

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.4
(197 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
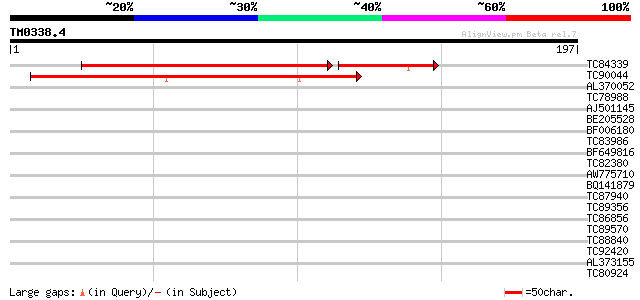
Score E
Sequences producing significant alignments: (bits) Value
TC84339 weakly similar to GP|17529298|gb|AAL38876.1 unknown prot... 137 1e-42
TC90044 93 6e-20
AL370052 weakly similar to GP|15450804|gb| endoplasmic reticulum... 31 0.27
TC78988 similar to GP|6862912|gb|AAF30301.1| unknown protein {Ar... 31 0.36
AJ501145 28 2.3
BE205528 similar to GP|12382006|d putative ABC transporter prote... 28 3.0
BF006180 similar to PIR|E86349|E86 hypothetical protein AAD41417... 28 3.0
TC83986 similar to GP|21618082|gb|AAM67132.1 unknown {Arabidopsi... 28 3.0
BF649816 weakly similar to GP|20466770|gb receptor protein kinas... 28 3.0
TC82380 similar to SP|P93236|ABA2_LYCES Zeaxanthin epoxidase ch... 27 3.9
AW775710 similar to PIR|T46096|T46 hypothetical protein T25B15.3... 27 5.1
BQ141879 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 23 5.6
TC87940 homologue to GP|12083236|gb|AAG48777.1 unknown protein {... 27 6.7
TC89356 similar to GP|19911207|dbj|BAB86930. glucosyltransferase... 27 6.7
TC86856 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase... 27 6.7
TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-bin... 26 8.8
TC88840 similar to GP|18087604|gb|AAL58932.1 AT3g58560/F14P22_15... 26 8.8
TC92420 similar to GP|6729042|gb|AAF27038.1| unknown protein {Ar... 26 8.8
AL373155 26 8.8
TC80924 similar to GP|21537146|gb|AAM61487.1 unknown {Arabidopsi... 26 8.8
>TC84339 weakly similar to GP|17529298|gb|AAL38876.1 unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 669
Score = 137 bits (344), Expect(2) = 1e-42
Identities = 64/87 (73%), Positives = 73/87 (83%)
Frame = +3
Query: 26 LDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSL 85
+ L V+SLPVVV VVD+LVPCVLISNFTC+ CY + RY+FKSSL DIPLVS++RS
Sbjct: 294 VSLIVTSLPVVVAVVDVLVPCVLISNFTCVNCYSFKQLLRRYSFKSSLMDIPLVSVLRSF 473
Query: 86 IILCVYSICDAPTLSHGPYLGTVTLCS 112
II+CVYSICD P LSHGPYLGTVTLCS
Sbjct: 474 IIICVYSICDGPALSHGPYLGTVTLCS 554
Score = 53.1 bits (126), Expect(2) = 1e-42
Identities = 29/36 (80%), Positives = 31/36 (85%), Gaps = 1/36 (2%)
Frame = +1
Query: 115 SIVLLSVKTCVFTVNSRIEAEAS-ASLKRQRLHLKK 149
SI+LLSVK C+FTVNS IEAEAS ASLKR RLHL K
Sbjct: 562 SIILLSVKACLFTVNSXIEAEASLASLKRXRLHLXK 669
>TC90044
Length = 581
Score = 93.2 bits (230), Expect = 6e-20
Identities = 52/119 (43%), Positives = 73/119 (60%), Gaps = 4/119 (3%)
Frame = +2
Query: 8 SKG*LVIIKT*SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFT--CIRCYGITKHFG 65
SK L ++KT L L + S PV++V+ D L+P L+S + + ++ HF
Sbjct: 191 SKRILRVVKTLFFLITMILSLLLFSAPVLLVIADALLPSALLSTLSPVSLSLTTLSSHFH 370
Query: 66 RYAFKSSLTDIPLVSLMRSLIILCVYSICDAPTL--SHGPYLGTVTLCSILSIVLLSVK 122
Y F+ SL DIPLVS++RS II CVYS+CD P L S GPYL T+CS+LS++ +S K
Sbjct: 371 NYDFRYSLIDIPLVSIIRSFIIFCVYSLCDGPRLSRSRGPYLCITTMCSVLSLLFVSFK 547
>AL370052 weakly similar to GP|15450804|gb| endoplasmic reticulum
alpha-mannosidase putative {Arabidopsis thaliana},
partial (4%)
Length = 440
Score = 31.2 bits (69), Expect = 0.27
Identities = 29/126 (23%), Positives = 55/126 (43%), Gaps = 3/126 (2%)
Frame = +3
Query: 48 LISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIIL---CVYSICDAPTLSHGPY 104
+ S ++C+R Y I + A ++ SL+ L C+ S+ D L + P
Sbjct: 66 IFSTYSCVRRYPICTKI*KCA*LKPANNLFFTSLLELLNYTEC*CLVSV*DVIYLEYYPS 245
Query: 105 LGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVVFS 164
LC +L+I C F ++ + + R+R + +S G LFL ++F
Sbjct: 246 CIKYVLCRLLTIKRKETNVCYFIF--QVMGQRFLTSNRKRYRISRSTG-KFLFL-ILIFG 413
Query: 165 LGHIVV 170
+ ++V+
Sbjct: 414 VTYLVL 431
>TC78988 similar to GP|6862912|gb|AAF30301.1| unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 2203
Score = 30.8 bits (68), Expect = 0.36
Identities = 23/52 (44%), Positives = 24/52 (45%)
Frame = -1
Query: 129 NSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARR 180
N RI E S LKR K W M V+FL S F L RTS R RR
Sbjct: 259 NPRIVKEGSKGLKRLNFLRKSMWVM-VVFLVSETFVLFIFQCERRTSSRRRR 107
>AJ501145
Length = 629
Score = 28.1 bits (61), Expect = 2.3
Identities = 28/108 (25%), Positives = 45/108 (40%)
Frame = +2
Query: 75 DIPLVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEA 134
DIP + + CVY + +L P +++L SIVL S K V + +
Sbjct: 122 DIP*STTKDGSVHYCVYQVRICGSLHSNPKRSSISLPLFTSIVLESFKGPVCRQSRIVHP 301
Query: 135 EASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTSCRARRSS 182
E S +R + + +P S V+ S CR++ RR+S
Sbjct: 302 EEKRSAAPRRNYPR----LPCQLASRVIRS-------CRSASTRRRTS 412
>BE205528 similar to GP|12382006|d putative ABC transporter protein {Oryza
sativa (japonica cultivar-group)}, partial (14%)
Length = 617
Score = 27.7 bits (60), Expect = 3.0
Identities = 15/50 (30%), Positives = 27/50 (54%)
Frame = +2
Query: 104 YLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGM 153
+LGTV+ ++LLS+ C ++ ++ LK +R L+KSW +
Sbjct: 62 FLGTVSKMISTLLMLLSMNLC--SIQHGSGCPQTSMLKLERCSLRKSWNL 205
>BF006180 similar to PIR|E86349|E86 hypothetical protein AAD41417.1
[imported] - Arabidopsis thaliana, partial (16%)
Length = 612
Score = 27.7 bits (60), Expect = 3.0
Identities = 15/63 (23%), Positives = 27/63 (42%), Gaps = 2/63 (3%)
Frame = -3
Query: 67 YAFKSSLTDIPLVSLMRSLIILCVYSICDAPTLSHGPYLGTV--TLCSILSIVLLSVKTC 124
+ F + +PL ++ +++ C C + S YLG TLC I + K+
Sbjct: 430 FGFPPRIISVPLPAIFVDIVMACFRPACATISASLAAYLGRAFRTLCGIFHSLSRPAKSS 251
Query: 125 VFT 127
F+
Sbjct: 250 EFS 242
>TC83986 similar to GP|21618082|gb|AAM67132.1 unknown {Arabidopsis
thaliana}, partial (45%)
Length = 783
Score = 27.7 bits (60), Expect = 3.0
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Frame = -3
Query: 30 VSSLPVVVVVVDLL--VPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLII 87
VSS P + + + C +ISN +R YG R K+++ D V ++
Sbjct: 205 VSSAPGLACIPGIFCRTSCFIISNINLLRSYGFVIRN*RDVDKNNVIDSVCV------VV 44
Query: 88 LCVYS 92
LC+YS
Sbjct: 43 LCLYS 29
>BF649816 weakly similar to GP|20466770|gb receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (12%)
Length = 658
Score = 27.7 bits (60), Expect = 3.0
Identities = 17/61 (27%), Positives = 31/61 (49%)
Frame = +1
Query: 116 IVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVVFSLGHIVVACRTS 175
+VLL + VF +E + L + L S MP++F+ V+F + ++V+ R +
Sbjct: 349 VVLLGSTSLVFEEQELVEETLNFLLTLWLISLYYSHLMPLVFIFPVLFLICLVLVSVRFA 528
Query: 176 C 176
C
Sbjct: 529 C 531
>TC82380 similar to SP|P93236|ABA2_LYCES Zeaxanthin epoxidase chloroplast
precursor (EC 1.14.-.-). [Tomato] {Lycopersicon
esculentum}, partial (11%)
Length = 624
Score = 27.3 bits (59), Expect = 3.9
Identities = 15/50 (30%), Positives = 25/50 (50%)
Frame = +3
Query: 78 LVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSVKTCVFT 127
L+S+ +I+ +S+C P + H L + LC +V K CVF+
Sbjct: 27 LLSIFAKIIVNAKHSLCFDPLIRHKDTLTKLNLCLC*FLVF---KVCVFS 167
>AW775710 similar to PIR|T46096|T46 hypothetical protein T25B15.30 -
Arabidopsis thaliana, partial (29%)
Length = 492
Score = 26.9 bits (58), Expect = 5.1
Identities = 10/16 (62%), Positives = 13/16 (80%)
Frame = -1
Query: 46 CVLISNFTCIRCYGIT 61
CV ISN +CI+CY +T
Sbjct: 228 CVWISNNSCIKCYLVT 181
>BQ141879 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (8%)
Length = 1161
Score = 23.5 bits (49), Expect(2) = 5.6
Identities = 9/27 (33%), Positives = 18/27 (66%), Gaps = 2/27 (7%)
Frame = -3
Query: 77 PLVSLMRSLIIL--CVYSICDAPTLSH 101
PL L+ ++IL C+Y++C A +++
Sbjct: 739 PLPDLVNRIVILYVCLYTVCSASLVTY 659
Score = 21.6 bits (44), Expect(2) = 5.6
Identities = 10/26 (38%), Positives = 15/26 (57%), Gaps = 4/26 (15%)
Frame = -1
Query: 40 VDLLVPCVLISNFTCIR----CYGIT 61
++ + CVLI+ TCI YG+T
Sbjct: 939 INYIYTCVLITLHTCIHHNCSIYGVT 862
>TC87940 homologue to GP|12083236|gb|AAG48777.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 790
Score = 26.6 bits (57), Expect = 6.7
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 3/60 (5%)
Frame = -1
Query: 93 ICDAPTLSHGPYLGTVTL---CSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKK 149
IC PT S LG++ + CS+LS+ L + CVF+ S + + K Q L +++
Sbjct: 268 ICSFPTFSFS--LGSILITSTCSLLSLTL*LLCFCVFSHRSNNSS*YKSIEKLQSLEIQE 95
>TC89356 similar to GP|19911207|dbj|BAB86930. glucosyltransferase-12 {Vigna
angularis}, partial (58%)
Length = 1642
Score = 26.6 bits (57), Expect = 6.7
Identities = 19/64 (29%), Positives = 28/64 (43%), Gaps = 6/64 (9%)
Frame = -1
Query: 32 SLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLT------DIPLVSLMRSL 85
SLP +++ LVPC+ S + CY I + SSL I L+S++ S
Sbjct: 1060 SLPQAHLIILFLVPCLA*SPIKTVFCYSINHAKSQLFLLSSLV*CFQS*HI*LISMVESQ 881
Query: 86 IILC 89
C
Sbjct: 880 ATPC 869
>TC86856 similar to GP|9759387|dbj|BAB10038.1 dihydropyrimidinase {Arabidopsis
thaliana}, partial (90%)
Length = 1911
Score = 26.6 bits (57), Expect = 6.7
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = +2
Query: 137 SASLKRQRLHLKKSWGMPVLFLSSVVFSL 165
S S K Q HLKK++G+ ++ ++VF +
Sbjct: 1610 SPSKKSQIYHLKKNYGIVFYYIVNIVFGI 1696
>TC89570 similar to GP|13936308|gb|AAK40307.1 putative methyl-binding domain
protein MBD105 {Zea mays}, partial (9%)
Length = 728
Score = 26.2 bits (56), Expect = 8.8
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +2
Query: 172 CRTSCRARRSSCFIESTLKL 191
CR SCR R +CF+ +T +
Sbjct: 104 CRCSCRCTRRACFV*ATCSI 163
>TC88840 similar to GP|18087604|gb|AAL58932.1 AT3g58560/F14P22_150
{Arabidopsis thaliana}, partial (20%)
Length = 788
Score = 26.2 bits (56), Expect = 8.8
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +2
Query: 97 PTLSHGPYLGTVTLCSILSIVLLSVKTCVFTV 128
PT+S P L V C+I IV +K +F V
Sbjct: 437 PTMSPNPLL*MVIFCAISGIVYRVIKRLLFVV 532
>TC92420 similar to GP|6729042|gb|AAF27038.1| unknown protein {Arabidopsis
thaliana}, partial (12%)
Length = 607
Score = 26.2 bits (56), Expect = 8.8
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -1
Query: 67 YAFKSSLTDIPLVSLMRSLIILCVYS 92
YA + LT +P +S +LII CV S
Sbjct: 241 YAKEGHLTSLPHISFTPNLIIFCVLS 164
>AL373155
Length = 485
Score = 26.2 bits (56), Expect = 8.8
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = +2
Query: 46 CVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLI 86
C I +C CY I HF ++ T +P+ L+R+L+
Sbjct: 302 CFRIQKSSCCCCYAILCHFSASMYELGGTLVPI--LVRTLV 418
>TC80924 similar to GP|21537146|gb|AAM61487.1 unknown {Arabidopsis
thaliana}, partial (21%)
Length = 887
Score = 26.2 bits (56), Expect = 8.8
Identities = 12/19 (63%), Positives = 13/19 (68%)
Frame = -1
Query: 146 HLKKSWGMPVLFLSSVVFS 164
H K SW +P LF SS VFS
Sbjct: 443 HKKLSWKIPPLFSSSKVFS 387
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.343 0.149 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,280,474
Number of Sequences: 36976
Number of extensions: 119535
Number of successful extensions: 1532
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 1518
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1529
length of query: 197
length of database: 9,014,727
effective HSP length: 91
effective length of query: 106
effective length of database: 5,649,911
effective search space: 598890566
effective search space used: 598890566
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0338.4


