26 (64%), Gaps = 1/26 (3%)
Frame = -2
Query: 35 FCQILLLQA*FPSRHSSSH-YSSFLF 59
+C +LLL + F HSSSH YS LF
Sbjct: 249 YCALLLL*SRFSRSHSSSHFYSRVLF 172
>BF644486 similar to GP|17933295|gb| AT5g67540/K9I9_10 {Arabidopsis
thaliana}, partial (7%)
Length = 610
Score = 25.0 bits (53), Expect = 6.0
Identities = 11/31 (35%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Frame = +1
Query: 11 GWAFPRKVYFNGDECMM---PPPTPTHFCQI 38
GW +Y N + + PPP P+HF Q+
Sbjct: 214 GWILLLHLYSNNNTSLNIIHPPPLPSHFHQL 306
>TC86983 similar to GP|9971154|dbj|BAB12429.1 mkpA {Dictyostelium
discoideum}, partial (0%)
Length = 1141
Score = 25.0 bits (53), Expect = 6.0
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSHY 54
K KN +S ++ ++ K F+ P P P H +LLL P+ H S Y
Sbjct: 275 KTKNLYSSEKSFS---KTQFHKTYSTSPSPIPQHLYLLLLLAP--PNYHFSHPY 421
>TC92601
Length = 455
Score = 24.6 bits (52), Expect = 7.8
Identities = 10/35 (28%), Positives = 17/35 (48%)
Frame = +3
Query: 2 DKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFC 36
D+ ++S+ A + D C+M T +HFC
Sbjct: 87 DRQSYSYPSSIANQKDYCLPTDLCIMKASTKSHFC 191
>AL366740 weakly similar to GP|4755186|gb unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 476
Score = 24.6 bits (52), Expect = 7.8
Identities = 11/30 (36%), Positives = 15/30 (49%)
Frame = -2
Query: 21 NGDECMMPPPTPTHFCQILLLQA*FPSRHS 50
N +C P P H CQ+ LQ+* + S
Sbjct: 361 NQPQCDQPSPEYRHLCQLQRLQS*IDIKDS 272
>BG581836 similar to GP|8778285|gb| F14D16.26 {Arabidopsis thaliana},
partial (18%)
Length = 749
Score = 24.6 bits (52), Expect = 7.8
Identities = 8/23 (34%), Positives = 12/23 (51%)
Frame = +3
Query: 11 GWAFPRKVYFNGDECMMPPPTPT 33
G +++YF D + PP PT
Sbjct: 60 GMKLSKRIYFGKDRAVAPPKVPT 128
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,330,781
Number of Sequences: 36976
Number of extensions: 57899
Number of successful extensions: 467
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 467
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 467
length of query: 61
length of database: 9,014,727
effective HSP length: 37
effective length of query: 24
effective length of database: 7,646,615
effective search space: 183518760
effective search space used: 183518760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0338.8


