

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0335b.5
(1648 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
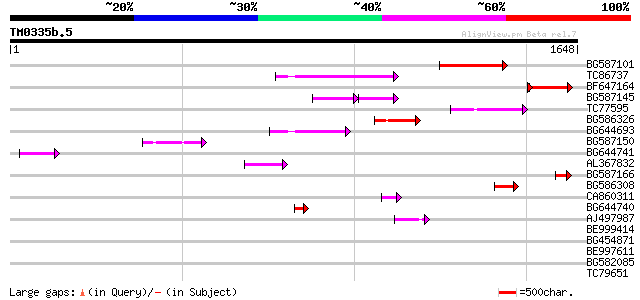
Score E
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 271 2e-72
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 182 9e-46
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 159 3e-44
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 86 8e-27
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 97 5e-20
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 93 7e-19
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 92 2e-18
BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, part... 76 1e-13
BG644741 63 1e-09
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 59 1e-08
BG587166 similar to GP|8778801|gb| T32E20.9 {Arabidopsis thalian... 56 1e-07
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 52 1e-06
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 49 1e-05
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 44 5e-04
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 44 7e-04
BE999414 40 0.006
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 39 0.017
BE997611 38 0.029
BG582085 36 0.11
TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (E... 32 1.6
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 271 bits (692), Expect = 2e-72
Identities = 121/199 (60%), Positives = 152/199 (75%)
Frame = +2
Query: 1249 LHDAKSYLWDDPFLFKICSDGVIRRCITEVDFEKILWHCHGSSYGGHFSGERTAAKVLQS 1308
L D + YLWD+PFL+K C+D + RC+ E + IL+HCHGS+Y GHF+ +T +K+ Q+
Sbjct: 2 LKDVRRYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQA 181
Query: 1309 GFYWPTLHRNSRAFVESCDRCQRTGNISRRNEMPLKNILEIELFDVWGIDFMGPFPPSFG 1368
GF+WPT+ +++ +F+ CD CQR GNIS RNEMP ILE+E+FDVWGIDFMGPFP S+
Sbjct: 182 GFWWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYN 361
Query: 1369 C*YILVAVDYVSKWVEASALSTNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFE 1428
YILVAVDYVSKWVEA A TND+ VVV K IF RFGVPR +ISDGG+HF N+ FE
Sbjct: 362 NKYILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFE 541
Query: 1429 SLLEKYGVKHKVSTPYHPQ 1447
LL+K GV+HKV+T YHPQ
Sbjct: 542 KLLKKNGVRHKVATAYHPQ 598
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 182 bits (462), Expect = 9e-46
Identities = 121/359 (33%), Positives = 185/359 (50%), Gaps = 4/359 (1%)
Frame = +1
Query: 774 VVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANENNELIPTRQVTKWRVCID 833
V++K + LLD G I S S +PV V K GG R C+D
Sbjct: 508 VLKKTLEDLLDKGFI-KASGSAAGAPVLFVRKPGGGI------------------RFCVD 630
Query: 834 YRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYG 893
YR LN++T+KD +PLP I + L R+AG +++ LD + ++++ + EDQEKTAF YG
Sbjct: 631 YRALNAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYG 810
Query: 894 VFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVF--GPNFDACLGNLALV 951
+F + PFGL APATFQR + + ++ + ++DD ++ G D + V
Sbjct: 811 LFEWIVCPFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDH-EAQVRRV 987
Query: 952 LKRCQETNLVLNWEKCHFMVRDGIVLGHKVSE-KGIEVDRAKIEVIEKLTPPTNIKGIRS 1010
L+R + L L+ +KC F V +G ++ KG+ D K+ I PP ++KG RS
Sbjct: 988 LRRLADAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARS 1167
Query: 1011 FLGHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDW 1070
FLG +Y+ FI +S++ +P+T L K+ PF + AF +K PV+ D
Sbjct: 1168 FLGFCNYYKDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDP 1347
Query: 1071 SLPFEIMCDASDLALGAVLCQKK-ERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFAC 1128
+ D S ALG VL Q+ + + + S+ L+ A+ NY +KELL V+AC
Sbjct: 1348 EAVTTVETDCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELL-AVWAC 1521
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 159 bits (402), Expect(2) = 3e-44
Identities = 81/125 (64%), Positives = 95/125 (75%)
Frame = +1
Query: 1512 LEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNRE 1571
L K I+ LN D+K A KR+L+LNEL+E RL AYE+A IYKE+TKKWHDR+I+ RE
Sbjct: 28 LSIKPIGPIKALNMDYKAAGGKRVLELNELEELRLDAYENAKIYKERTKKWHDRRIIRRE 207
Query: 1572 FVSGQLVLLFNSRLRLFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKV 1631
F G+LVLLFNSRL+LFPGKL+S WSGPF VK V P GAVEV + E+ FTVNGQRLK
Sbjct: 208 FREGELVLLFNSRLKLFPGKLRSHWSGPFQVKNVMPSGAVEVWS-ESTGPFTVNGQRLKH 384
Query: 1632 YQGGE 1636
Y GE
Sbjct: 385 YTSGE 399
Score = 39.7 bits (91), Expect(2) = 3e-44
Identities = 15/16 (93%), Positives = 16/16 (99%)
Frame = +3
Query: 1504 KACHLPVELEHKAYWA 1519
K+CHLPVELEHKAYWA
Sbjct: 3 KSCHLPVELEHKAYWA 50
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 85.5 bits (210), Expect(2) = 8e-27
Identities = 50/134 (37%), Positives = 73/134 (54%)
Frame = +2
Query: 880 PEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSVFGP 939
P+D EKTAF G + YK MPFGL NA +T+QR + +F+D + +E+++DD V
Sbjct: 8 PDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKSL 187
Query: 940 NFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVDRAKIEVIEKL 999
L +L K E + LN KC F V G LG+ V+++GIEV+ +I I L
Sbjct: 188 RATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAILDL 367
Query: 1000 TPPTNIKGIRSFLG 1013
P N + ++ G
Sbjct: 368 PSPKNSREVQRLTG 409
Score = 55.1 bits (131), Expect(2) = 8e-27
Identities = 33/118 (27%), Positives = 51/118 (42%)
Frame = +3
Query: 1013 GHAGFYRRFIKDFSKLAKPMTNLLEKEAPFTFDENCLRAFESIKESLVTAPVIVAPDWSL 1072
G RFI + P LL F +DE C AFE +K+ L T PV+ P+
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 1073 PFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEK 1130
+ S A+ +VL ++ I+Y S+ + + + Y T EK V+ + K
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 97.1 bits (240), Expect = 5e-20
Identities = 65/229 (28%), Positives = 105/229 (45%), Gaps = 7/229 (3%)
Frame = +2
Query: 1282 KILWHCHGSSYGGHFSGERTAAKVLQSGFYWPTLHRNSRAFVESCDRCQ-----RTGNIS 1336
K++ H S+ GH G +++ F+WP + R FV +CD C R
Sbjct: 176 KLVQESHDSTAAGH-PGRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRG 352
Query: 1337 RRNEMPLKNILEIELFDVWGIDFMGPFPPSFG--C*YILVAVDYVSKWVEASALSTNDSK 1394
+P+ N L +L +DF+ PP+ G Y+ V VD +SK V + T +++
Sbjct: 353 FLKPLPVPNRLHSDL----SMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAE 520
Query: 1395 VVVAFLKKNIFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEI 1454
+ G+P++I+SD G+++ R + GV +ST YHPQT G E
Sbjct: 521 ACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 1455 SNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKTPIGTSPFHLVFG 1503
N+E++ +L V S+ +W L A R + IG +PF + G
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHG 847
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 93.2 bits (230), Expect = 7e-19
Identities = 53/132 (40%), Positives = 81/132 (61%)
Frame = +2
Query: 1061 TAPVIVAPDWSLPFEIMCDASDLALGAVLCQKKERVLYVIYYASRVLNEAQRNYTTTEKE 1120
+AP++V P+ + + + DAS LG VL Q ++ VI YASR L + + NY T + E
Sbjct: 11 SAPILVLPEL-ITYVVYTDASITGLGCVLTQHEK----VIAYASRQLRKHEGNYPTHDLE 175
Query: 1121 LLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIIDRR 1180
+ VVFA + +R Y+ G KV +HTDH +L+++F + + R RW+ + ++DL+I
Sbjct: 176 MAAVVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYP 355
Query: 1181 GKDNSVADHLSR 1192
GK N VAD LSR
Sbjct: 356 GKANLVADALSR 391
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 91.7 bits (226), Expect = 2e-18
Identities = 70/234 (29%), Positives = 107/234 (44%)
Frame = +2
Query: 756 NYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPVQVVPKKGGITVVANE 815
N PI P R+NP V++ ++ LL+ G I P V V + KK G
Sbjct: 62 NMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFL----- 223
Query: 816 NNELIPTRQVTKWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQ 875
R+ IDY +LN+V K +PLP ID++ D L G +++ +D G +Q
Sbjct: 224 -------------RMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQ 364
Query: 876 ICVAPEDQEKTAFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFS 935
V ED KTAF YG + M FG N P F M +F D +++ + +F +D
Sbjct: 365 HRVIGEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDIL 544
Query: 936 VFGPNFDACLGNLALVLKRCQETNLVLNWEKCHFMVRDGIVLGHKVSEKGIEVD 989
++ N + +L L LK ++ L +V G H +S +G++VD
Sbjct: 545 IYSKNENEHENHLRLALKVLKDIGL-CQISYV*ILVEVGFFSLHVISGEGLKVD 703
>BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, partial (1%)
Length = 754
Score = 75.9 bits (185), Expect = 1e-13
Identities = 59/191 (30%), Positives = 96/191 (49%), Gaps = 3/191 (1%)
Frame = +2
Query: 385 KALVKKNLDKQFS-KFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETI 443
K V K+L + + K +E F+K IP + E+ A F + R+ E +E I
Sbjct: 110 KKKVAKHLKRGANEKEMESFQKRVFMIPLEKPFEE----AYFTHRLWMFFRETRETEEDI 277
Query: 444 --LMTEECSAILQRKMPKKRRDPGSFTIPVEIEGLTMVEALCDLGASINLMSLRMFKRLN 501
+ E + +R KK+ D G F I ++G+ ALC+ GAS+ ++ M L
Sbjct: 278 RRMFCEAREKMKKRITLKKKSDSGKFAISWTMKGIEFPHALCNTGASVIILPRDMADHLG 457
Query: 502 LGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVDFVILDMEEDSKVPLILGRPF 561
L +V P+ S D S ++ GII D+ V++ + PVDF +LD++ + L+L R F
Sbjct: 458 L-KVDPSKESFTFVDCSQRSSGGIIRDLEVQIGNALVPVDFHVLDIKINWNSSLLLRRAF 634
Query: 562 LATGEAEIKVA 572
L+TG ++ A
Sbjct: 635 LSTGRGSVQHA 667
>BG644741
Length = 735
Score = 62.8 bits (151), Expect = 1e-09
Identities = 31/114 (27%), Positives = 49/114 (42%)
Frame = -2
Query: 30 LSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAEKFTTRFFPRSLLRKLKNDIMTFTQSTDE 89
L +FP SL A+ W P S+ TW L + F R++P S + + F E
Sbjct: 587 LRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGE 408
Query: 90 NLYEAWEHFKKLLRKCPQHNLTQAECVAKFYDGLLYSSRFGLDAASSGEFDALP 143
++ +W+ F LR P H + FY G +++ LD + G + P
Sbjct: 407 SVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGSYGVCP 246
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 59.3 bits (142), Expect = 1e-08
Identities = 34/128 (26%), Positives = 64/128 (49%), Gaps = 1/128 (0%)
Frame = -1
Query: 682 EEKSELKSLPSSLKYAYLEEGENKPVI-LNSVLTPLKEEKLLKVLRDHKSALGWTIDDIK 740
+E+ ++ ++ L ENK I + + L + K+ ++LR++ + +D+
Sbjct: 384 QERKAIQPHQEEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMP 205
Query: 741 GISPAICMHKILLEENYKPIVQPQRRLNPSMKDVVRKEIIKLLDAGVIYPISDSEWVSPV 800
G+ P I H+I + P+ RR +P M ++ E+ K +DAG + + EWV+ +
Sbjct: 204 GLDPKIVEHRIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANI 25
Query: 801 QVVPKKGG 808
VPKK G
Sbjct: 24 VPVPKKDG 1
>BG587166 similar to GP|8778801|gb| T32E20.9 {Arabidopsis thaliana}, partial
(2%)
Length = 756
Score = 55.8 bits (133), Expect = 1e-07
Identities = 28/46 (60%), Positives = 37/46 (79%)
Frame = +1
Query: 1587 LFPGKLKSRWSGPFVVKRVFPHGAVEVENPETKNIFTVNGQRLKVY 1632
LFPGKLKSR SGPF VK V P+GA+ + + + ++ FTVNGQR+K+Y
Sbjct: 1 LFPGKLKSR*SGPFKVKEVRPYGAIVLWSTDGRD-FTVNGQRVKLY 135
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 52.4 bits (124), Expect = 1e-06
Identities = 22/71 (30%), Positives = 44/71 (60%)
Frame = -2
Query: 1409 GVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVN 1468
G+P I++D G+HF + F E++ ++ ++P +PQ++GQ E SN+ + L+K ++
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 1469 SSRKDWSRKLD 1479
+ W+ +LD
Sbjct: 502 LKKGCWADELD 470
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 49.3 bits (116), Expect = 1e-05
Identities = 28/61 (45%), Positives = 37/61 (59%), Gaps = 1/61 (1%)
Frame = +1
Query: 1080 ASDLALGAVLCQK-KERVLYVIYYASRVLNEAQRNYTTTEKELLGVVFACEKFRPYILGF 1138
A D +GA L QK +E + I YASR+L A+RNYT E+E L ++A FR Y+ G
Sbjct: 1 APDWGIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGP 180
Query: 1139 K 1139
K
Sbjct: 181 K 183
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 43.9 bits (102), Expect = 5e-04
Identities = 18/41 (43%), Positives = 28/41 (67%)
Frame = -1
Query: 828 WRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLD 868
+R+CIDYR+ N VT K+ +PLP ID + D++ Y+ +D
Sbjct: 202 FRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCYF*NID 80
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 43.5 bits (101), Expect = 7e-04
Identities = 27/106 (25%), Positives = 55/106 (51%), Gaps = 5/106 (4%)
Frame = -2
Query: 1119 KELLGVVFACEKFRPYILGFKVIVHTDHAALRHLFAKQDSKPRLIRWVLLLQEFDLEIID 1178
K + +A ++ R Y++ + + ++++F K R+ RW +LL E+D+E
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1179 RRG-KDNSVADHLSRLEGGACSPI----PIQEEFSDEKLLAVSTKE 1219
++ K + +ADHL A P+ PI+ +F DE+++ + K+
Sbjct: 455 QKAIKGSILADHL------AHQPLEDYRPIKFDFPDEEIMYLKMKD 336
>BE999414
Length = 613
Score = 40.4 bits (93), Expect = 0.006
Identities = 24/65 (36%), Positives = 33/65 (49%)
Frame = +3
Query: 3 DPHAHMEKFIRNCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAEK 62
DP HME I TYR V A++ LF +L+ A W + + S+ +W DL +
Sbjct: 33 DPDEHMEN-IEVVLTYRSVR---GAVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGDLCHE 200
Query: 63 FTTRF 67
FTT F
Sbjct: 201FTTHF 215
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 38.9 bits (89), Expect = 0.017
Identities = 20/88 (22%), Positives = 43/88 (48%)
Frame = +2
Query: 1417 DGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSR 1476
+G + + ++ L + +G +S+ YHP + GQ E N+ + L ++ + WS+
Sbjct: 11 EG*SSLYSNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSK 190
Query: 1477 KLDDALWAYRTAFKTPIGTSPFHLVFGK 1504
A + Y T++ +PF ++G+
Sbjct: 191 AFPWAEYWYNTSYNISAAMTPFKALYGR 274
>BE997611
Length = 547
Score = 38.1 bits (87), Expect = 0.029
Identities = 17/59 (28%), Positives = 33/59 (55%)
Frame = -2
Query: 389 KKNLDKQFSKFLEVFKKLHINIPFSEALEQMPIYAKFMKDILSKRRKLSEVDETILMTE 447
+K + +F+ + K +N+P +E +EQ+P KF++++L LSE + L +E
Sbjct: 444 EKEVPPKFNALHHILKGYKVNMPITETVEQIPCCIKFLQELLKTNANLSEEEFISLSSE 268
>BG582085
Length = 824
Score = 36.2 bits (82), Expect = 0.11
Identities = 26/73 (35%), Positives = 37/73 (50%), Gaps = 2/73 (2%)
Frame = -3
Query: 1160 PRLIRWVLLLQEFDLEIIDRR--GKDNSVADHLSRLEGGACSPIPIQEEFSDEKLLAVST 1217
P +R +LL QEFDLE R G +A R + + I I+E F +E+L A+S
Sbjct: 771 P*RLRCILLCQEFDLESRQERETGMW*LIAFPCLRNKQVTQNEISIKERFPNEQLFAISQ 592
Query: 1218 KEPLPWYVHFANF 1230
+ PW+ NF
Sbjct: 591 R---PWFADMTNF 562
>TC79651 similar to PIR|T52581|T52581 Ca2+-transporting ATPase (EC 3.6.1.38)
ECA3 [imported] - Arabidopsis thaliana, partial (38%)
Length = 1546
Score = 32.3 bits (72), Expect = 1.6
Identities = 18/48 (37%), Positives = 27/48 (55%)
Frame = -1
Query: 493 SLRMFKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPV 540
SLR+FK LN+ T T+I+ RS +G + +V +V KF+ V
Sbjct: 886 SLRLFKALNISTTTSTVIATVDGCRSSNILHG*VVSLVGQVSKFISSV 743
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,874,594
Number of Sequences: 36976
Number of extensions: 737140
Number of successful extensions: 3315
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 3268
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3309
length of query: 1648
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1539
effective length of database: 4,984,343
effective search space: 7670903877
effective search space used: 7670903877
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0335b.5


