

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0336.12
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
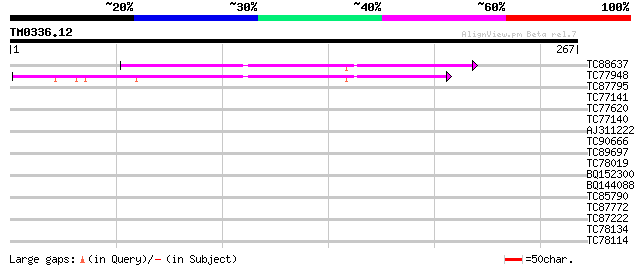
Score E
Sequences producing significant alignments: (bits) Value
TC88637 similar to GP|14596217|gb|AAK68836.1 Unknown protein {Ar... 115 2e-26
TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30... 112 1e-25
TC87795 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde ... 32 0.26
TC77141 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde ... 30 0.74
TC77620 similar to GP|10129643|emb|CAC08239. Pspzf zinc finger p... 30 1.3
TC77140 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde ... 29 2.2
AJ311222 similar to GP|19850241|gb cytosolic aldehyde dehydrogen... 28 3.7
TC90666 weakly similar to GP|21594566|gb|AAM66022.1 unknown {Ara... 28 3.7
TC89697 similar to GP|4886756|gb|AAD32031.1| cellulose synthase ... 28 4.8
TC78019 similar to GP|9294410|dbj|BAB02491.1 protein kinase-like... 27 6.3
BQ152300 homologue to PIR|H84602|H84 40S ribosomal protein S25 [... 27 6.3
BQ144088 homologue to PIR|H84602|H84 40S ribosomal protein S25 [... 27 6.3
TC85790 homologue to PIR|H84602|H84602 40S ribosomal protein S25... 27 6.3
TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper ester... 27 6.3
TC87222 homologue to PIR|H84602|H84602 40S ribosomal protein S25... 27 6.3
TC78134 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Ar... 27 8.2
TC78114 similar to GP|12643052|gb|AAK00441.1 unknown protein {Or... 27 8.2
>TC88637 similar to GP|14596217|gb|AAK68836.1 Unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 1207
Score = 115 bits (287), Expect = 2e-26
Identities = 63/169 (37%), Positives = 95/169 (55%), Gaps = 1/169 (0%)
Frame = +1
Query: 53 KSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNK 112
K+ K ++ K + F A L +G + + LV KC FA++KL+WE+L+
Sbjct: 394 KANKNGRASVEKLKASNFPATHLKIGSWEYKSKYEGDLVAKCYFAKQKLVWEVLEGEL-- 567
Query: 113 KFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGG 171
K K E+ W IS ++A ++ L + + QP+F +E N Q HT W+ S DFTGG
Sbjct: 568 KSKIEIQWSDISQLKANCPDDGPSTLTLMVARQPLFFRETNPQPRKHTLWQ-STTDFTGG 744
Query: 172 HAMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYFGP 220
A +R H L+ G L +HYE +++C++RL LSQQP +DSP+F P
Sbjct: 745 QASIHRRHVLQCEQGLLIKHYEKLVQCNDRLKFLSQQPEIMVDSPHFDP 891
>TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30780
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1916
Score = 112 bits (280), Expect = 1e-25
Identities = 75/225 (33%), Positives = 115/225 (50%), Gaps = 18/225 (8%)
Frame = +2
Query: 2 NSKPPFGLELSSSTMNERE---PHLRNVCPKP-PLGL--KRTTSAMKEREKMWRKMQKSK 55
N +P F S + + P N +P PLGL K++ S + + + +SK
Sbjct: 185 NKRPKFDFNSDSPKCDSGDDSGPISYNPLDEPSPLGLRLKKSPSLLDLIQMKLSQTYESK 364
Query: 56 KQN-----------SSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWE 104
K++ ++ K + F A +L +G + + LV KC FA+ KL+WE
Sbjct: 365 KKDQKGSASASAAAAAADSKLKASNFPATVLKIGTWEYKSRYEGDLVAKCYFAKHKLVWE 544
Query: 105 ILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEI 163
+LD K K E+PW I A++A ++ LE+ L +P+F +EIN Q HT W+
Sbjct: 545 VLDGCL--KNKIEIPWSDIMALKANYPDDAPGTLEVILARRPLFFREINPQPRKHTLWQ- 715
Query: 164 SDHDFTGGHAMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQ 208
S DFTGG A +R H ++ G LG+H+E +++CD RL LSQQ
Sbjct: 716 STTDFTGGQASMHRRHFVQCPQGLLGRHFEKLIQCDPRLNFLSQQ 850
>TC87795 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde dehydrogenase
{Oryza sativa (japonica cultivar-group)}, partial (92%)
Length = 1788
Score = 32.0 bits (71), Expect = 0.26
Identities = 15/44 (34%), Positives = 23/44 (52%)
Frame = +3
Query: 217 YFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTIN 260
Y+ P +F NI+D + DE+ VMAL + + +KS N
Sbjct: 1245 YYIEPTIFSNIKDDMVIAQDEIFGPVMALKKFKTIEEAIKSANN 1376
>TC77141 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde dehydrogenase
{Oryza sativa (japonica cultivar-group)}, partial (93%)
Length = 1699
Score = 30.4 bits (67), Expect = 0.74
Identities = 14/47 (29%), Positives = 24/47 (50%)
Frame = +3
Query: 214 DSPYFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTIN 260
D Y+ P +F N+++ + DE+ VMAL + + +KS N
Sbjct: 1206 DKGYYIEPTIFSNVKEDMRIAQDEIFGPVMALMKFKTIEEAIKSANN 1346
>TC77620 similar to GP|10129643|emb|CAC08239. Pspzf zinc finger protein-like
{Arabidopsis thaliana}, partial (26%)
Length = 1958
Score = 29.6 bits (65), Expect = 1.3
Identities = 14/51 (27%), Positives = 26/51 (50%)
Frame = +1
Query: 21 PHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFY 71
PH + + P + ++ ++A R MWRKM+ S+ + FA ++ Y
Sbjct: 1558 PHPQTMQPSVAMRIRMISNAASARRSMWRKMKWGVYYASTNIMLFAYSSGY 1710
>TC77140 similar to GP|14164407|dbj|BAB55806. cytosolic aldehyde dehydrogenase
{Oryza sativa (japonica cultivar-group)}, partial (94%)
Length = 1825
Score = 28.9 bits (63), Expect = 2.2
Identities = 13/44 (29%), Positives = 23/44 (51%)
Frame = +3
Query: 217 YFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTIN 260
Y+ P +F N+++ + DE+ VMAL + + +KS N
Sbjct: 1215 YYIEPTIFSNVKEDMLIAQDEIFGPVMALMKFKTIEEAIKSANN 1346
>AJ311222 similar to GP|19850241|gb cytosolic aldehyde dehydrogenase RF2C
{Zea mays}, partial (27%)
Length = 420
Score = 28.1 bits (61), Expect = 3.7
Identities = 13/44 (29%), Positives = 23/44 (51%)
Frame = -3
Query: 217 YFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTIN 260
Y+ P +F N+++ + DE+ VMAL + + +KS N
Sbjct: 319 YYIEPTIFSNVKEDMLIAKDEIFGPVMALMKFKTVEEAIKSANN 188
>TC90666 weakly similar to GP|21594566|gb|AAM66022.1 unknown {Arabidopsis
thaliana}, partial (58%)
Length = 664
Score = 28.1 bits (61), Expect = 3.7
Identities = 17/39 (43%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Frame = +2
Query: 38 TSAMKEREKMWRKMQKSKK-QNSSTTLKFAPATFYAKLL 75
+S K+REK WR Q SKK +N LK A + K L
Sbjct: 248 SSMKKKREKEWRGKQYSKKRRNREKKLKHAEKVYGKKCL 364
>TC89697 similar to GP|4886756|gb|AAD32031.1| cellulose synthase catalytic
subunit {Arabidopsis thaliana}, partial (26%)
Length = 1011
Score = 27.7 bits (60), Expect = 4.8
Identities = 15/59 (25%), Positives = 31/59 (52%), Gaps = 3/59 (5%)
Frame = +3
Query: 45 EKMWRKMQKS---KKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRK 100
+KMW + K+ K++++S +K + N+G +I T + +++V+ RRK
Sbjct: 306 KKMWMILNKNLKWKRKSTSLCIKII*IALMMMIQNIGSNPYILTVLEKIMVQN*ITRRK 482
>TC78019 similar to GP|9294410|dbj|BAB02491.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (58%)
Length = 1454
Score = 27.3 bits (59), Expect = 6.3
Identities = 16/76 (21%), Positives = 32/76 (42%)
Frame = -1
Query: 25 NVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIP 84
N CP P + T SA+ E + +K + + ++ + ++L + +
Sbjct: 770 NSCPSPRITSGATYSAVPTNELDFSPSEKLNPRRLTGLIRRLSGILHRRVLR*DSWILLS 591
Query: 85 TTVNQLVVKCQFARRK 100
+TV L V C+ R +
Sbjct: 590 STVKVLTVLCESLRER 543
>BQ152300 homologue to PIR|H84602|H84 40S ribosomal protein S25 [imported] -
Arabidopsis thaliana, partial (92%)
Length = 645
Score = 27.3 bits (59), Expect = 6.3
Identities = 19/74 (25%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Frame = +3
Query: 30 PPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTV-- 87
PP + K+++K W K ++ +K N+ + F T+ L KY I +V
Sbjct: 15 PPSSKPAKSGGGKQKKKKWSKGKQKEKVNN--MVLFDQGTYDKLLTEAPKYKLITPSVLS 188
Query: 88 NQLVVKCQFARRKL 101
++L + ARR +
Sbjct: 189 DRLRINGSLARRAI 230
>BQ144088 homologue to PIR|H84602|H84 40S ribosomal protein S25 [imported] -
Arabidopsis thaliana, partial (92%)
Length = 773
Score = 27.3 bits (59), Expect = 6.3
Identities = 19/74 (25%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Frame = +1
Query: 30 PPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTV-- 87
PP + K+++K W K ++ +K N+ + F T+ L KY I +V
Sbjct: 82 PPSSKPAKSGGGKQKKKKWSKGKQKEKVNN--MVLFDQGTYDKLLTEAPKYKLITPSVLS 255
Query: 88 NQLVVKCQFARRKL 101
++L + ARR +
Sbjct: 256 DRLRINGSLARRAI 297
>TC85790 homologue to PIR|H84602|H84602 40S ribosomal protein S25 [imported]
- Arabidopsis thaliana, partial (92%)
Length = 695
Score = 27.3 bits (59), Expect = 6.3
Identities = 19/74 (25%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Frame = +1
Query: 30 PPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTV-- 87
PP + K+++K W K ++ +K N+ + F T+ L KY I +V
Sbjct: 115 PPSSKPAKSGGGKQKKKKWSKGKQKEKVNN--MVLFDQGTYDKLLTEAPKYKLITPSVLS 288
Query: 88 NQLVVKCQFARRKL 101
++L + ARR +
Sbjct: 289 DRLRINGSLARRAI 330
>TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper esterase
{Capsicum annuum}, partial (34%)
Length = 1313
Score = 27.3 bits (59), Expect = 6.3
Identities = 17/47 (36%), Positives = 25/47 (53%)
Frame = -2
Query: 221 PVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTINHELSQHL 267
P L + DYNT HD+L++ +T L++ ST+N SQ L
Sbjct: 907 PYLHHSPLDYNTHNHDQLLKV-----KTNSTLNQ*LSTVNSNPSQVL 782
>TC87222 homologue to PIR|H84602|H84602 40S ribosomal protein S25 [imported]
- Arabidopsis thaliana, partial (92%)
Length = 751
Score = 27.3 bits (59), Expect = 6.3
Identities = 19/74 (25%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Frame = +1
Query: 30 PPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTV-- 87
PP + K+++K W K ++ +K N+ + F T+ L KY I +V
Sbjct: 100 PPSSKPAKSGGGKQKKKKWSKGKQKEKVNN--MVLFDQGTYDKLLTEAPKYKLITPSVLS 273
Query: 88 NQLVVKCQFARRKL 101
++L + ARR +
Sbjct: 274 DRLRINGSLARRAI 315
>TC78134 similar to GP|9795609|gb|AAF98427.1| Unknown protein {Arabidopsis
thaliana}, partial (46%)
Length = 1133
Score = 26.9 bits (58), Expect = 8.2
Identities = 19/79 (24%), Positives = 35/79 (44%), Gaps = 1/79 (1%)
Frame = +2
Query: 190 QHYENMLKCDERLMK-LSQQPFPRLDSPYFGPPVLFKNIRDYNTVQHDELMEQVMALDRT 248
QH+ ++ K L++ L QQP +L P +R QH + Q
Sbjct: 251 QHHLSLQKQSPDLIQILIQQPLFKLTQPLSNK*FKC*LVRQKQPQQHQQQQNQQQNQTNN 430
Query: 249 QPFLSELKSTINHELSQHL 267
+ +L++ ++ +H+ QHL
Sbjct: 431 KIYLNKQETLTSHQSKQHL 487
>TC78114 similar to GP|12643052|gb|AAK00441.1 unknown protein {Oryza
sativa}, partial (94%)
Length = 1077
Score = 26.9 bits (58), Expect = 8.2
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = -3
Query: 175 TYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPY 217
T + H ++G HY ++R+ + S+QPFP SP+
Sbjct: 241 TMGVDHRRRKEWEVGHHYN-----EQRVCRNSKQPFPCSPSPF 128
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,291,184
Number of Sequences: 36976
Number of extensions: 110840
Number of successful extensions: 590
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 586
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 588
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0336.12


