

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.2
(282 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
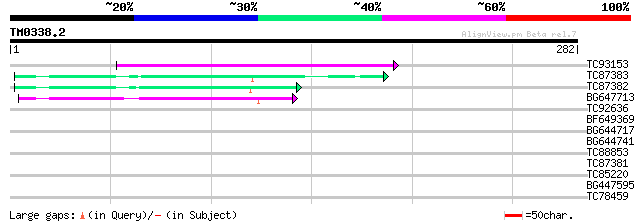
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 108 3e-24
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 49 2e-06
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 47 1e-05
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 47 1e-05
TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iride... 38 0.005
BF649369 34 0.055
BG644717 32 0.36
BG644741 30 0.80
TC88853 similar to SP|O04350|TBCA_ARATH Tubulin-specific chapero... 30 0.80
TC87381 30 1.0
TC85220 weakly similar to PIR|S33520|S33520 Lea protein - soybea... 29 1.8
BG447595 28 5.2
TC78459 similar to GP|12643061|gb|AAK00450.1 unknown protein {Or... 27 8.8
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 108 bits (269), Expect = 3e-24
Identities = 55/140 (39%), Positives = 77/140 (54%)
Frame = +2
Query: 54 DQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAE 113
D R L F R PP F G PD A W++EIE+IF V+Q E KV T++L +A+
Sbjct: 92 DGTRMLETFLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQCFETQKVQFGTHMLAEEAD 271
Query: 114 YWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKL 173
WW ++E + V W FR FL +YFP+ R ++E +FL L+QG M++ EYAAK
Sbjct: 272 DWWISLLPVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKF 451
Query: 174 ESLAKHFRFFRDQVDEPYMC 193
LA + + + E C
Sbjct: 452 VELATFYPHYSAETAEFSKC 511
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 48.9 bits (115), Expect = 2e-06
Identities = 46/188 (24%), Positives = 74/188 (38%), Gaps = 2/188 (1%)
Frame = -2
Query: 3 KRIQNMVNANQLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDF 62
+++Q +VNA Q L++A + D+ + +R +RE + ND
Sbjct: 880 QQLQEIVNAQQ------ALLEAQQKRFKDHVSSSDSLSSRSSRSQRREFQM------NDI 737
Query: 63 RRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGA--R 120
+ D P F G PD+ W+Q +E++F+ + E KV + L A WW R
Sbjct: 736 K*-DIPDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRR 560
Query: 121 GMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHF 180
E W R KY P Q + T P+ + K S +HF
Sbjct: 559 RKREGKSKIKTWEKMRQKLTRKYLPPH-----------YYQDNYTQPQLSKK--SSYRHF 419
Query: 181 RFFRDQVD 188
++Q+D
Sbjct: 418 SPTKNQID 395
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 46.6 bits (109), Expect = 1e-05
Identities = 36/145 (24%), Positives = 58/145 (39%), Gaps = 2/145 (1%)
Frame = +2
Query: 3 KRIQNMVNANQLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDF 62
+++Q ++NA Q L++A + + + +R H +R+ + ND
Sbjct: 473 QQLQEIINAQQ------ALLEAEQRRFEGDVSYSDSSSSRSSHSQRRQLQM------NDI 616
Query: 63 RRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRG--AR 120
+ D P F G D+ W+Q IE++FE + E KV + L A WW R
Sbjct: 617 K-VDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRR 793
Query: 121 GMMEANHVEVNWNSFRVAFLEKYFP 145
E W+ R KY P
Sbjct: 794 RKREGKSKIKTWDKMRQKLTRKYLP 868
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 46.6 bits (109), Expect = 1e-05
Identities = 34/141 (24%), Positives = 57/141 (40%), Gaps = 2/141 (1%)
Frame = +2
Query: 5 IQNMVNANQLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRR 64
+Q VNA Q L++A + +D+ + +R +R+ + + +
Sbjct: 197 LQETVNAQQ------ALLEAQRRRNDDDGSGSDSSSSRSSRSHRRQTRMSKIK------- 337
Query: 65 QDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGM-- 122
D P F G PDE W+Q IE++F+ + +E KV + L A WW+ +
Sbjct: 338 VDIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRN 517
Query: 123 MEANHVEVNWNSFRVAFLEKY 143
E W+ R KY
Sbjct: 518 CEGKSKIKTWDKMRQKLTRKY 580
>TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iridescent
virus}, partial (1%)
Length = 772
Score = 37.7 bits (86), Expect = 0.005
Identities = 28/96 (29%), Positives = 40/96 (41%), Gaps = 8/96 (8%)
Frame = +3
Query: 13 QLAEMVATLVQAMTVQTNDNAQ--------RRAAEDARELHRLQREAALDQNRGLNDFRR 64
Q EM Q +Q NAQ RR +D R + + + L + +
Sbjct: 249 QEMEMEEMRRQIQELQETVNAQQAILEAERRRVDDDGSSDSSSSRSSRSHRRKTLMNDIK 428
Query: 65 QDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAK 100
D P F G PDE W+Q IE++FE + GA+
Sbjct: 429 VDIPDFEGELQPDEFVDWLQAIERVFEYKEIPRGAQ 536
>BF649369
Length = 631
Score = 34.3 bits (77), Expect = 0.055
Identities = 35/160 (21%), Positives = 65/160 (39%)
Frame = +3
Query: 68 PKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANH 127
P F G D+ WI E F+V T + +V L+ + G +W+ ++
Sbjct: 135 PLFEG----DDPVAWITRAEIYFDVQNTPDDMRVKLSRLSMEGPTIHWF----NLLMETE 290
Query: 128 VEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQV 187
+++ + A + +Y D R E + L+ + ++ E+ E L+ ++
Sbjct: 291 DDLSREKLKKALIARY--DGRRLENPFEELSTLRQIGSVEEFVEAFELLSSQV----GRL 452
Query: 188 DEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
E F+ GL+A I VR L ++ A +VE
Sbjct: 453 PEEQYLGYFMSGLKAHIRRRVRTLNPTTRMQMMRIAKDVE 572
>BG644717
Length = 267
Score = 31.6 bits (70), Expect = 0.36
Identities = 16/41 (39%), Positives = 24/41 (58%)
Frame = -2
Query: 68 PKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLL 108
P+F G ++ ++ EI+KIFEV+ S V LA+Y L
Sbjct: 260 PEFLGSQINEDPQNFLDEIKKIFEVMHVSGNDLVELASYQL 138
>BG644741
Length = 735
Score = 30.4 bits (67), Expect = 0.80
Identities = 24/97 (24%), Positives = 46/97 (46%), Gaps = 5/97 (5%)
Frame = -2
Query: 108 LLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSAR---DERESQFLTL--RQG 162
L+G+A+ W+ + N + WN R FL +Y+P S + ++R + F+ L
Sbjct: 566 LMGEADIWFTE----LPYNSI-FTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGESV 402
Query: 163 SMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRG 199
S + + + L S+ H ++D+ + + F RG
Sbjct: 401 SSSWDRFTSFLRSVPNH------RIDDDSLKEYFYRG 309
>TC88853 similar to SP|O04350|TBCA_ARATH Tubulin-specific chaperone A
(Tubulin-folding cofactor A) (CFA) (TCP1-chaperonin
cofactor A homolog), partial (73%)
Length = 689
Score = 30.4 bits (67), Expect = 0.80
Identities = 20/59 (33%), Positives = 33/59 (55%), Gaps = 5/59 (8%)
Frame = +3
Query: 46 RLQREAALDQNRG-----LNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGA 99
R + EAAL+ +G LN+ +++ P+ DEA I E+EK+FE ++ EG+
Sbjct: 384 RKRLEAALEDLKGILAELLNETDKKESPEI------DEARNTIVEVEKVFETIEA*EGS 542
>TC87381
Length = 814
Score = 30.0 bits (66), Expect = 1.0
Identities = 20/62 (32%), Positives = 27/62 (43%)
Frame = +3
Query: 59 LNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRG 118
+ND + D P F G DE +Q IE +FE + E KV + L A WW
Sbjct: 612 MNDIK-VDIPDFEGELQSDEFVD*LQAIECVFEYKEIPEDHKVKVVAV*LKKHALIWWEN 788
Query: 119 AR 120
+
Sbjct: 789 LK 794
>TC85220 weakly similar to PIR|S33520|S33520 Lea protein - soybean, partial
(69%)
Length = 1880
Score = 29.3 bits (64), Expect = 1.8
Identities = 12/36 (33%), Positives = 19/36 (52%)
Frame = +2
Query: 230 KNRRMDRAGTGGPMRSSPRSYQGKGKLQMKKPYQRP 265
K ++ R G GP R SP++ K K++ P +P
Sbjct: 242 KGKKPQRHGLSGPKRKSPKALASKTKIRTAPPLIKP 349
>BG447595
Length = 309
Score = 27.7 bits (60), Expect = 5.2
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = -2
Query: 127 HVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGS 163
H E NW SFR F EK S R + + LR+GS
Sbjct: 278 HSERNWGSFRA*FGEKQ*IYSRRGQNWQEKWKLREGS 168
>TC78459 similar to GP|12643061|gb|AAK00450.1 unknown protein {Oryza
sativa}, partial (67%)
Length = 1619
Score = 26.9 bits (58), Expect = 8.8
Identities = 8/23 (34%), Positives = 16/23 (68%)
Frame = +1
Query: 123 MEANHVEVNWNSFRVAFLEKYFP 145
+++NH+ +N SF ++F +FP
Sbjct: 112 IKSNHIHINQKSFSISFSVSFFP 180
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,428,233
Number of Sequences: 36976
Number of extensions: 69754
Number of successful extensions: 293
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 292
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 293
length of query: 282
length of database: 9,014,727
effective HSP length: 95
effective length of query: 187
effective length of database: 5,502,007
effective search space: 1028875309
effective search space used: 1028875309
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0338.2


